Key Points
-
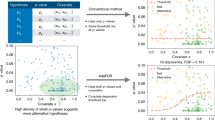
Gene-set analysis of GWAS data can best be understood as an analysis using genes as data points, carrying out a test of the relationship between a gene set and the genetic associations of genes with a phenotype.
-
Self-contained gene-set analysis does not provide information about the gene set itself, but only about the genes it contains. As such it cannot be used to draw biologically meaningful conclusions and is therefore unsuitable for the research questions gene-set analysis is generally used to address.
-
Competitive gene-set analysis can be biologically informative but is vulnerable to various forms of confounding and biases as a result of linkage disequilibrium. Of evaluated competitive gene-set analysis methods, only INRICH and MAGMA show consistently good statistical performance.
-
Statistical power for competitive gene-set analysis is strongly dependent on the heritability of a phenotype. Gene-set effect sizes for more strongly heritable phenotypes must be higher to achieve the same level of power as less strongly heritable phenotypes.
-
As a result of the structure of competitive gene-set analysis, increasing sample size will only improve its statistical power to a limited extent, especially for more strongly heritable phenotypes.
-
Gene-set analysis applied to GWAS data and gene expression data show the same kind of statistical behaviour. Both are instances of a broader framework of gene-level analysis, which also includes other approaches such as gene-network analysis.
Abstract
The rapid increase in loci discovered in genome-wide association studies has created a need to understand the biological implications of these results. Gene-set analysis provides a means of gaining such understanding, but the statistical properties of gene-set analysis are not well understood, which compromises our ability to interpret its results. In this Analysis article, we provide an extensive statistical evaluation of the core structure that is inherent to all gene- set analyses and we examine current implementations in available tools. We show which factors affect valid and successful detection of gene sets and which provide a solid foundation for performing and interpreting gene-set analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Visscher, P. M., Brown, M. A., McCarthy, M. I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Wang, K., Li, M. & Hakonarson, H. Analysing biological pathways in genome-wide association studies. Nat. Rev. Genet. 11, 843–854 (2010). This review provides a comprehensive overview of the basic concepts and issues relevant to gene-set analysis.
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Sullivan, P. F. & Posthuma, D. Biological pathways and networks implicated in psychiatric disorders. Curr. Opin. Behav. Sci. 2, 58–68 (2015).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Nurnberger, J. I. et al. Identification of pathways for bipolar disorder: a meta-analysis. JAMA Psychiatry 71, 657–664 (2014).
Askland, K., Read, C. & Moore, J. Pathway-based analyses of whole-genome association study data in bipolar disorder reveal genes mediating ion channel activity and synaptic neurotransmission. Hum. Genet. 125, 63–79 (2009).
Wang, K. et al. Diverse genome-wide association studies associate the IL12/IL23 pathway with Crohn disease. Am. J. Hum. Genet. 84, 399–405 (2009).
Eleftherohorinou, H., Hoggart, C. J., Wright, V. J., Levin, M. & Coin, L. J. Pathway-driven gene stability selection of two rheumatoid arthritis GWAS identifies and validates new susceptibility genes in receptor mediated signalling pathways. Hum. Mol. Genet. 20, 3494–3506 (2011).
Menashe, I. et al. Pathway analysis of breast cancer genome-wide association study highlights three pathways and one canonical signaling cascade. Cancer Res. 70, 4453–4459 (2010).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Sullivan, P. F., Daly, M. J. & O'Donovan, M. Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat. Rev. Genet. 13, 537–551 (2012).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Manolio, T. A. Genomewide association studies and assessment of the risk of disease. N. Engl. J. Med. 363, 166–176 (2010).
Manolio, T. A. Bringing genome-wide association findings into clinical use. Nat. Rev. Genet. 14, 549–558 (2013).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Qian, D. C. et al. Identification of shared and unique susceptibility pathways among cancers of the lung, breast, and prostate from genome-wide association studies and tissue-specific protein interactions. Hum. Mol. Genet. 24, 7406–7420 (2015).
Wang, L., Jia, P., Wolfinger, R. D., Chen, X. & Zhao, Z. Gene set analysis of genome-wide association studies: methodological issues and perspectives. Genomics 98, 1–8 (2011).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Croft, D., Mundo, A. F., Haw, R., Milacic, M. & Weiser, J. The Reactome pathway knowledgebase. Nucleic Acids Res. 42, D472–D477 (2014). This is a clear practical guide to carrying out a gene-set analysis.
