Key Points
-
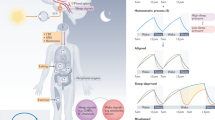
Circadian rhythms in mammals are regulated by a master circadian pacemaker located in the hypothalamic suprachiasmatic nucleus (SCN), which coordinates rhythmic processes throughout the organism.
-
Circadian clocks are cell autonomous and these cellular clocks are located in SCN neurons as well as in almost every cell in the body.
-
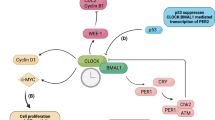
The molecular mechanism of circadian clocks in mammals involves an autoregulatory transcriptional feedback loop involving the positive elements CLOCK and BMAL1, which transcriptionally activate the negative feedback elements period (PER) and cryptochrome (CRY), which inhibit their own transcription by repressing the CLOCK–BMAL1 complex.
-
Post-translational regulation (for example, phosphorylation, acetylation and ubiquitylation) of clock proteins have important roles in regulating the stability, localization and turnover of clock components.
-
The sleep disorder familial advanced sleep phase syndrome (FASPS) has been found to be caused by mutations in two core clock genes, period homologue 2 (PER2) and casein kinase 1 delta (CSNK1D), in humans.
-
There is weak but emerging evidence for allelic variants in clock genes to be associated with diurnal preference, mood disorders, sleep and metabolic disorders.
-
Peripheral circadian oscillators are controlled by signals arising from the SCN and from proximal signals related to feeding behaviour, hormonal signals and body-temperature fluctuations.
-
In addition to their primary role in the generation of circadian rhythms, recent work has shown that circadian clock genes can affect a wide variety of other physiological processes.
-
Emerging examples of circadian regulation of physiological pathways include diverse aspects of cellular metabolism, cell growth and DNA-damage control, xenobiotic responses, and the modulation of behavioural responses to drugs and alcohol.
-
The knowledge that circadian clocks are cell autonomous and distributed throughout the body provide a new perspective to target central as well as peripheral circadian oscillators for therapeutic intervention.
Abstract
Circadian cycles affect a variety of physiological processes, and disruptions of normal circadian biology therefore have the potential to influence a range of disease-related pathways. The genetic basis of circadian rhythms is well studied in model organisms and, more recently, studies of the genetic basis of circadian disorders has confirmed the conservation of key players in circadian biology from invertebrates to humans. In addition, important advances have been made in understanding how these molecules influence physiological functions in tissues throughout the body. Together, these studies set the scene for applying our knowledge of circadian biology to the understanding and treatment of a range of human diseases, including cancer and metabolic and behavioural disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bell-Pedersen, D. et al. Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nature Rev. Genet. 6, 544–556 (2005).
Lowrey, P. L. & Takahashi, J. S. Mammalian circadian biology: elucidating genome-wide levels of temporal organization. Annu. Rev. Genomics Hum. Genet. 5, 407–441 (2004).
Hastings, M. H., Reddy, A. B. & Maywood, E. S. A clockwork web: circadian timing in brain and periphery, in health and disease. Nature Rev. Neurosci. 4, 649–661 (2003).
Antle, M. C. & Silver, R. Orchestrating time: arrangements of the brain circadian clock. Trends Neurosci. 28, 145–151 (2005).
Herzog, E. D. Neurons and networks in daily rhythms. Nature Rev. Neurosci. 8, 790–802 (2007).
Berson, D. M. Phototransduction in ganglion-cell photoreceptors. Pflugers Arch. 454, 849–855 (2007).
Hankins, M. W., Peirson, S. N. & Foster, R. G. Melanopsin: an exciting photopigment. Trends Neurosci. 31, 27–36 (2008).
Saper, C. B., Scammell, T. E. & Lu, J. Hypothalamic regulation of sleep and circadian rhythms. Nature 437, 1257–1263 (2005).
Kramer, A. et al. Regulation of daily locomotor activity and sleep by hypothalamic EGF receptor signaling. Science 294, 2511–2515 (2001).
Kraves, S. & Weitz, C. J. A role for cardiotrophin-like cytokine in the circadian control of mammalian locomotor activity. Nature Neurosci. 9, 212–219 (2006).
Cheng, M. Y. et al. Prokineticin 2 transmits the behavioural circadian rhythm of the suprachiasmatic nucleus. Nature 417, 405–410 (2002).
