Key Points
-
The human transcriptome is the product of a coordinated series of transcriptional, co-transcriptional and post-transcriptional regulatory events.
-
RNA splicing is a key regulatory step in gene expression that allows a limited genome to express an impressive diversity of coding and non-coding RNAs.
-
RNA mis-splicing causes a large array of human diseases due to hereditary and somatic mutations.
-
Mis-splicing may result from mutations to RNA cis-regulatory elements, core spliceosomal components or trans-acting regulatory factors.
-
Mutations in some genes, such as lamin A (LMNA), cause multiple types of diseases, from muscular dystrophy to premature ageing syndromes.
-
A key small nuclear RNA (snRNA) component of the minor spliceosome functions as a stress-activated switch to control expression levels of genes containing minor introns.
-
Some splicing factors linked to diseases, such as amyotrophic lateral sclerosis, contain low-complexity regions with prion-like domains that are susceptible to abnormal aggregation.
-
Splicing modulatory therapeutic strategies have been developed that target a range of diseases, including muscular dystrophies and motor neuron diseases, and are currently being tested in clinical trials.
Abstract
The human transcriptome is composed of a vast RNA population that undergoes further diversification by splicing. Detecting specific splice sites in this large sequence pool is the responsibility of the major and minor spliceosomes in collaboration with numerous splicing factors. This complexity makes splicing susceptible to sequence polymorphisms and deleterious mutations. Indeed, RNA mis-splicing underlies a growing number of human diseases with substantial societal consequences. Here, we provide an overview of RNA splicing mechanisms followed by a discussion of disease-associated errors, with an emphasis on recently described mutations that have provided new insights into splicing regulation. We also discuss emerging strategies for splicing-modulating therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kim, M. S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Treutlein, B., Gokce, O., Quake, S. R. & Sudhof, T. C. Cartography of neurexin alternative splicing mapped by single-molecule long-read mRNA sequencing. Proc. Natl Acad. Sci. USA 111, E1291–E1299 (2014). Long-read sequencing of full-length neurexin mRNAs from pre-frontal cortex is performed to determine the extent of alternative splicing and provide evidence for thousands of neurexin isoforms.
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Gerstein, M. B. et al. Comparative analysis of the transcriptome across distant species. Nature 512, 445–448 (2014).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Singh, R. K. & Cooper, T. A. Pre-mRNA splicing in disease and therapeutics. Trends Mol. Med. 18, 472–482 (2012).
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 56, 55–66 (2014). Demonstrates that circRNAs are generated co-transcriptionally, that the circularization process competes with linear splicing and then MBNL functions as an auto-regulatory factor in circRNA biogenesis.
Barry, G. et al. The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing. Mol. Psychiatry 19, 486–494 (2014).
Sharp, P. A. Split genes and RNA splicing. Cell 77, 805–815 (1994).
Dujardin, G. et al. How slow RNA polymerase II elongation favors alternative exon skipping. Mol. Cell 54, 683–690 (2014).
Maquat, L. E. et al. Processing of human β-globin mRNA precursor to mRNA is defective in three patients with β+-thalassemia. Proc. Natl Acad. Sci. USA 77, 4287–4291 (1980).
Spritz, R. A. et al. Base substitution in an intervening sequence of a β+-thalassemic human globin gene. Proc. Natl Acad. Sci. USA 78, 2455–2459 (1981).
Busslinger, M., Moschonas, N. & Flavell, R. A. β+ thalassemia: aberrant splicing results from a single point mutation in an intron. Cell 27, 289–298 (1981).
Takeshima, Y. et al. Mutation spectrum of the dystrophin gene in 442 Duchenne/Becker muscular dystrophy cases from one Japanese referral center. J. Hum. Genet. 55, 379–388 (2010).
Fletcher, S. et al. Antisense suppression of donor splice site mutations in the dystrophin gene transcript. Mol. Genet. Genom. Med. 1, 162–173 (2013).
Chu, C. S., Trapnell, B. C., Curristin, S., Cutting, G. R. & Crystal, R. G. Genetic basis of variable exon 9 skipping in cystic fibrosis transmembrane conductance regulator mRNA. Nat. Genet. 3, 151–156 (1993).
Tsui, L. C. & Dorfman, R. The cystic fibrosis gene: a molecular genetic perspective. Cold Spring Harb. Perspect. Med. 3, a009472 (2013).
