Key Points
-
Ancient DNA provides transformative insight into the history of human adaptation via the ability to directly track genetic variant frequency changes across space and time.
-
Analyses of human, archaic hominin, and domesticated plant and animal ancient genomic data sets can each inform our understanding of past human evolution and behaviour.
-
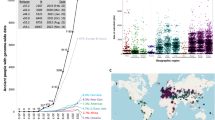
The number of published ancient genomic data sets is growing substantially each year, contributing expanded precision and power to evolutionary analyses based on these data.
-
Human ancient genome data have already helped characterize the histories of biological adaptations to northern latitudes and cold climates, agriculture-associated dietary shifts, and a changing infectious disease landscape.
-
After migrating out of Africa, ancient human populations acquired genetic variants conferring fitness advantages in Eurasian environments through adaptive introgression with archaic hominin populations who had already been inhabiting this region for hundreds of thousands of years.
-
Ancient genome data reveal some substantial time lags between documented environmental or cultural changes and the appearance and spread of genetic variants associated with human biological adaptations, with possible implications for intervening human health and/or potential compensatory cultural behaviours.
Abstract
The past several years have witnessed an explosion of successful ancient human genome-sequencing projects, with genomic-scale ancient DNA data sets now available for more than 1,100 ancient human and archaic hominin (for example, Neandertal) individuals. Recent 'evolution in action' analyses have started using these data sets to identify and track the spatiotemporal trajectories of genetic variants associated with human adaptations to novel and changing environments, agricultural lifestyles, and introduced or co-evolving pathogens. Together with evidence of adaptive introgression of genetic variants from archaic hominins to humans and emerging ancient genome data sets for domesticated animals and plants, these studies provide novel insights into human evolution and the evolutionary consequences of human behaviour that go well beyond those that can be obtained from modern genomic data or the fossil and archaeological records alone.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Huxley, T. H. Evidence as to Man's Place in Nature (Williams and Norgate, 1863).
Darwin, C. The Descent of Man, and Selection in Relation to Sex (John Murray, 1871).
Fan, S., Hansen, M. E. B., Lo, Y. & Tishkoff, S. A. Going global by adapting local: A review of recent human adaptation. Science 354, 54–59 (2016).
Laland, K. N., Odling-Smee, J. & Myles, S. How culture shaped the human genome: bringing genetics and the human sciences together. Nat. Rev. Genet. 11, 137–148 (2010).
Scheinfeldt, L. B. & Tishkoff, S. A. Recent human adaptation: genomic approaches, interpretation and insights. Nat. Rev. Genet. 14, 692–702 (2013).
Sabeti, P. C. et al. Positive natural selection in the human lineage. Science 312, 1614–1620 (2006).
Raj, T. et al. Common risk alleles for inflammatory diseases are targets of recent positive selection. Am. J. Hum. Genet. 92, 517–529 (2013).
Papandreou, I., Cairns, R. A., Fontana, L., Lim, A. L. & Denko, N. C. HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metab. 3, 187–197 (2006).
Natarajan, V. T. et al. IFN-γ signaling maintains skin pigmentation homeostasis through regulation of melanosome maturation. Proc. Natl Acad. Sci. USA 111, 2301–2306 (2014).
Norton, H. L. et al. Genetic evidence for the convergent evolution of light skin in Europeans and East Asians. Mol. Biol. Evol. 24, 710–722 (2006).
Pickrell, J. K. & Reich, D. Toward a new history and geography of human genes informed by ancient DNA. Trends Genet. 30, 377–389 (2014).
Nielsen, R. et al. Tracing the peopling of the world through genomics. Nature 541, 302–310 (2017).
Orlando, L., Gilbert, M. T. P. & Willerslev, E. Reconstructing ancient genomes and epigenomes. Nat. Rev. Genet. 16, 395–408 (2015).
Llamas, B., Willerslev, E. & Orlando, L. Human evolution: a tale from ancient genomes. Phil. Trans. R. Soc. B 372, 20150484 (2016).
Nakagome, S. et al. Estimating the ages of selection signals from different epochs in human history. Mol. Biol. Evol. 33, 657–669 (2016).
Field, Y. et al. Detection of human adaptation during the past 2000 years. Science 354, 760–764 (2016).
Malaspinas, A.-S., Malaspinas, O., Evans, S. N. & Slatkin, M. Estimating allele age and selection coefficient from time-serial data. Genetics 192, 599–607 (2012).
Sams, A. J., Hawks, J. & Keinan, A. The utility of ancient human DNA for improving allele age estimates, with implications for demographic models and tests of natural selection. J. Hum. Evol. 79, 64–72 (2015).
Gamba, C. et al. Genome flux and stasis in a five millennium transect of European prehistory. Nat. Commun. 5, 5257 (2014).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015). This paper performed a genome-wide scan for signatures of positive selection with ancient genome data from 230 Europeans (!8,500–2,300 years BP ) and characterized the spatiotemporal frequency trajectories of adaptive alleles related to diet, skin pigmentation, stature and the immune response.