Li, J. D. et al. Attenuated circadian rhythms in mice lacking the prokineticin 2 gene. J. Neurosci. 26, 11615–11623 (2006).
Prosser, H. M. et al. Prokineticin receptor 2 (Prokr2) is essential for the regulation of circadian behavior by the suprachiasmatic nuclei. Proc. Natl Acad. Sci. USA 104, 648–653 (2007).
Lee, C., Etchegaray, J. P., Cagampang, F. R., Loudon, A. S. & Reppert, S. M. Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107, 855–867 (2001).
Preitner, N. et al. The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110, 251–260 (2002).
Sato, T. K. et al. A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron 43, 527–537 (2004).
Gallego, M. & Virshup, D. M. Post-translational modifications regulate the ticking of the circadian clock. Nature Rev. Mol. Cell Biol. 8, 139–148 (2007).
Lowrey, P. L. et al. Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 288, 483–492 (2000).
Meng, Q. J. et al. Setting clock speed in mammals: the CK1 epsilon tau mutation in mice accelerates circadian pacemakers by selectively destabilizing PERIOD proteins. Neuron 58, 78–88 (2008).
Eide, E. J. et al. Control of mammalian circadian rhythm by CKIepsilon-regulated proteasome-mediated PER2 degradation. Mol. Cell Biol. 25, 2795–2807 (2005).
Shirogane, T., Jin, J., Ang, X. L. & Harper, J. W. SCFbeta-TRCP controls clock-dependent transcription via casein kinase 1-dependent degradation of the mammalian period-1 (Per1) protein. J. Biol. Chem. 280, 26863–26872 (2005).
Reischl, S. et al. Beta-TrCP1-mediated degradation of PERIOD2 is essential for circadian dynamics. J. Biol. Rhythms 22, 375–386 (2007).
Busino, L. et al. SCFFbxl3 controls the oscillation of the circadian clock by directing the degradation of cryptochrome proteins. Science 316, 900–904 (2007).
Siepka, S. M. et al. Circadian mutant Overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell 129, 1011–1023 (2007).
Godinho, S. I. et al. The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science 316, 897–900 (2007). References 23–25 identify a novel SCF E3 ubiquitin ligase complex in which the F-box protein FBXL3 interacts specifically with the CRY proteins to target them for degradation by the proteasome.
Etchegaray, J. P., Lee, C., Wade, P. A. & Reppert, S. M. Rhythmic histone acetylation underlies transcription in the mammalian circadian clock. Nature 421, 177–182 (2003).
Ripperger, J. A. & Schibler, U. Rhythmic CLOCK-BMAL1 binding to multiple E-box motifs drives circadian Dbp transcription and chromatin transitions. Nature Genet. 38, 369–374 (2006).
Doi, M., Hirayama, J. & Sassone-Corsi, P. Circadian regulator CLOCK is a histone acetyltransferase. Cell 125, 497–508 (2006).
Hirayama, J. et al. CLOCK-mediated acetylation of BMAL1 controls circadian function. Nature 450, 1086–1090 (2007).
Nakahata, Y. et al. The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell 134, 329–340 (2008).
Asher, G. et al. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell 134, 317–328 (2008). References 30–31 show that the NAD+ dependent deacetylase SIRT1 interacts with the CLOCK protein and has a role in the regulation of BMAL1 and PER2 deacetylation.
Roenneberg, T., Wirz-Justice, A. & Merrow, M. Life between clocks: daily temporal patterns of human chronotypes. J. Biol. Rhythms 18, 80–90 (2003).
Roenneberg, T. et al. Epidemiology of the human circadian clock. Sleep Med. Rev. 11, 429–438 (2007).
Borbely, A. A. & Achermann, P. Sleep homeostasis and models of sleep regulation. J. Biol. Rhythms 14, 557–568 (1999).
Sack, R. L. et al. Circadian rhythm sleep disorders: part II, advanced sleep phase disorder, delayed sleep phase disorder, free-running disorder, and irregular sleep–wake rhythm. An American Academy of Sleep Medicine review. Sleep 30, 1484–1501 (2007).
Jones, C. R. et al. Familial advanced sleep-phase syndrome: A short-period circadian rhythm variant in humans. Nature Med. 5, 1062–1065 (1999).
Reid, K. J. et al. Familial advanced sleep phase syndrome. Arch. Neurol. 58, 1089–1094 (2001).