Niblock, M. & Gallo, J. M. Tau alternative splicing in familial and sporadic tauopathies. Biochem. Soc. Trans. 40, 677–680 (2012).
Gruenbaum, Y. & Medalia, O. Lamins: the structure and protein complexes. Curr. Opin. Cell Biol. 32, 7–12 (2015).
Luo, Y. B., Mastaglia, F. L. & Wilton, S. D. Normal and aberrant splicing of LMNA. J. Med. Genet. 51, 215–223 (2014).
Muchir, A. et al. Identification of mutations in the gene encoding lamins A/C in autosomal dominant limb girdle muscular dystrophy with atrioventricular conduction disturbances (LGMD1B). Hum. Mol. Genet. 9, 1453–1459 (2000).
Morel, C. F. et al. A LMNA splicing mutation in two sisters with severe Dunnigan-type familial partial lipodystrophy type 2. J. Clin. Endocrinol. Metab. 91, 2689–2695 (2006).
Eriksson, M. et al. Recurrent de novo point mutations in lamin A cause Hutchinson–Gilford progeria syndrome. Nature 423, 293–298 (2003).
De Sandre-Giovannoli, A. et al. Lamin a truncation in Hutchinson–Gilford progeria. Science 300, 2055 (2003).
Otomo, J. et al. Electrophysiological and histopathological characteristics of progressive atrioventricular block accompanied by familial dilated cardiomyopathy caused by a novel mutation of lamin A/C gene. J. Cardiovasc. Electrophysiol 16, 137–145 (2005).
Liu, M. M. & Zack, D. J. Alternative splicing and retinal degeneration. Clin. Genet. 84, 142–149 (2013).
Tanackovic, G. et al. PRPF mutations are associated with generalized defects in spliceosome formation and pre-mRNA splicing in patients with retinitis pigmentosa. Hum. Mol. Genet. 20, 2116–2130 (2011).
Linder, B. et al. Identification of a PRPF4 loss-of-function variant that abrogates U4/U6.U5 tri-snRNP integration and is associated with retinitis pigmentosa. PLoS ONE 9, e111754 (2014).
Vaclavik, V., Gaillard, M. C., Tiab, L., Schorderet, D. F. & Munier, F. L. Variable phenotypic expressivity in a Swiss family with autosomal dominant retinitis pigmentosa due to a T494M mutation in the PRPF3 gene. Mol. Vis. 16, 467–475 (2010).
Maubaret, C. G. et al. Autosomal dominant retinitis pigmentosa with intrafamilial variability and incomplete penetrance in two families carrying mutations in PRPF8. Invest. Ophthalmol. Vis. Sci. 52, 9304–9309 (2011).
Venturini, G., Rose, A. M., Shah, A. Z., Bhattacharya, S. S. & Rivolta, C. CNOT3 is a modifier of PRPF31 mutations in retinitis pigmentosa with incomplete penetrance. PLoS Genet. 8, e1003040 (2012).
Comitato, A. et al. Mutations in splicing factor PRPF3, causing retinal degeneration, form detrimental aggregates in photoreceptor cells. Hum. Mol. Genet. 16, 1699–1707 (2007).
Mozaffari-Jovin, S. et al. Inhibition of RNA helicase Brr2 by the C-terminal tail of the spliceosomal protein Prp8. Science 341, 80–84 (2013).
Chen, X. et al. PRPF4 mutations cause autosomal dominant retinitis pigmentosa. Hum. Mol. Genet. 23, 2926–2939 (2014).
Kevany, B. M. & Palczewski, K. Phagocytosis of retinal rod and cone photoreceptors. Physiology (Bethesda) 25, 8–15 (2010).
Farkas, M. H. et al. Mutations in pre-mRNA processing factors 3, 8, and 31 cause dysfunction of the retinal pigment epithelium. Am. J. Pathol. 184, 2641–2652 (2014).
David, C. J. & Manley, J. L. Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged. Genes Dev. 24, 2343–2364 (2010).
Adler, A. S. et al. An integrative analysis of colon cancer identifies an essential function for PRPF6 in tumor growth. Genes Dev. 28, 1068–1084 (2014). This study identifies the tri-snRNP complex protein PRPF6 as an oncogenic driver of colon cancer proliferation by promoting selective splicing of growth regulatory gene transcripts.