Gelabert, P., Olalde, I., de-Dios, T., Civit, S. & Lalueza-Fox, C. Malaria was a weak selective force in ancient Europeans. Sci. Rep. 7, 1377 (2017).
Buckley, M. T. et al. Selection in Europeans on fatty acid desaturases associated with dietary changes. Mol. Biol. Evol. 34, 1307–1318 (2017).
Ye, K., Gao, F., Wang, D., Bar-Yosef, O. & Keinan, A. Dietary adaptation of FADS genes in Europe varied across time and geography. Nat. Ecol. Evol. 1, 0167 (2017).
Sverrisdóttir, O. Ó. et al. Direct estimates of natural selection in Iberia indicate calcium absorption was not the only driver of lactase persistence in Europe. Mol. Biol. Evol. 31, 975–983 (2014).
Günther, T. et al. Genomics of Mesolithic Scandinavia reveal colonization routes and high-latitude adaptation. Preprint at bioRxiv http://dx.doi.org/10.1101/164400 (2017). This study conducted a genome-wide scan of positive selection using ancient genome data from seven Scandinavian individuals (!9,500–6,000 years BP ) to reveal a haplotype that the authors propose may underlie physiological adaptation to cold climate.
Stephan, W. Signatures of positive selection: from selective sweeps at individual loci to subtle allele frequency changes in polygenic adaptation. Mol. Ecol. 25, 79–88 (2016).
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Haak, W. et al. Massive migration from the steppe was a source for Indo–European languages in Europe. Nature 522, 207–211 (2015).
Olalde, I. et al. The Beaker phenomenon and the genomic transformation of northwest Europe. Preprint at bioRxiv http://dx.doi.org/10.1101/135962 (2017).
Cassidy, L. M. et al. Neolithic and Bronze Age migration to Ireland and establishment of the insular Atlantic genome. Proc. Natl Acad. Sci. USA 113, 368–373 (2016).
Shennan, S. Evolutionary demography and the population history of the European early Neolithic. Hum. Biol. 81, 339–355 (2009).
Leonardi, M. et al. Evolutionary patterns and processes: lessons from ancient DNA. Syst. Biol. 66, e1–e29 (2017).
Mirazón Lahr, M. The shaping of human diversity: filters, boundaries and transitions. Phil. Trans. R. Soc. B 371, http://dx.doi.org/10.1098/rstb.2015.0241 (2016).
Stewart, J. R. & Stringer, C. B. Human evolution out of Africa: the role of refugia and climate change. Science 335, 1317–1321 (2012).
Richards, M. A brief review of the archaeological evidence for Palaeolithic and Neolithic subsistence. Eur. J. Clin. Nutr. 56, 1262–1278 (2002).
Perry, G. H. Parasites and human evolution. Evol. Anthropol. 23, 218–228 (2014).
Pearce-Duvet, J. M. C. The origin of human pathogens: evaluating the role of agriculture and domestic animals in the evolution of human disease. Biol. Rev. 81, 369–382 (2006).
Jablonski, N. G. & Chaplin, G. Human skin pigmentation as an adaptation to UV radiation. Proc. Natl Acad. Sci. USA 107, 8962–8968 (2010).
Chaplin, G. & Jablonski, N. G. Vitamin D and the evolution of human depigmentation. Am. J. Phys. Anthropol. 139, 451–461 (2009).
Brickley, M. B. et al. Ancient vitamin D deficiency: long-term trends. Curr. Anthropol. 58, 420–427 (2017).
Ovesen, L., Brot, C. & Jakobsen, J. Food contents and biological activity of 25-hydroxyvitamin D: a vitamin D metabolite to be reckoned with? Ann. Nutr. Metab. 47, 107–113 (2003).
Bodnar, L. M. et al. Maternal vitamin D deficiency increases the risk of preeclampsia. J. Clin. Endocrinol. Metab. 92, 3517–3522 (2007).
Wang, T. J. et al. Common genetic determinants of vitamin D insufficiency: a genome-wide association study. Lancet 376, 180–188 (2010).
Lao, O., de Gruijter, J. M., van Duijn, K., Navarro, A. & Kayser, M. Signatures of positive selection in genes associated with human skin pigmentation as revealed from analyses of single nucleotide polymorphisms. Ann. Hum. Genet. 71, 354–369 (2007).
Sturm, R. A. et al. Human pigmentation genes under environmental selection. Genome Biol. 13, 248–263 (2012).
Beleza, S. et al. The timing of pigmentation lightening in Europeans. Mol. Biol. Evol. 30, 24–35 (2013).
Günther, T. et al. Ancient genomes link early farmers from Atapuerca in Spain to modern-day Basques. Proc. Natl Acad. Sci. USA 112, 11917–11922 (2015).
González-Fortes, G. et al. Paleogenomic evidence for multi-generational mixing between Neolithic farmers and Mesolithic hunter–gatherers in the Lower Danube Basin. Curr. Biol. 27, 1801–1810.e10 (2017).