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001).
Xu, Y. et al. Modeling of a human circadian mutation yields insights into clock regulation by PER2. Cell 128, 59–70 (2007).
Vanselow, K. et al. Differential effects of PER2 phosphorylation: molecular basis for the human familial advanced sleep phase syndrome (FASPS). Genes Dev. 20, 2660–2672 (2006).
Gallego, M., Eide, E. J., Woolf, M. F., Virshup, D. M. & Forger, D. B. An opposite role for tau in circadian rhythms revealed by mathematical modeling. Proc. Natl Acad. Sci. USA 103, 10618–10623 (2006).
Satoh, K., Mishima, K., Inoue, Y., Ebisawa, T. & Shimizu, T. Two pedigrees of familial advanced sleep phase syndrome in Japan. Sleep 26, 416–417 (2003).
Xu, Y. et al. Functional consequences of a CKIdelta mutation causing familial advanced sleep phase syndrome. Nature 434, 640–644 (2005). References 38–40 and 43 provide the first evidence that the human disorder FASPS involves mutations in the core circadian genes, PER2 and CSNK1D.
Ancoli-Israel, S., Schnierow, B., Kelsoe, J. & Fink, R. A pedigree of one family with delayed sleep phase syndrome. Chronobiol. Int. 18, 831–840 (2001).
Iwase, T. et al. Mutation screening of the human Clock gene in circadian rhythm sleep disorders. Psychiatry Res. 109, 121–128 (2002).
Katzenberg, D. et al. A CLOCK polymorphism associated with human diurnal preference. Sleep 21, 569–576 (1998).
Robilliard, D. L. et al. The 3111 Clock gene polymorphism is not associated with sleep and circadian rhythmicity in phenotypically characterized human subjects. J. Sleep Res. 11, 305–312 (2002).
Mishima, K., Tozawa, T., Satoh, K., Saitoh, H. & Mishima, Y. The 3111T/C polymorphism of hClock is associated with evening preference and delayed sleep timing in a Japanese population sample. Am. J. Med. Genet. B Neuropsychiatr. Genet. 133, 101–104 (2005).
Takano, A. et al. A missense variation in human casein kinase I epsilon gene that induces functional alteration and shows an inverse association with circadian rhythm sleep disorders. Neuropsychopharmacology 29, 1901–1909 (2004).
Carpen, J. D., Archer, S. N., Skene, D. J., Smits, M. & von Schantz, M. A single-nucleotide polymorphism in the 5′-untranslated region of the hPER2 gene is associated with diurnal preference. J. Sleep Res. 14, 293–297 (2005).
Archer, S. N. et al. A length polymorphism in the circadian clock gene Per3 is linked to delayed sleep phase syndrome and extreme diurnal preference. Sleep 26, 413–415 (2003).
Pereira, D. S. et al. Association of the length polymorphism in the human Per3 gene with the delayed sleep-phase syndrome: does latitude have an influence upon it? Sleep 28, 29–32 (2005).
Sandrelli, F. et al. A molecular basis for natural selection at the timeless locus in Drosophila melanogaster. Science 316, 1898–1900 (2007).
Viola, A. U. et al. PER3 polymorphism predicts sleep structure and waking performance. Curr. Biol. 17, 613–618 (2007).
Naylor, E. et al. The circadian clock mutation alters sleep homeostasis in the mouse. J. Neurosci. 20, 8138–8143 (2000).
Franken, P., Lopez-Molina, L., Marcacci, L., Schibler, U. & Tafti, M. The transcription factor DBP affects circadian sleep consolidation and rhythmic EEG activity. J. Neurosci. 20, 617–625 (2000).
Wisor, J. P. et al. A role for cryptochromes in sleep regulation. BMC Neurosci. 3, 20 (2002).
Laposky, A. et al. Deletion of the mammalian circadian clock gene BMAL1/Mop3 alters baseline sleep architecture and the response to sleep deprivation. Sleep 28, 395–409 (2005).
Franken, P. et al. NPAS2 as a transcriptional regulator of non-rapid eye movement sleep: genotype and sex interactions. Proc. Natl Acad. Sci. USA 103, 7118–7123 (2006).
Taheri, S., Zeitzer, J. M. & Mignot, E. The role of hypocretins (orexins) in sleep regulation and narcolepsy. Annu. Rev. Neurosci. 25, 283–313 (2002).