Mooney, M. A., Nigg, J. T., McWeeney, S. K. & Wilmot, B. Functional and genomic context in pathway analysis of GWAS data. Trends Genet. 30, 390–400 (2014).
Holmans, P. Statistical methods for pathway analysis of genome-wide data for association with complex genetic traits. Adv. Genet. 72, 141–179 (2010). This paper provides an extensive theoretical overview of different types of methods and sources of gene sets.
Evangelou, M., Rendon, A., Ouwehand, W. H., Wernisch, L. & Dudbridge, F. Comparison of methods for competitive tests of pathway analysis. PLoS ONE 7, e41018 (2012).
Gui, H., Li, M., Sham, P. C. & Cherny, S. S. Comparison of seven algorithms for pathway analysis using the WTCCC Crohn's Disease dataset. BMC Res. Notes 4, 386 (2011).
Elbers, C. C. et al. Using genome-wide pathway analysis to unravel the etiology of complex diseases. Genet. Epidemiol. 33, 419–431 (2009).
Tintle, N. L., Borchers, B., Brown, M. & Bekmetjev, A. Comparing gene set analysis methods on single-nucleotide polymorphism data from Genetic Analysis Workshop 16. BMC Proc. 3, S96 (2009).
Ballard, D., Abraham, C., Cho, J. & Zhao, H. Pathway analysis comparison using Crohn's disease genome wide association studies. BMC Med. Genom. 3, 25 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Moskvina, V. et al. Evaluation of an approximation method for assessment of overall significance of multiple-dependent tests in a genome-wide association study. Genet. Epidemiol. 35, 861–866 (2011).
O'Dushlaine, C. et al. The SNP ratio test: pathway analysis of genome-wide association datasets. Bioinformatics 25, 2762–2763 (2009).
Lips, E. S. et al. Functional gene group analysis identifies synaptic gene groups as risk factor for schizophrenia. Mol. Psychiatry 20, 22–34 (2011).
Li, M., Kwan, J. S. H. & Sham, P. C. HYST: a hybrid set-based test for genome-wide association studies, with application to protein-protein interaction-based association analysis. Am. J. Hum. Genet. 91, 478–488 (2012).
Pedroso, I. et al. Common genetic variants and gene-expression changes associated with bipolar disorder are over-represented in brain signaling pathway genes. Biol. Psychiatry 72, 311–317 (2012).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Wang, K., Li, M. & Bucan, M. Pathway-based approaches for analysis of genome-wide association studies. Am. J. Hum. Genet. 81, 1278–1283 (2007).
Holmans, P. et al. Gene ontology analysis of GWA study data sets provides insights into the biology of bipolar disorder. Am. J. Hum. Genet. 85, 13–24 (2009).
Lee, P. H., O'Dushlaine, C., Thomas, B. & Purcell, S. M. INRICH: Interval-based enrichment analysis for genome-wide association studies. Bioinformatics 28, 1797–1799 (2012).
Segrè, A. V. et al. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Bhatti, G. KEGGandMetacoreDzPathwaysGEO: Disease Datasets from GEO. R package version 0.104.0 (2014).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Jia, P. & Zhao, Z. Network-assisted analysis to prioritize GWAS results: princples, methods and perspectives. Hum. Genet. 133, 125–138 (2014).
Mitrea, C. et al. Methods and approaches in the topology-based analysis of biological pathways. Front. Physiol. 4, 278 (2013).
