Key Points
-
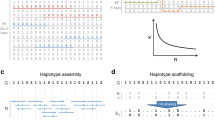
Converting next-generation sequencing (NGS) image files into a set of called SNPs involves a number of steps including image analysis, alignment and assembly, SNP calling and genotype calling.
-
Genotype probabilities for a single individual can be calculated from alignments using recalibrated quality scores.
-
SNP calling and genotype calling is best done using information from multiple individuals simultaneously. The pattern of linkage disequilibrium should be used to call SNPs and genotypes when possible.
-
Analyses of low coverage data can proceed by taking uncertainty in the genotype calls into account, rather than assuming any particular genotype call is correct.
-
The methods used for calling SNPs and for taking uncertainty in SNP genotypes into account can have a strong effect on downstream analyses, including association mapping analyses.
Abstract
Meaningful analysis of next-generation sequencing (NGS) data, which are produced extensively by genetics and genomics studies, relies crucially on the accurate calling of SNPs and genotypes. Recently developed statistical methods both improve and quantify the considerable uncertainty associated with genotype calling, and will especially benefit the growing number of studies using low- to medium-coverage data. We review these methods and provide a guide for their use in NGS studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Metzker, M. Sequencing technologies — the next generation. Nature Rev. Genet. 11, 31–46 (2010). This article provides an excellent Review of NGS technologies and their applications.
Li, R. et al. The sequence and de novo assembly of the giant panda genome. Nature 463, 311–317 (2010).
Ng, S. B. et al. Exome sequencing identifies the cause of a mendelian disorder. Nature Genet. 42, 30–35 (2010).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nature Biotech. 28, 503–510 (2010).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature Biotech. 28, 511–515 (2010).
Liti, G. et al. Population genomics of domestic and wild yeasts. Nature 458, 337–341 (2009).
Li, Y. et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nature Genet. 42, 969–972 (2010).
Durbin, R. M. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010). This 1000Genomes paper provides an application of many of the state-of-the-art methods for analysis of NGS data.
Flicek, P. & Birney, E. Sense from sequence reads: methods for alignment and assembly. Nature Methods 6, S6–S12 (2009).
Kim, S. Y. et al. Design of association studies with pooled or un-pooled next-generation sequencing data. Genet. Epidemiol. 34, 479–491 (2010).
Li, H., Ruan, J. & Durbin, R. M. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008). This paper describes MAQ, a forerunner of efficient, hash-based alignment algorithms for short reads. MAQ also produces genotype calls. The concept of read-mapping quality is introduced in this paper.
Li, J. B. et al. Multiplex padlock targeted sequencing reveal human hypermutable CpG variations. Genome Res. 19, 1606–1615 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Quinlan, A. R. et al. Pyrobayes: an improved base caller for SNP discovery in pyrosequences. Nature Methods 5, 179–181 (2008).
Wu, H, Irizarry, R. A. & Bravo, H. C. Intensity normalization improves color calling in SOLiD sequencing. Nature Methods 7, 336–337 (2010).
Kircher, M., Stenzel, U. & Kelso, J. Improved base calling for the Illumina Genome Analyzer using machine learning strategies. Genome Biol. 10, R83 (2009).
Kao, W. C., Stevens, K. & Song, Y. S. BayesCall: a model-based basecalling algorithm for high-throughput short-read sequencing. Genome Res. 19, 1884–1895 (2009).
Kao, W. C. & Song, Y. S. naiveBayesCall: an efficient model-based base-calling algorithm for high-throughput sequencing. Lect. Notes Comp. Sci. 6044, 233–247 (2010).
Burrows, M. & Wheeler, D. A block-sorting lossless data compression algorithm. Technical Report 124, Digital Equipment Corporation. HP Labs Technical Reports [online], (1994).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Lunter, G. & Goodson, M. Stampy: a statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome Res. 27 Oct 2010 (doi:10.1101/gr.111120.110).
Sundquist, A., Ronaghi, M., Tang, H., Pevzner, P. & Batzoglou, S. Whole-genome sequencing and assembly with high-throughput, short-read technologies. PLoS ONE 2, e484 (2007).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Butler, J. et al. ALLPATHS: de novo assembly of whole-genome shotgun microreads. Genome Res. 18, 810–820 (2008).
Simpson, J. T. et al. ABySS: a parallel assembler for short read sequence data. Genome Res. 19, 1117–1123 (2009).
Chaisson, M. J. P., Brinza, D. & Pevzner, P. A. De novo fragment assembly with short mate-paired reads: does the read length matter? Genome Res. 19, 336–346 (2009).
Brockman, W. et al. Quality scores and SNP detection in sequencing-by-synthesis systems. Genome Res. 18, 763–770 (2008).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 10 Apr 2011 (doi:10.1038/ng.806).
Harismendy, O. et al. Evaluation of next generation sequencing platforms for population targeted sequencing studies. Genome Biol. 10, R32 (2009).
Wang, J. et al. The diploid sequence of an Asian individual. Nature 456, 60–65 (2009).
Hedges, D. et al. Exome sequencing of a multigenerational human pedigree. PLoS ONE 4, e8232 (2009).
Martin, E. R. et al. SeqEM: an adaptive genotype-calling approach for next- generation sequencing studies. Bioinformatics 26, 2803–2810 (2010).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Dai, J. Y. et al. Imputation methods to improve inference in SNP association studies. Genet. Epidemiol. 30, 690–702 (2006).