Sakurai, T. The neural circuit of orexin (hypocretin): maintaining sleep and wakefulness. Nature Rev. Neurosci. 8, 171–181 (2007).
Hendricks, J. C. & Sehgal, A. Why a fly? Using Drosophila to understand the genetics of circadian rhythms and sleep. Sleep 27, 334–342 (2004).
Cirelli, C. et al. Reduced sleep in Drosophila Shaker mutants. Nature 434, 1087–1092 (2005).
Dauvilliers, Y., Maret, S. & Tafti, M. Genetics of normal and pathological sleep in humans. Sleep Med. Rev. 9, 91–100 (2005).
Mignot, E. Why we sleep: the temporal organization of recovery. PLoS Biol. 6, e106 (2008).
Levi, F. & Schibler, U. Circadian rhythms: mechanisms and therapeutic implications. Annu. Rev. Pharmacol. Toxicol. 47, 593–628 (2007).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Storch, K. F. et al. Extensive and divergent circadian gene expression in liver and heart. Nature 417, 78–83 (2002).
Ueda, H. R. et al. A transcription factor response element for gene expression during circadian night. Nature 418, 534–539 (2002).
Yamazaki, S. et al. Resetting central and peripheral circadian oscillators in transgenic rats. Science 288, 682–685 (2000).
Yoo, S. H. et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl Acad. Sci. USA 101, 5339–5346 (2004).
Nagoshi, E. et al. Circadian gene expression in individual fibroblasts: cell-autonomous and self-sustained oscillators pass time to daughter cells. Cell 119, 693–705 (2004).
Welsh, D. K., Yoo, S. H., Liu, A. C., Takahashi, J. S. & Kay, S. A. Bioluminescence imaging of individual fibroblasts reveals persistent, independently phased circadian rhythms of clock gene expression. Curr. Biol. 14, 2289–2295 (2004).
Damiola, F., Le Minh, N., Preitner, N. & Kornmann, B. Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev. 14, 2950–2961 (2000).
Stokkan, K. A., Yamazaki, S., Tei, H., Sakaki, Y. & Menaker, M. Entrainment of the circadian clock in the liver by feeding. Science 291, 490–493 (2001).
Balsalobre, A. et al. Resetting of circadian time in peripheral tissues by glucocorticoid signaling. Science 289, 2344–2347 (2000).
Brown, S. A., Zumbrunn, G., Fleury-Olela, F. & Preitner, N. Rhythms of mammalian body temperature can sustain peripheral circadian clocks. Curr. Biol. 12, 1574–1583 (2002).
Bunger, M. K. et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000).
McDearmon, E. L. et al. Dissecting the functions of the mammalian clock protein BMAL1 by tissue-specific rescue in mice. Science 314, 1304–1308 (2006).
Kornmann, B., Schaad, O., Bujard, H., Takahashi, J. S. & Schibler, U. System-driven and oscillator-dependent circadian transcription in mice with a conditionally active liver clock. PLoS Biol. 5, e34 (2007). Conditional inactivation of the cell-autonomous clock in the liver reveals the role of local control versus systemic cues for regulating cycling gene expression in the liver.
Ramsey, K. M., Marcheva, B., Kohsaka, A. & Bass, J. The clockwork of metabolism. Annu. Rev. Nutr. 27, 219–240 (2007).
Rutter, J., Reick, M. & McKnight, S. L. Metabolism and the control of circadian rhythms. Annu. Rev. Biochem. 71, 307–331 (2002).
Wijnen, H. & Young, M. W. Interplay of circadian clocks and metabolic rhythms. Annu. Rev. Genet. 40, 409–448 (2006).
Yang, X. et al. Nuclear receptor expression links the circadian clock to metabolism. Cell 126, 801–810 (2006). This comprehensive analysis of nuclear receptor gene expression reveals extensive circadian regulation of this superfamily of proteins.
Turek, F. W. et al. Obesity and metabolic syndrome in circadian Clock mutant mice. Science 308, 1043–1045 (2005).
Scott, E. M., Carter, A. M. & Grant, P. J. Association between polymorphisms in the Clock gene, obesity and the metabolic syndrome in man. Int. J. Obes. (Lond.) 32, 658–662 (2007).
Rudic, R. D. et al. BMAL1 and CLOCK, two essential components of the circadian clock, are involved in glucose homeostasis. PLoS Biol. 2, e377 (2004).