Acknowledgements
This study makes use of data generated by the Wellcome Trust Case-Control Consortium. A full list of the investigators who contributed to the generation of the data is available from www.wtccc.org.uk. Funding for the project was provided by the Wellcome Trust under award 076113 and 085475. This work was funded by The Netherlands Organization for Scientific Research (NWO VICI 453-14-005, 645-000-003).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (methods)
The statistical properties of gene-set analysis (PDF 1267 kb)
Supplementary information S2 (figure)
Mean significance rates for different types of method at sample size of about 100,000. (PDF 171 kb)
Supplementary information S3 (figure)
Mean significance rates for self-contained analysis at different levels of polygenicity. (PDF 133 kb)
Supplementary information S4 (figure)
Gene association scores as a function of gene size. (PDF 177 kb)
Supplementary information S5 (figure)
Accounting for gene density in competitive analysis. (PDF 128 kb)
Supplementary information S6 (figure)
Additional simulation results for self-contained and competitive analysis tools. (PDF 196 kb)
Supplementary information S7 (figure)
Results for competitive analysis tools at different settings. (PDF 222 kb)
Supplementary information S8 (figure)
Type 1 error rates for MAGMA and INRICH at lower significance thresholds for analysis of Reactome gene sets. (PDF 154 kb)
Supplementary information S9 (figure)
Result for MHC simulation for MAGMA and INRICH. (PDF 155 kb)
Supplementary information S10 (figure)
Effect of SNP effect distribution on power. (PDF 149 kb)
Supplementary information S11 (figure)
Effect of gene effect distribution on power. (PDF 175 kb)
Supplementary information S12 (figure)
QQ-plots for genes in associated gene sets. (PDF 181 kb)
Supplementary information S13 (figure)
Comparison of power for different types of competitive analysis method. (PDF 325 kb)
Supplementary information S14 (figure)
Comparison of power between self-contained and competitive analysis at 0% background heritability. (PDF 135 kb)
Supplementary information S15 (figure)
Comparison in power for competitive analysis with MAGMA and INRICH. (PDF 312 kb)
Glossary
- Gene sets
-
Groups of genes that share a particular property, typically their involvement in a particular biological process.
- Gene-association scores
-
Measures of the strength of the association, or the evidence for that association, between a gene and a phenotype of interest.
- Self-contained GSA
-
A type of gene-set analysis (GSA) that tests the null hypothesis that none of the genes in the gene set are associated with the phenotype.
- Linkage disequilibrium
-
The presence of statistical associations between alleles at different loci.
- Competitive GSA
-
A type of gene-set analysis (GSA) that tests the null hypothesis that the genes in the gene set are no more strongly associated with the phenotype than other genes.
- Genetic architecture
-
The pattern of genetic variants underlying a phenotype, including the number of variants, the allele frequencies, the effect sizes and the nature of effects (for example, additive or non-additive).
- Heritability
-
The proportion of the phenotypic variance that can be attributed to genetic differences among individuals.
- Phenotype permutation
-
A permutation scheme in which phenotypes are permuted, implying that phenotypes are independent of the genotypes.
- Gene permutation
-
A permutation scheme in which genes are permuted, implying that gene-association scores are independent of the target gene set. This is equivalent to randomly drawing gene sets of the same size.
- Polygenic phenotypes
-
Phenotypes influenced by large numbers of genetic variants, each with individually small effects.
- Biological confounding
-
Confounding that occurs in gene-set analysis (GSA) when the biological process according to which a gene set is defined has no causal role in the phenotype but contains many genes also involved with another biological process that does.
- Methodological confounding
-
Confounding that occurs in gene-set analysis (GSA) as a result of choices in data collection or computation of the gene-association scores, which creates a gene-set association that biologically does not exist.
- Population stratification
-
The presence of systematic differences in allele frequencies in subpopulations of the sample, possibly as a result of different ancestry. This can lead to inflation of gene-association scores when correlated to the phenotype, potentially resulting in spurious gene-set associations.
Rights and permissions
About this article
Cite this article
de Leeuw, C., Neale, B., Heskes, T. et al. The statistical properties of gene-set analysis. Nat Rev Genet 17, 353–364 (2016). https://doi.org/10.1038/nrg.2016.29
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2016.29
This article is cited by
-
Neuroimaging and multiomics reveal cross-scale circuit abnormalities in schizophrenia
Nature Mental Health (2023)
-
Integrative pathway and network analysis provide insights on flooding-tolerance genes in soybean
Scientific Reports (2023)
-
Excitatory/inhibitory imbalance in autism: the role of glutamate and GABA gene-sets in symptoms and cortical brain structure
Translational Psychiatry (2023)
-
New insights into the genetic etiology of Alzheimer’s disease and related dementias
Nature Genetics (2022)
-
Mapping gene and gene pathways associated with coronary artery disease: a CARDIoGRAM exome and multi-ancestry UK biobank analysis
Scientific Reports (2021)