Jones, E. R. et al. Upper Palaeolithic genomes reveal deep roots of modern Eurasians. Nat. Commun. 6, 8912 (2015).
Broushaki, F. et al. Early Neolithic genomes from the eastern Fertile Crescent. Science 353, 499–503 (2016).
Olalde, I. et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature 507, 225–228 (2014).
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
Wilde, S. et al. Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 y. Proc. Natl Acad. Sci. USA 111, 4832–4837 (2014). This study used ancient DNA data for alleles known to be involved in human pigmentation variation to identify a history of positive natural selection and estimate the strength of selection for each locus.
Jeong, C. et al. Long-term genetic stability and a high-altitude East Asian origin for the peoples of the high valleys of the Himalayan arc. Proc. Natl Acad. Sci. USA 113, 7485–7490 (2016). This is an ancient genome study of eight individuals from Nepal (3,150–1,250 years BP ) that found staggered appearances and frequency increases for several genetic variants known from studies of modern regional populations to be associated with physiological adaptation to high altitude.
Lorenzo, F. R. et al. A genetic mechanism for Tibetan high-altitude adaptation. Nat. Genet. 46, 951–956 (2014).
Peng, Y. et al. Genetic variations in Tibetan populations and high-altitude adaptation at the Himalayas. Mol. Biol. Evol. 28, 1075–1081 (2011).
Ralph, P. & Coop, G. Parallel adaptation: one or many waves of advance of an advantageous allele? Genetics 186, 647–668 (2010).
Novembre, J., Galvani, A. P. & Slatkin, M. The geographic spread of the CCR5 Δ32 HIV-resistance allele. PLoS Biol. 3, e339 (2005).
Richards, M. P., Schulting, R. J. & Hedges, R. E. M. Archaeology: sharp shift in diet at onset of Neolithic. Nature 425, 366–366 (2003).
Chaplin, G. & Jablonski, N. G. The human environment and the vitamin D compromise: Scotland as a case study in human biocultural adaptation and disease susceptibility. Hum. Biol. 85, 529–552 (2013).
Druzhkova, A. S. et al. Ancient DNA analysis affirms the canid from Altai as a primitive dog. PLoS ONE 8, e57754 (2013).
Snir, A. et al. The origin of cultivation and proto-weeds, long before Neolithic farming. PLoS ONE 10, e0131422 (2015).
Boivin, N. L. et al. Ecological consequences of human niche construction: examining long-term anthropogenic shaping of global species distributions. Proc. Natl Acad. Sci. USA 113, 6388–6396 (2016).
Copeland, L., Blazek, J., Salman, H. & Tang, M. C. Form and functionality of starch. Food Hydrocoll. 23, 1527–1534 (2009).
Gerbault, P. et al. Evolution of lactase persistence: an example of human niche construction. Phil. Trans. R. Soc. B. 366, 863–877 (2011).
Campbell, A. K., Waud, J. P. & Matthews, S. B. The molecular basis of lactose intolerance. Sci. Prog. 88, 157–202 (2005).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nat. Genet. 39, 31–40 (2007).
Itan, Y., Powell, A., Beaumont, M. A., Burger, J. & Thomas, M. G. The origins of lactase persistence in Europe. PLoS Comput. Biol. 5, e1000491 (2009).
Hofmanová, Z. et al. Early farmers from across Europe directly descended from Neolithic Aegeans. Proc. Natl Acad. Sci. USA 113, 6886–6891 (2016).
Burger, J., Kirchner, M., Bramanti, B., Haak, W. & Thomas, M. G. Absence of the lactase-persistence-associated allele in early Neolithic Europeans. Proc. Natl Acad. Sci. USA 104, 3736–3741 (2007). This was the first ancient DNA-based study of the history of the European lactase persistence allele; this study reported that the allele was not present in eight early Neolithic individuals (!7,500 years BP ) from four geographic sites, suggesting that the ability of individuals to digest lactose across their lifetimes likely post-dated the origin and spread of European dairying practices.
Craig, O. E. et al. Did the first farmers of central and eastern Europe produce dairy foods? Antiquity 79, 882–894 (2005).
Copley, M. S. et al. Direct chemical evidence for widespread dairying in prehistoric Britain. Proc. Natl Acad. Sci. USA 100, 1524–1529 (2003).
Treuil, R. Dikili Tash, village préhistorique de Macédoine orientale. 1, Fouilles de Jean Deshayes (1961–1975), vol. 2, Bulletin de Correspondance Hellénique Supplément 37 (Ecole française d'Athènes, 2004).
Salque, M. et al. Earliest evidence for cheese making in the sixth millennium BC in northern Europe. Nature 493, 522–525 (2013).
Perry, G. H. et al. Diet and the evolution of human amylase gene copy number variation. Nat. Genet. 39, 1256–1260 (2007).
Inchley, C. E. et al. Selective sweep on human amylase genes postdates the split with Neanderthals. Sci. Rep. 6, 37198 (2016).