Woon, P. Y. et al. Aryl hydrocarbon receptor nuclear translocator-like (BMAL1) is associated with susceptibility to hypertension and type 2 diabetes. Proc. Natl Acad. Sci. USA 104, 14412–14417 (2007).
Shimba, S. et al. Brain and muscle Arnt-like protein-1 (BMAL1), a component of the molecular clock, regulates adipogenesis. Proc. Natl Acad. Sci. USA 102, 12071–12076 (2005).
Bunger, M. K. et al. Progressive arthropathy in mice with a targeted disruption of the Mop3/Bmal-1 locus. Genesis 41, 122–132 (2005).
Kondratov, R. V., Kondratova, A. A., Gorbacheva, V. Y., Vykhovanets, O. V. & Antoch, M. P. Early aging and age-related pathologies in mice deficient in BMAL1, the core component of the circadian clock. Genes Dev. 20, 1868–1873 (2006).
Lucas, R. J., Stirland, J. A., Mohammad, Y. N. & Loudon, A. S. Postnatal growth rate and gonadal development in circadian tau mutant hamsters reared in constant dim red light. J. Reprod. Fertil. 118, 327–330 (2000).
Oklejewicz, M., Hut, R. A., Daan, S., Loudon, A. S. & Stirland, A. J. Metabolic rate changes proportionally to circadian frequency in tau mutant Syrian hamsters. J. Biol. Rhythms 12, 413–422 (1997).
Green, C. B. et al. Loss of nocturnin, a circadian deadenylase, confers resistance to hepatic steatosis and diet-induced obesity. Proc. Natl Acad. Sci. USA 104, 9888–9893 (2007).
Rutter, J., Reick, M., Wu, L. C. & McKnight, S. L. Regulation of clock and NPAS2 DNA binding by the redox state of NAD cofactors. Science 293, 510–514 (2001).
Dioum, E. M. et al. NPAS2: a gas-responsive transcription factor. Science 298, 2385–2387 (2002).
Raghuram, S. et al. Identification of heme as the ligand for the orphan nuclear receptors REV-ERBalpha and REV-ERBbeta. Nature Struct. Mol. Biol. 14, 1207–1213 (2007).
Yin, L. et al. Rev-erbα, a heme sensor that coordinates metabolic and circadian pathways. Science 318, 1786–1789 (2007).
Kaasik, K. & Lee, C. C. Reciprocal regulation of haem biosynthesis and the circadian clock in mammals. Nature 430, 467–471 (2004).
Liu, C., Li, S., Liu, T., Borjigin, J. & Lin, J. D. Transcriptional coactivator PGC-1alpha integrates the mammalian clock and energy metabolism. Nature 447, 477–481 (2007). This paper identifies a key link between the circadian and metabolic systems.
Kohsaka, A. et al. High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab. 6, 414–421 (2007).
Pittendrigh, C. S. Temporal organization: reflections of a Darwinian clock-watcher. Annu. Rev. Physiol. 55, 16–54 (1993).
Matsuo, T. et al. Control mechanism of the circadian clock for timing of cell division in vivo. Science 302, 255–259 (2003).
Miller, B. H. et al. Circadian and CLOCK-controlled regulation of the mouse transcriptome and cell proliferation. Proc. Natl Acad. Sci. USA 104, 3342–3347 (2007).
Fu, L., Pelicano, H., Liu, J., Huang, P. & Lee, C. The circadian gene Period2 plays an important role in tumor suppression and DNA damage response in vivo. Cell 111, 41–50 (2002).
Fu, L., Patel, M. S., Bradley, A., Wagner, E. F. & Karsenty, G. The molecular clock mediates leptin-regulated bone formation. Cell 122, 803–15 (2005).
Gery, S. et al. The circadian gene per1 plays an important role in cell growth and DNA damage control in human cancer cells. Mol. Cell 22, 375–382 (2006).
Unsal-Kacmaz, K., Mullen, T. E., Kaufmann, W. K. & Sancar, A. Coupling of human circadian and cell cycles by the timeless protein. Mol. Cell Biol. 25, 3109–3116 (2005).
Oklejewicz, M. et al. Phase resetting of the mammalian circadian clock by DNA damage. Curr. Biol. 18, 286–291 (2008).