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Kurtovic-Kozaric, A. et al. PRPF8 defects cause missplicing in myeloid malignancies. Leukemia 29, 126–136 (2014).
Yoshida, K. & Ogawa, S. Splicing factor mutations and cancer. Wiley Interdiscip. Rev. RNA 5, 445–459 (2014).
Malcovati, L. et al. SF3B1 mutation identifies a distinct subset of myelodysplastic syndrome with ring sideroblasts. Blood 126, 233–241 (2015).
Shirai, C. L. et al. Mutant U2AF1 expression alters hematopoiesis and pre-mRNA splicing in vivo. Cancer Cell 27, 631–643 (2015).
Okeyo-Owuor, T. et al. U2AF1 mutations alter sequence specificity of pre-mRNA binding and splicing. Leukemia 29, 909–917 (2015).
Ilagan, J. O. et al. U2AF1 mutations alter splice site recognition in hematological malignancies. Genome Res. 25, 14–26 (2015).
Zhang, J. et al. Disease-associated mutation in SRSF2 misregulates splicing by altering RNA-binding affinities. Proc. Natl Acad. Sci. USA 112, E4726–E4734 (2015).
Kim, E. et al. SRSF2 mutations contribute to myelodysplasia by mutant-specific effects on exon recognition. Cancer Cell 27, 617–630 (2015).
Komeno, Y. et al. SRSF2 is essential for hematopoiesis, and its myelodysplastic syndrome-related mutations dysregulate alternative pre-mRNA splicing. Mol. Cell. Biol. 35, 3071–3082 (2015).
Hsu, T. Y. et al. The spliceosome is a therapeutic vulnerability in MYC-driven cancer. Nature 525, 384–388 (2015).
Bonnal, S., Vigevani, L. & Valcarcel, J. The spliceosome as a target of novel antitumour drugs. Nat. Rev. Drug Discov. 11, 847–859 (2012).
Abdel-Salam, G. M. et al. A homozygous mutation in RNU4ATAC as a cause of microcephalic osteodysplastic primordial dwarfism type I (MOPD I) with associated pigmentary disorder. Am. J. Med. Genet. A 155, 2885–2896 (2011).
Edery, P. et al. Association of TALS developmental disorder with defect in minor splicing component U4atac snRNA. Science 332, 240–243 (2011).
He, H. et al. Mutations in U4atac snRNA, a component of the minor spliceosome, in the developmental disorder MOPD I. Science 332, 238–240 (2011).
Jafarifar, F., Dietrich, R. C., Hiznay, J. M. & Padgett, R. A. Biochemical defects in minor spliceosome function in the developmental disorder MOPD I. RNA 20, 1078–1089 (2014).
Younis, I. et al. Minor introns are embedded molecular switches regulated by highly unstable U6atac snRNA. eLife 2, e00780 (2013). The authors propose that minor introns function as molecular switches which are regulated by U6atac stability, which is itself controlled by the stress-activated kinase p38 MAPK.
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Konig, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Kwon, S. C. et al. The RNA-binding protein repertoire of embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1122–1130 (2013).
Moore, M. J. et al. Mapping Argonaute and conventional RNA-binding protein interactions with RNA at single-nucleotide resolution using HITS-CLIP and CIMS analysis. Nat. Protoc. 9, 263–293 (2014).
Daneholt, B. Assembly and transport of a premessenger RNP particle. Proc. Natl Acad. Sci. USA 98, 7012–7017 (2001).
King, O. D., Gitler, A. D. & Shorter, J. The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease. Brain Res. 1462, 61–80 (2012).
Kim, H. J. et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495, 467–473 (2013). Reports that hereditary mutations in the prion-like domains of HNRNPA1 and HNRNPA2B1 cause multisystem proteinopathy and ALS and drive the formation of cytoplasmic inclusions.
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Han, T. W. et al. Cell-free formation of RNA granules: bound RNAs identify features and components of cellular assemblies. Cell 149, 768–779 (2012).
Marangi, G. & Traynor, B. J. Genetic causes of amyotrophic lateral sclerosis: new genetic analysis methodologies entailing new opportunities and challenges. Brain Res. 1607, 75–93 (2014).
Renton, A. E., Chio, A. & Traynor, B. J. State of play in amyotrophic lateral sclerosis genetics. Nat. Neurosci. 17, 17–23 (2014).