Minichiello, M. J. & Durbin, R. Mapping trait loci by use of inferred ancestral recombination graphs. Am. J. Hum. Genet. 79, 910–922 (2006).
Scheet, P. & Stephens, M. A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. Am. J. Hum. Genet. 78, 629–644 (2006).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnely, P. A new multipoint method for genome-wide association studies via imputation of genotypes. Nature Genet. 39, 906–913 (2007).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nature Rev. Genet. 11, 499–511 (2010). This Review provides a comprehensive overview of available statistical methods for imputing genotypes and discusses various uses of imputation.
Huang, L. et al. The relationship between imputation error and statistical power in genetic association studies in diverse populations. Am. J. Hum. Genet. 85, 692–698 (2009).
Schaid, D. J., Rowland, C. M., Tines, D. E., Jacobson, R. M. & Poland, G. A. Score tests for association between traits and haplotypes when linkage phase is ambiguous. Am. J. Hum. Genet. 70, 425–434 (2002).
Servin, B. & Stephens, M. Imputation-based analysis of association studies: candidate genes and quantitative traits. PLoS Genet. 3, e114 (2007).
Hellmann, I. et al. Population genetic analysis of shotgun assemblies of genomic sequences from multiple individuals. Genome Res. 18, 1020–1029 (2008).
Johnson, P. L. F. & Slatkin, M. Accounting for bias from sequencing error in population genetic estimates. Mol. Biol. Evol. 25, 199–206 (2008).
Johnson, P. L. F. & Slatkin, M. Inference of population genetic parameters in metagenomics. A clean look at messy data. Genome Res. 16, 1320–1327 (2006).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Li, H. et al. The sequence alignment/map (SAM) format and SAMtools. Bioinformatics. 25, 2078–2079 (2009).
Le, S. Q. & Durbin, R. SNP detection and genotyping from low-coverage sequencing data on multiple diploid samples. Genome Res. 27 Oct 2010 (doi:10.1101/gr.113084.110).
Acknowledgements
This work was supported in part by NIH grants NIGMS R01-HG003229–05 and R01-HG003229–0551 to R.N., an NSF CAREER grant DBI-0846015 to Y.S.S. and an NIH National Research Service Award Trainee appointment on T32-HG00047 to J.S.P.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
FURTHER INFORMATION
Glossary
- Likelihoods
-
Functions expressing the probability of observing the data — for example, next-generation sequencing data — given a parameter, such as a genotype or an allele frequency.
- Posterior probabilities
-
In this context, these are the probabilities of a particular genotype given observed data: they are calculated by incorporating the information from the next-generation sequencing data as well as some prior information.
- Hashing
-
A procedure of creating a data structure that helps to accelerate alignment. It stores information about which reads or where in the reference genome a particular substring or subsequence occurs. Some hash-based aligners hash the reads, while others hash the reference genome.
- Paired-end reads
-
Sequencing of both the forward and reverse template of a DNA molecule, which is possible by inserting a primer sequence between the two ends of the read. The use of this technique greatly helps to increase assembly and alignment accuracy.
- CEU individuals
-
One of the 11 populations in HapMap phase three. It consists of Utah residents with Northern and Western European ancestry from the Centre d'Etude du Polymorphisme Humain (CEPH) collection.
- Bayes' formula
-
A mathematical expression showing that a posterior probability can be found as the prior probability multiplied by the likelihood divided by a constant.
- Correlated errors
-
Errors that do not occur independently of each other. An error that is observed in one position might increase the chance of observing another error in a neighbouring position.
- Prior probability
-
In the context of this Review, the probability of a genotype calculated without incorporating information from the next-generation sequencing data. Prior probabilities can be obtained from a set of reference data.
- Maximum likelihood
-
The statistical principle of estimating a parameter by finding the value of the parameters that maximizes the likelihood function.
- Imputation
-
The substitution of some value for a missing data point. In this context, it is the use of a set of reference haplotypes to infer a genotype for an individual, when data are missing or incomplete.
- Likelihood ratio test
-
A method for testing statistical hypotheses based on comparing the maximum likelihood under two different models. In this context, the allele frequency in one model equals zero, whereas the frequency in the second model might be larger than zero.
Rights and permissions
About this article
Cite this article
Nielsen, R., Paul, J., Albrechtsen, A. et al. Genotype and SNP calling from next-generation sequencing data. Nat Rev Genet 12, 443–451 (2011). https://doi.org/10.1038/nrg2986
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2986
This article is cited by
-
A high-performance computational workflow to accelerate GATK SNP detection across a 25-genome dataset
BMC Biology (2024)
-
A cautionary tale of low-pass sequencing and imputation with respect to haplotype accuracy
Genetics Selection Evolution (2024)
-
Signatures of adaptation at key insecticide resistance loci in Anopheles gambiae in Southern Ghana revealed by reduced-coverage WGS
Scientific Reports (2024)
-
Investigating the potential roles of intra-colonial genetic variability in Pocillopora corals using genomics
Scientific Reports (2024)
-
Identification of eQTLs using different sets of single nucleotide polymorphisms associated with carcass and body composition traits in pigs
BMC Genomics (2024)