Pregueiro, A. M., Liu, Q., Baker, C. L., Dunlap, J. C. & Loros, J. J. The Neurospora checkpoint kinase 2: a regulatory link between the circadian and cell cycles. Science 313, 644–649 (2006).
Filipski, E. et al. Host circadian clock as a control point in tumor progression. J. Natl Cancer Inst. 94, 690–697 (2002).
Gorbacheva, V. Y. et al. Circadian sensitivity to the chemotherapeutic agent cyclophosphamide depends on the functional status of the CLOCK/BMAL1 transactivation complex. Proc. Natl Acad. Sci. USA 102, 3407–3412 (2005).
Schernhammer, E. S. et al. Rotating night shifts and risk of breast cancer in women participating in the nurses' health study. J. Natl Cancer Inst. 93, 1563–1568 (2001).
Megdal, S. P., Kroenke, C. H., Laden, F., Pukkala, E. & Schernhammer, E. S. Night work and breast cancer risk: a systematic review and meta-analysis. Eur. J. Cancer 41, 2023–2032 (2005).
Kubo, T. et al. Prospective cohort study of the risk of prostate cancer among rotating-shift workers: findings from the Japan collaborative cohort study. Am. J. Epidemiol. 164, 549–555 (2006).
Antoch, M. P., Kondratov, R. V. & Takahashi, J. S. Circadian clock genes as modulators of sensitivity to genotoxic stress. Cell Cycle 4, 901–907 (2005).
Lis, C. G. et al. Circadian timing in cancer treatment: the biological foundation for an integrative approach. Integr. Cancer Ther. 2, 105–111 (2003).
Méndez-Ferrer, S., Lucas, D., Battista, M. & Frenette, P. S. Haematopoietic stem cell release is regulated by circadian oscillations. Nature 452, 442–447 (2008). This paper describes the surprising observation of circadian regulation of haematopoietic stem-cell release and transplantation efficiency.
Spanagel, R. et al. The clock gene Per2 influences the glutamatergic system and modulates alcohol consumption. Nature Med. 11, 35–42 (2005).
McClung, C. A. et al. Regulation of dopaminergic transmission and cocaine reward by the Clock gene. Proc. Natl Acad. Sci. USA 102, 9377–9381 (2005).
Roybal, K. et al. Mania-like behavior induced by disruption of CLOCK. Proc. Natl Acad. Sci. USA 104, 6406–6411 (2007).
Wehr, T. A., Sack, D., Rosenthal, N., Duncan, W. & Gillin, J. C. Circadian rhythm disturbances in manic-depressive illness. Fed. Proc. 42, 2809–2814 (1983).
Jones, S. H. Circadian rhythms, multilevel models of emotion and bipolar disorder — an initial step towards integration? Clin. Psychol. Rev. 21, 1193–1209 (2001).
Johansson, C. et al. Circadian clock-related polymorphisms in seasonal affective disorder and their relevance to diurnal preference. Neuropsychopharmacology 28, 734–739 (2003).
Nievergelt, C. M. et al. Suggestive evidence for association of the circadian genes PERIOD3 and ARNTL with bipolar disorder. Am. J. Med. Genet. B Neuropsychiatr. Genet. 141, 234–241 (2006).
Desan, P. H. et al. Genetic polymorphism at the CLOCK gene locus and major depression. Am. J. Med. Genet. 96, 418–421 (2000).
Bailer, U. et al. No association of clock gene T3111C polymorphism and affective disorders. Eur. Neuropsychopharmacol. 15, 51–55 (2005).
Nievergelt, C. M. et al. Examination of the clock gene Cryptochrome 1 in bipolar disorder: mutational analysis and absence of evidence for linkage or association. Psychiatr. Genet. 15, 45–52 (2005).
Taheri, S., Lin, L., Austin, D., Young, T. & Mignot, E. Short sleep duration is associated with reduced leptin, elevated ghrelin, and increased body mass index. PLoS Med. 1, e62 (2004).
Spiegel, K., Tasali, E., Penev, P. & Van Cauter, E. Brief communication: sleep curtailment in healthy young men is associated with decreased leptin levels, elevated ghrelin levels, and increased hunger and appetite. Ann. Intern. Med. 141, 846–850 (2004).
Ancoli-Israel, S. et al. The role of actigraphy in the study of sleep and circadian rhythms. Sleep 26, 342–392 (2003).
Horne, J. A. & Ostberg, O. A self-assessment questionnaire to determine morningness–eveningness in human circadian rhythms. Int. J. Chronobiol. 4, 97–110 (1976).