Perry, G. H., Kistler, L., Kelaita, M. A. & Sams, A. J. Insights into hominin phenotypic and dietary evolution from ancient DNA sequence data. J. Hum. Evol. 79, 55–63 (2015).
Mathias, R. A. et al. Adaptive evolution of the FADS gene cluster within Africa. PLoS ONE 7, e44926 (2012).
Paul, B. D. & Snyder, S. H. The unusual amino acid l-ergothioneine is a physiologic cytoprotectant. Cell Death Differ. 17, 1134–1140 (2010).
Huff, C. D. et al. Crohn's disease and genetic hitchhiking at IBD5. Mol. Biol. Evol. 29, 101–111 (2012).
Lindo, J. et al. A time transect of exomes from a Native American population before and after European contact. Nat. Commun. 7, 13175 (2016). In this paper, the authors sequenced the exomes of 25 ancient First Nations individuals (!6,200–800 years BP ) from British Columbia, Canada, to identify an HLA-DQA1 gene haplotype with a substantial frequency difference compared with the Tsimshian descendant population living in the region today, potentially reflecting adaptation to disease outbreaks associated with European colonization (which post-dated the ancient DNA time series).
Harkins, K. M. & Stone, A. C. Ancient pathogen genomics: insights into timing and adaptation. J. Hum. Evol. 79, 137–149 (2015).
Bos, K. I. et al. Pre-Columbian mycobacterial genomes reveal seals as a source of New World human tuberculosis. Nature 514, 494–497 (2014).
Roffey, S. et al. Investigation of a medieval pilgrim burial excavated from the leprosarium of St Mary Magdalen Winchester. PLoS Negl. Trop. Dis. 11, e0005186 (2017).
Spyrou, M. A. et al. Historical Y. pestis genomes reveal the European Black Death as the source of ancient and modern plague pandemics. Cell Host Microbe 19, 874–881 (2016).
Marciniak, S. et al. Plasmodium falciparum malaria in 1 st−2nd century CE southern Italy. Curr. Biol. 26, R1220–R1222 (2016).
Gelabert, P. et al. Mitochondrial DNA from the eradicated European Plasmodium vivax and P. falciparum from 70-year-old slides from the Ebro Delta in Spain. Proc. Natl Acad. Sci. USA 113, 11495–11500 (2016).
Duggan, A. T. et al. 17 th century Variola virus reveals the recent history of smallpox. Curr. Biol. 26, 3407–3412 (2016).
Devault, A. M. et al. Second-pandemic strain of Vibrio cholerae from the Philadelphia cholera outbreak of 1849. N. Engl. J. Med. 370, 334–340 (2014).
Devault, A. M. et al. A molecular portrait of maternal sepsis from Byzantine Troy. eLife 6, e20983 (2017).
Warinner, C. et al. Pathogens and host immunity in the ancient human oral cavity. Nat. Genet. 46, 336–344 (2014). This study reported ancient DNA results from dental calculus collected from four European individuals (!1,000–750 years BP ), including documentation of the presence of various pathogenic bacteria and producing direct evidence that pig, sheep, wheat and cruciferous vegetables were consumed as part of the diet.
Rasmussen, S. et al. Early divergent strains of Yersinia pestis in Eurasia 5,000 years ago. Cell 163, 571–582 (2015). This paper sequenced seven Eurasian plague ( Yersinia pestis ) ancient genomes (!5,000–2,800 years BP ) and, among other findings, discovered that a gene encoding a protein necessary for Y. pestis viability in the flea gut was absent from genomes prior to !3,600 years BP ; the subsequent acquisition of this gene through horizontal transfer likely helped facilitate the bubonic plague transmission cycle.
Hinnebusch, B. J. et al. Role of Yersinia murine toxin in survival of Yersinia pestis in the midgut of the flea vector. Science 296, 733–735 (2002).
Bos, K. I. et al. A draft genome of Yersinia pestis from victims of the Black Death. Nature 478, 506–510 (2011).
Hummel, S., Schmidt, D., Kremeyer, B., Herrmann, B. & Oppermann, M. Detection of the CCR5-Δ32 HIV resistance gene in Bronze Age skeletons. Genes Immun. 6, 371–374 (2005).
Wolpoff, M. H., Thorne, A. G., Smith, F. H., Frayer, D. W. & Pope, G. G. in Origins of Anatomically Modern Humans (eds Nitecki, M. H. & Nitecki, D.) V.) 175–199 (Plenum Press, 1994).
Tattersall, I. Out of Africa: modern human origins special feature: human origins: out of Africa. Proc. Natl Acad. Sci. USA 106, 16018–16021 (2009).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010).
Vernot, B. & Akey, J. M. Resurrecting surviving Neandertal lineages from modern human genomes. Science 343, 1017–1021 (2014).
Dannemann, M., Prüfer, K. & Kelso, J. Functional implications of Neandertal introgression in modern humans. Genome Biol. 18, 61–72 (2017).
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014).