Ling, S. C., Polymenidou, M. & Cleveland, D. W. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79, 416–438 (2013).
Goodwin, M. & Swanson, M. S. RNA-binding protein misregulation in microsatellite expansion disorders. Adv. Exp. Med. Biol. 825, 353–388 (2014).
Prudencio, M. et al. Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat. Neurosci. 18, 1175–1182 (2015).
Lee, Y. B. et al. Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep. 5, 1178–1186 (2013).
Buratti, E. & Baralle, F. E. TDP-43: gumming up neurons through protein–protein and protein–RNA interactions. Trends Biochem. Sci. 37, 237–247 (2012).
Arnold, E. S. et al. ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc. Natl Acad. Sci. USA 110, E736–E745 (2013). Demonstration that controlled expression of ALS-associated TDP-43 mutations in a mouse model causes splicing dysregulation of specific targets and progressive motor neuron loss in the absence of TDP-43 aggregation or nuclear depletion.
Lagier-Tourenne, C. et al. Divergent roles of ALS-linked proteins FUS/TLS and TDP-43 intersect in processing long pre-mRNAs. Nat. Neurosci. 15, 1488–1497 (2012).
Nakaya, T., Alexiou, P., Maragkakis, M., Chang, A. & Mourelatos, Z. FUS regulates genes coding for RNA-binding proteins in neurons by binding to their highly conserved introns. RNA 19, 498–509 (2013).
Bai, B. et al. U1 small nuclear ribonucleoprotein complex and RNA splicing alterations in Alzheimer's disease. Proc. Natl Acad. Sci. USA 110, 16562–16567 (2013).
Li, Y. I., Sanchez-Pulido, L., Haerty, W. & Ponting, C. P. RBFOX and PTBP1 proteins regulate the alternative splicing of micro-exons in human brain transcripts. Genome Res. 25, 1–13 (2015).
Sibley, C. R. et al. Recursive splicing in long vertebrate genes. Nature 521, 371–375 (2015).
Duff, M. O. et al. Genome-wide identification of zero nucleotide recursive splicing in Drosophila. Nature 521, 376–379 (2015).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Tollervey, J. R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Ling, J. P., Pletnikova, O., Troncoso, J. C. & Wong, P. C. TDP-43 repression of nonconserved cryptic exons is compromised in ALS-FTD. Science 349, 650–655 (2015). Shows that TDP-43 depletion results in splicing of non-conserved cryptic exons that are often located in distal regions of long introns and that loss of TDP-43-mediated cryptic exon splicing repression is a potential pathogenic factor in ALS and FTD.
Nelson, S. B. & Valakh, V. Excitatory/inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron 87, 684–698 (2015).
Irimia, M. et al. A highly conserved program of neuronal microexons is misregulated in autistic brains. Cell 159, 1511–1523 (2014).
Gehman, L. T. et al. The splicing regulator Rbfox1 (A2BP1) controls neuronal excitation in the mammalian brain. Nat. Genet. 43, 706–711 (2011).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Martin, C. L. et al. Cytogenetic and molecular characterization of A2BP1/FOX1 as a candidate gene for autism. Am. J. Med. Genet. B Neuropsychiatr. Genet. 144, 869–876 (2007).
Weyn-Vanhentenryck, S. M. et al. HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Rep. 6, 1139–1152 (2014).
Kole, R., Krainer, A. R. & Altman, S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat. Rev. Drug Discov. 11, 125–140 (2012).
Ruegg, U. T. Pharmacological prospects in the treatment of Duchenne muscular dystrophy. Curr. Opin. Neurol. 26, 577–584 (2013).
Kole, R. & Krieg, A. M. Exon skipping therapy for Duchenne muscular dystrophy. Adv. Drug Deliv. Rev. 87, 104–107 (2015).
Osorio, F. G. et al. Splicing-directed therapy in a new mouse model of human accelerated aging. Sci. Transl. Med. 3, 106ra107 (2011).
Faravelli, I., Nizzardo, M., Comi, G. P. & Corti, S. Spinal muscular atrophy — recent therapeutic advances for an old challenge. Nat. Rev. Neurol. 11, 351–359 (2015).
Li, D. K., Tisdale, S., Lotti, F. & Pellizzoni, L. SMN control of RNP assembly: from post-transcriptional gene regulation to motor neuron disease. Semin. Cell Dev. Biol. 32, 22–29 (2014).