Duffy, J. F., Rimmer, D. W. & Czeisler, C. A. Association of intrinsic circadian period with morningness–eveningness, usual wake time, and circadian phase. Behav. Neurosci. 115, 895–899 (2001).
Aschoff, J. Circadian rhythms in man. Science 148, 1427–1432 (1965).
Czeisler, C. A. et al. Stability, precision, and near-24-hour period of the human circadian pacemaker. Science 284, 2177–2181 (1999).
Sack, R. L. et al. Circadian rhythm sleep disorders: part I, basic principles, shift work and jet lag disorders. An American Academy of Sleep Medicine review. Sleep 30, 1460–1483 (2007).
Brown, S. A. et al. The period length of fibroblast circadian gene expression varies widely among human individuals. PLoS Biol. 3, e338 (2005).
Brown, S. A., Kunz, D., Dumas, A. & Westermark, P. O. Molecular insights into human daily behavior. Proc. Natl Acad. Sci. USA 105, 1602–1607 (2008).
Liu, A. C. et al. Intercellular coupling confers robustness against mutations in the SCN circadian clock network. Cell 129, 605–616 (2007).
Ko, C. H. & Takahashi, J. S. Molecular components of the mammalian circadian clock. Hum. Mol. Genet. 15 (Suppl 2), R271–R277 (2006).
Acknowledgements
We thank the anonymous reviewers for their editorial suggestions. Research is supported by grants from the National Institutes of Health (U01 MH61915 and R01 MH078024) and a Silvio O. Conte Center grant (P50 MH074924) to J.S.T. J.S.T. is an Investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary information S1 (table)
Human circadian gene mutation and associated phenotypic effects/disorders. (PDF 0 kb)
Supplementary information S2 (table)
Mouse circadian mutants and observed circadian and physiological phenotypes. (PDF 164 kb)
Related links
Related links
DATABASES
OMIM
familial advanced sleep-phase syndrome
FURTHER INFORMATION
Joseph S. Takahashi at Northwestern University
Glossary
- Suprachiasmatic nucleus
-
(SCN). The master circadian pacemaker in the mammalian brain, which is composed of a group of neurons located in the anterior hypothalamus immediately dorsal to the optic chiasm.
- Retinal ganglion cells
-
A class of neuronal cells in the mammalian retina that relay information to the central nervous system via the optic tracts.
- Sleep homeostatic process
-
A formal description of a control process underlying the drive to sleep, typically estimated from the intensity of slow-wave sleep as measured from the Fourier spectrum of electroencephagram recordings from the brain.
- Narcolepsy
-
A sleep disorder characterized by sleepiness, cataplexy and abnormal transitions from wakefulness into rapid eye movement sleep.
- Hyperphagia
-
Excessive eating.
- Arcuate nucleus
-
A group of neurons in the mediobasal hypothalamus of the brain that are involved in a number of neuroendocrine functions and feeding.
- Hyperlipidaemia
-
Elevation of lipids (fats) in the bloodstream.
- Hepatic steatosis
-
Fatty liver.
- Hyperglycaemia
-
Elevation of glucose in the bloodstream.
- Hypoinsulinaemia
-
Abnormally low insulin in the bloodstream.
- Sarcopaenia
-
Age-related loss of skeletal muscle mass and strength.
Rights and permissions
About this article
Cite this article
Takahashi, J., Hong, HK., Ko, C. et al. The genetics of mammalian circadian order and disorder: implications for physiology and disease. Nat Rev Genet 9, 764–775 (2008). https://doi.org/10.1038/nrg2430
Issue Date:
DOI: https://doi.org/10.1038/nrg2430
This article is cited by
-
The nature and distribution of putative non-functional alleles suggest only two independent events at the origins of Astyanax mexicanus cavefish populations
BMC Ecology and Evolution (2024)
-
Synthesis of causal and surrogate models by non-equilibrium thermodynamics in biological systems
Scientific Reports (2024)
-
Pressure Building Against the Clock: The Impact of Circadian Misalignment on Blood Pressure
Current Hypertension Reports (2024)
-
Role of polygenic risk scores in the association between chronotype and health risk behaviors
BMC Psychiatry (2023)
-
Genetics of circadian rhythms and sleep in human health and disease
Nature Reviews Genetics (2023)