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Huerta-Sánchez, E. et al. Altitude adaptation in Tibet caused by introgression of Denisovan-like DNA. Nature 512, 194–197 (2014). This paper demonstrated that a genetic haplotype surrounding the EPAS1 gene that underlies a physiological adaptation to high altitude in modern Tibetans is the result of adaptive introgression from Denisovans or a related archaic hominin population.
Racimo, F. et al. Archaic adaptive introgression in TBX15/WARS2. Mol. Biol. Evol. 34, 509–524 (2017).
Skoglund, P. & Jakobsson, M. Archaic human ancestry in East Asia. Proc. Natl Acad. Sci. USA 108, 18301–18306 (2011).
Reich, D. et al. Denisova admixture and the first modern human dispersals into Southeast Asia and Oceania. Am. J. Hum. Genet. 89, 516–528 (2011).
Castellano, S. et al. Patterns of coding variation in the complete exomes of three Neandertals. Proc. Natl Acad. Sci. USA 111, 6666–6671 (2014).
Lalueza-Fox, C. et al. A melanocortin 1 receptor allele suggests varying pigmentation among Neanderthals. Science 318, 1453–1455 (2007).
Lalueza-Fox, C., Gigli, E., de la Rasilla, M., Fortea, J. & Rosas, A. Bitter taste perception in Neanderthals through the analysis of the TAS2R38 gene. Biol. Lett. 5, 809–811 (2009).
McCoy, R. C., Wakefield, J. & Akey, J. M. Impacts of Neanderthal-introgressed sequences on the landscape of human gene expression. Cell 168, 916–927 (2017).
Simonti, C. N. et al. The phenotypic legacy of admixture between modern humans and Neandertals. Science 351, 737–741 (2016). This study used electronic health record phenotypes from a large sample of modern human patients to associate alleles originally introgressed from Neandertals with increased risk of depression, skin lesions associated with sun exposure (actinic keratosis), and other phenotypes.
Gittelman, R. M. et al. Archaic hominin admixture facilitated adaptation to Out-of-Africa environments. Curr. Biol. 26, 3375–3382 (2016). This paper analysed 126 genomic regions containing strong signatures of adaptive introgression from archaic hominins identified in a sample of geographically diverse human populations; these loci are significantly enriched for genes involved in the immune response and also contain multiple genes with known roles in skin pigmentation.
Racimo, F., Sankararaman, S., Nielsen, R. & Huerta-Sánchez, E. Evidence for archaic adaptive introgression in humans. Nat. Rev. Genet. 16, 359–371 (2015).
Sankararaman, S. et al. The landscape of Neandertal ancestry in present-day humans. Nature 507, 354–357 (2014).
Vernot, B. et al. Excavating Neandertal and Denisovan DNA from the genomes of Melanesian individuals. Science 352, 235–239 (2016).
Fumagalli, M. et al. Greenlandic Inuit show genetic signatures of diet and climate adaptation. Science 349, 1343–1347 (2015).
Gburcik, V., Cawthorn, W. P., Nedergaard, J., Timmons, J. A. & Cannon, B. An essential role for Tbx15 in the differentiation of brown and 'brite' but not white adipocytes. Am. J. Physiol. Endocrinol. Metab. 303, E1053–E1060 (2012).
Deschamps, M. et al. Genomic signatures of selective pressures and introgression from archaic hominins at human innate immunity genes. Am. J. Hum. Genet. 98, 5–21 (2016).
Enard, D. & Petrov, D. A. RNA viruses drove adaptive introgressions between Neanderthals and modern humans. Preprint at bioRxivhttp://dx.doi.org/10.1101/120477 (2017).
Abi-Rached, L. et al. The shaping of modern human immune systems by multiregional admixture with archaic humans. Science 334, 89–94 (2011).
Sams, A. J. et al. Adaptively introgressed Neandertal haplotype at the OAS locus functionally impacts innate immune responses in humans. Genome Biol. 17, 246–261 (2016).
Sullivan, A. P., de Manuel, M., Marques-Bonet, T. & Perry, G. H. An evolutionary medicine perspective on Neandertal extinction. J. Hum. Evol. 108, 62–71 (2017).
Houldcroft, C. J. & Underdown, S. J. Neanderthal genomics suggests a pleistocene time frame for the first epidemiologic transition. Am. J. Phys. Anthropol. 160, 379–388 (2016).
Key, F. M., Teixeira, J. C., de Filippo, C. & Andrés, A. M. Advantageous diversity maintained by balancing selection in humans. Curr. Opin. Genet. Dev. 29, 45–51 (2014).
Nédélec, Y. et al. Genetic ancestry and natural selection drive population differences in immune responses to pathogens. Cell 167, 657–669 (2016).
Quach, H. et al. Genetic adaptation and Neandertal admixture shaped the immune system of human populations. Cell 167, 643–656.e17 (2016).
Park, S. D. E. et al. Genome sequencing of the extinct Eurasian wild aurochs, Bos primigenius, illuminates the phylogeography and evolution of cattle. Genome Biol. 16, 234–249 (2015).
Loog, L. et al. Inferring allele frequency trajectories from ancient DNA indicates that selection on a chicken gene coincided with changes in medieval husbandry practices. Mol. Biol. Evol. http://dx.doi.org/10.1093/molbev/msx142 (2017). This study of domestic chickens connected the ancient DNA-informed timing of a significant change in frequency for an allele associated with increased egg production to concomitant increases in the intensity of chicken husbandry as documented by historical and archaeological records.
MacHugh, D. E., Larson, G. & Orlando, L. Taming the past: ancient DNA and the study of animal domestication. Annu. Rev. Anim. Biosci. 5, 329–351 (2017).
Ramos-Madrigal, J. et al. Genome sequence of a 5,310-year-old maize cob provides insights into the early stages of maize domestication. Curr. Biol. 26, 3195–3201 (2016). This study identified a mix of ancestral and derived functional genetic variants in maize from Mexico !5,300 years BP , highlighting the gradual temporal process of trait evolution in this domestic species.
Schubert, M. et al. Prehistoric genomes reveal the genetic foundation and cost of horse domestication. Proc. Natl Acad. Sci. USA 111, E5661–E5669 (2014).
Ollivier, M. et al. Amy2B copy number variation reveals starch diet adaptations in ancient European dogs. R. Soc. Open Sci. 3, 160449 (2016).
Librado, P. et al. Tracking the origins of Yakutian horses and the genetic basis for their fast adaptation to subarctic environments. Proc. Natl Acad. Sci. USA 112, E6889–E6897 (2015).
Jaenicke-Després, V. et al. Early allelic selection in maize as revealed by ancient DNA. Science 302, 1206–1208 (2003).
Flink, L. G. et al. Establishing the validity of domestication genes using DNA from ancient chickens. Proc. Natl Acad. Sci. USA 111, 6184–6189 (2013).
West, B. & Zhou, B.-X. Did chickens go North? New evidence for domestication. J. Archaeol. Sci. 15, 515–533 (1988).
Librado, P. et al. Ancient genomic changes associated with domestication of the horse. Science 356, 442–445 (2017).
Outram, A. K. et al. The earliest horse harnessing and milking. Science 323, 1332–1335 (2009).
Ludwig, A. et al. Coat colour variation at the beginning of horse domestication. Science 324, 485 (2009).
Ludwig, A. et al. Twenty-five thousand years of fluctuating selection on leopard complex spotting and congenital night blindness in horses. Phil. Trans. R. Soc. B. 370, 20130386 (2014).
Ottoni, C. et al. The palaeogenetics of cat dispersal in the ancient world. Nat. Ecol. Evol. 1, 0139 (2017).
Axelsson, E. et al. The genomic signature of dog domestication reveals adaptation to a starch-rich diet. Nature 495, 360–364 (2013).
Arendt, M., Cairns, K. M., Ballard, J. W. O., Savolainen, P. & Axelsson, E. Diet adaptation in dog reflects spread of prehistoric agriculture. Heredity 117, 301–306 (2016).
Botigué, L. R. et al. Ancient European dog genomes reveal continuity since the Early Neolithic. Nat. Commun. 8, 16082 (2017).
Frantz, L. A. F. et al. Genomic and archaeological evidence suggest a dual origin of domestic dogs. Science 352, 1228–1231 (2016).
Kistler, L., Ware, R., Smith, O., Collins, M. & Allaby, R. G. A new model for ancient DNA decay based on paleogenomic meta-analysis. Nucleic Acids Res. 45, 6310–6320 (2017).
Fehren-Schmitz, L. & Georges, L. Ancient DNA reveals selection acting on genes associated with hypoxia response in pre-Columbian Peruvian Highlanders in the last 8500 years. Sci. Rep. 6, 23485 (2016).
Sullivan, A. P., Bird, D. W. & Perry, G. H. Human behaviour as a long-term ecological driver of non-human evolution. Nat. Ecol. Evol. 1, 0065 (2017).
Noonan, J. P. et al. Sequencing and analysis of Neanderthal genomic DNA. Science 314, 1113–1118 (2006).
Green, R. E. et al. Analysis of one million base pairs of Neanderthal DNA. Nature 444, 330–336 (2006).
Ramírez, O. et al. Paleogenomics in a temperate environment: shotgun sequencing from an extinct Mediterranean Caprine. PLoS ONE 4, e5670 (2009).
Miller, W. et al. Sequencing the nuclear genome of the extinct woolly mammoth. Nature 456, 387–390 (2008).
Lambert, D. M. & Millar, C. D. Evolutionary biology: ancient genomics is born. Nature 444, 275–276 (2006).
Wall, J. D. & Kim, S. K. Inconsistencies in Neanderthal genomic DNA sequences. PLoS Genet. 3, 1862–1866 (2007).
Rasmussen, M. et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463, 757–762 (2010).
Lazaridis, I. et al. Genomic insights into the origin of farming in the ancient Near East. Nature 536, 419–424 (2016).
Pinhasi, R. et al. Optimal ancient DNA yields from the inner ear part of the human petrous bone. PLoS ONE 10, e0129102 (2015).
Gansauge, M.-T. & Meyer, M. Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat. Protoc. 8, 737–748 (2013).
Carpenter, M. L. et al. Pulling out the 1%: whole-genome capture for the targeted enrichment of ancient DNA sequencing libraries. Am. J. Hum. Genet. 93, 852–864 (2013).
Hofreiter, M. et al. The future of ancient DNA: technical advances and conceptual shifts. BioEssays 37, 284–293 (2015).
Gron, K. J. et al. Cattle management for dairying in Scandinavia's earliest Neolithic. PLoS ONE 10, e0131267 (2015).
Zeder, M. A. & Hesse, B. The initial domestication of goats (Capra hircus) in the Zagros mountains 10,000 years ago. Science 287, 2254–2257 (2000).
Cramp, L. J. E. et al. Neolithic dairy farming at the extreme of agriculture in northern Europe. Proc. Biol. Sci. 281, 20140819 (2014).
Warinner, C. et al. Direct evidence of milk consumption from ancient human dental calculus. Sci. Rep. 4, 7104 (2015).
Yang, Y. et al. Proteomics evidence for kefir dairy in Early Bronze Age China. J. Archaeol. Sci. 45, 178–186 (2014).
Burbano, H. A. et al. Targeted investigation of the Neandertal genome by array-based sequence capture. Science 328, 723–725 (2010).
Rasmussen, M. et al. An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334, 94–98 (2011).
Keller, A. et al. New insights into the Tyrolean Iceman's origin and phenotype as inferred by whole-genome sequencing. Nat. Commun. 3, 698 (2012).
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012).
Fu, Q. et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl Acad. Sci. USA 110, 2223–2227 (2013).
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014).
Schroeder, H. et al. Genome-wide ancestry of 17 th-century enslaved Africans from the Caribbean. Proc. Natl Acad. Sci. USA 112, 3669–3673 (2015).
Malaspinas, A.-S. et al. Two ancient human genomes reveal Polynesian ancestry among the indigenous Botocudos of Brazil. Curr. Biol. 24, R1035–R1037 (2014).
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014).
Rasmussen, M. et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225–229 (2014).
Seguin-Orlando, A. et al. Genomic structure in Europeans dating back at least 36,200 years. Science 346, 1113–1118 (2014).
Skoglund, P. et al. Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and farmers. Science 344, 747–750 (2014).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Gallego Llorente, M. et al. Ancient Ethiopian genome reveals extensive Eurasian admixture in Eastern Africa. Science 350, 820–822 (2015).
Olalde, I. et al. A common genetic origin for early farmers from Mediterranean Cardial and Central European LBK cultures. Mol. Biol. Evol. 32, 3132–3142 (2015).
Raghavan, M. et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science 349, aab3884 (2015).
Sawyer, S. et al. Nuclear and mitochondrial DNA sequences from two Denisovan individuals. Proc. Natl Acad. Sci. USA 112, 15696–15700 (2015).
Rasmussen, M. et al. The ancestry and affiliations of Kennewick Man. Nature 523, 455–458 (2015).
Martiniano, R. et al. Genomic signals of migration and continuity in Britain before the Anglo-Saxons. Nat. Commun. 7, 10326 (2016).
Schiffels, S. et al. Iron Age and Anglo-Saxon genomes from East England reveal British migration history. Nat. Commun. 7, 10408 (2016).
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016).
Gallego Llorente, M. et al. The genetics of an early Neolithic pastoralist from the Zagros, Iran. Sci. Rep. 6, 31326 (2016).
Kılınç, G. M. et al. The demographic development of the first farmers in Anatolia. Curr. Biol. 26, 2659–2666 (2016).
Omrak, A. et al. Genomic evidence establishes Anatolia as the source of the European Neolithic gene pool. Curr. Biol. 26, 270–275 (2016).
Meyer, M. et al. Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature 531, 504–507 (2016).
Skoglund, P. et al. Genomic insights into the peopling of the Southwest Pacific. Nature 538, 510–513 (2016).
Jones, E. R. et al. The Neolithic transition in the Baltic was not driven by admixture with early European farmers. Curr. Biol. 27, 576–582 (2017).
Unterländer, M. et al. Ancestry and demography and descendants of Iron Age nomads of the Eurasian Steppe. Nat. Commun. 8, 14615 (2017).
Lipson, M. et al. Parallel ancient genomic transects reveal complex population history of early European farmers. Preprint at bioRxivhttp://dx.doi.org/10.1101/114488 (2017).
Lindo, J. et al. Ancient individuals from the North American Northwest Coast reveal 10,000 years of regional genetic continuity. Proc. Natl Acad. Sci. USA 114, 4093–4098 (2017).
Kennett, D. J. et al. Archaeogenomic evidence reveals prehistoric matrilineal dynasty. Nat. Commun. 8, 14115 (2017).
Mathieson, I. et al. The genomic history of southeastern Europe. Preprint at bioRxivhttp://dx.doi.org/10.1101/135616 (2017).
Martiniano, R. et al. The population genomics of archaeological transition in west Iberia: Investigation of ancient substructure using imputation and haplotype-based methods. PLoS Genet. 13, e1006852 (2017).
Siska, V. et al. Genome-wide data from two early Neolithic East Asian individuals dating to 7700 years ago. Sci. Adv. 3, e1601877 (2017).
Haber, M. et al. Continuity and admixture in the last five millennia of Levantine history from ancient Canaanite and present-day Lebanese genome sequences. Am. J. Hum. Genet. http://dx.doi.org/10.1016/j.ajhg.2017.06.013 (2017).
Schuenemann, V. J. et al. Ancient Egyptian mummy genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods. Nat. Commun. 8, 15694 (2017).
Schlebusch, C. M. et al. Ancient genomes from southern Africa pushes modern human divergence beyond 260,000 years ago. Preprint at bioRxivhttp://dx.doi.org/10.1101/145409 (2017).
Mittnik, A. et al. The genetic history of northern Europe. Preprint at bioRxivhttp://dx.doi.org/10.1101/113241 (2017).
Slon, V. et al. A fourth Denisovan individual. Sci. Adv. 3, e1700186 (2017).
Kuhlwilm, M. et al. Ancient gene flow from early modern humans into Eastern Neanderthals. Nature 530, 429–433 (2016).
Acknowledgements
The authors thank C. Bergey and R. George for discussion about the manuscript. This work was supported by grants from the National Science Foundation (BCS-1554834 and BCS-1317163; to G.H.P.).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (table)
Spatiotemporal frequencies of the European lactase persistence allele. (PDF 186 kb)
Glossary
- Adaptation
-
A process of phenotypic and corresponding genetic change over time for traits that confer increased reproductive fitness in a given environmental context.
- Positive natural selection
-
A mechanism of evolution in which a genetically mediated trait that confers a relative fitness advantage increases in frequency over time because of that advantage. In this Review, we refer to positive selection as an adaptive process that can act on new or previously existing genetic variants.
- Phenotype
-
Physical traits of an organism; often refers to externally visible traits but may include internal and microscopic or biochemical traits.
- Ancient DNA
-
DNA from palaeontological, archaeological, or historical but pre-modern biological specimens that is often damaged and degraded and recovered in small quantities.
- Exome
-
All or nearly all protein-coding gene regions of the nuclear genome; in humans, representing approximately 1% of the genome.
- Single-nucleotide polymorphism
-
(SNP). A single position in the reference genome at which the specific nucleotide present (thymine, guanine, cytosine, or adenine) varies among individuals in a population or species.
- Archaic hominins
-
Now-extinct populations or species that are distinct from anatomically modern humans but that share a more recent common ancestor with modern humans than with chimpanzees — for example, Neandertals and Denisovans.
- Anatomically modern humans
-
Hominins recognizable phenotypically as early members of our own species, Homo sapiens, first appearing >200,000 years BP in Africa.
- Adaptive introgression
-
The process of a genetic variant that was originally introduced into a population via admixture increasing in frequency by positive natural selection because it confers a fitness advantage.
- Genetic drift
-
Changes in genetic variation over time that are due to random (chance) processes, apart from natural selection.
- Gene flow
-
Movement of genetic variation between populations, for example, through migration or admixture.
- Neolithic
-
A cultural period in human prehistory characterized by early technological and demographic shifts associated with the transition to farming and pastoralism, occurring at different times across regions.
- Domestication
-
A process of plant and animal evolution mediated by human selection for particular phenotypes (artificial selection), sometimes combined with commensal adaptation to human-constructed niches.
- Biocultural adaptation
-
The process of interaction between human cultural and adaptive biological change (for example, dairying and the ability of adults to digest milk sugars).
- Zoonotic
-
The ability of a pathogen to be directly or indirectly transmitted to humans from animals sharing the same habitat.
- Admixture
-
Interbreeding between previously isolated populations.
Rights and permissions
About this article
Cite this article
Marciniak, S., Perry, G. Harnessing ancient genomes to study the history of human adaptation. Nat Rev Genet 18, 659–674 (2017). https://doi.org/10.1038/nrg.2017.65
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2017.65
This article is cited by
-
Assessing the impact of post-mortem damage and contamination on imputation performance in ancient DNA
Scientific Reports (2024)
-
Ancient dolphin genomes reveal rapid repeated adaptation to coastal waters
Nature Communications (2023)
-
Climate Change Predictive of Body Size and Proportionality in Humans
Evolutionary Biology (2023)
-
Performance of innovative nanomaterials for bone remains consolidation and effect on 14C dating and on palaeogenetic analysis
Scientific Reports (2022)
-
Genetic load: genomic estimates and applications in non-model animals
Nature Reviews Genetics (2022)