Cartegni, L., Hastings, M. L., Calarco, J. A., de Stanchina, E. & Krainer, A. R. Determinants of exon 7 splicing in the spinal muscular atrophy genes, SMN1 and SMN2. Am. J. Hum. Genet. 78, 63–77 (2006).
Hua, Y., Vickers, T. A., Okunola, H. L., Bennett, C. F. & Krainer, A. R. Antisense masking of an hnRNP A1/A2 intronic splicing silencer corrects SMN2 splicing in transgenic mice. Am. J. Hum. Genet. 82, 834–848 (2008).
Hua, Y. et al. Motor neuron cell-nonautonomous rescue of spinal muscular atrophy phenotypes in mild and severe transgenic mouse models. Genes Dev. 29, 288–297 (2015). Although SMA is a motor neuron disease, this study uses a combination of splice-switching and decoy oligonucleotides to provide evidence that disease is not specific to motor neurons in a mouse model of SMA.
Lopez Castel, A., Cleary, J. D. & Pearson, C. E. Repeat instability as the basis for human diseases and as a potential target for therapy. Nat. Rev. Mol. Cell Biol. 11, 165–170 (2010).
Wang, E. T. et al. Transcriptome-wide regulation of pre-mRNA splicing and mRNA localization by muscleblind proteins. Cell 150, 710–724 (2012).
Charizanis, K. et al. Muscleblind-like 2-mediated alternative splicing in the developing brain and dysregulation in myotonic dystrophy. Neuron 75, 437–450 (2012).
Wang, E. T. et al. Antagonistic regulation of mRNA expression and splicing by CELF and MBNL proteins. Genome Res. 25, 858–871 (2015).
Wheeler, T. M. et al. Reversal of RNA dominance by displacement of protein sequestered on triplet repeat RNA. Science 325, 336–339 (2009).
Leger, A. J. et al. Systemic delivery of a peptide-linked morpholino oligonucleotide neutralizes mutant RNA toxicity in a mouse model of myotonic dystrophy. Nucleic Acid. Ther. 23, 109–117 (2013).
Wheeler, T. M. et al. Targeting nuclear RNA for in vivo correction of myotonic dystrophy. Nature 488, 111–115 (2012).
Kendall, G. C. et al. Dantrolene enhances antisense-mediated exon skipping in human and mouse models of Duchenne muscular dystrophy. Sci. Transl. Med. 4, 164ra160 (2012).
Cherry, J. J. et al. Enhancement of SMN protein levels in a mouse model of spinal muscular atrophy using novel drug-like compounds. EMBO Mol. Med. 5, 1035–1050 (2013).
Naryshkin, N. A. et al. SMN2 splicing modifiers improve motor function and longevity in mice with spinal muscular atrophy. Science 345, 688–693 (2014). A minigene splicing reporter combined with chemical screening and optimization were used to identify small molecule compounds that activate SMN2 exon 7 splicing and increase SMN levels.
Palacino, J. et al. SMN2 splice modulators enhance U1-pre-mRNA association and rescue SMA mice. Nat. Chem. Biol. 11, 511–517 (2015); erratum 11, 741 (2015).
Childs-Disney, J. L. et al. Induction and reversal of myotonic dystrophy type 1 pre-mRNA splicing defects by small molecules. Nat. Commun. 4, 2044 (2013).
Childs-Disney, J. L. et al. Structure of the myotonic dystrophy type 2 RNA and designed small molecules that reduce toxicity. ACS Chem. Biol. 9, 538–550 (2014).
Warf, M. B., Nakamori, M., Matthys, C. M., Thornton, C. A. & Berglund, J. A. Pentamidine reverses the splicing defects associated with myotonic dystrophy. Proc. Natl Acad. Sci. USA 106, 18551–18556 (2009).
Hoskins, J. W. et al. Lomofungin and dilomofungin: inhibitors of MBNL1–CUG RNA binding with distinct cellular effects. Nucleic Acids Res. 42, 6591–6602 (2014).
Jahromi, A. H. et al. A novel CUGexp. MBNL1 inhibitor with therapeutic potential for myotonic dystrophy type 1. ACS Chem. Biol. 8, 1037–1043 (2013).
Niland, C. N., Merry, C. R. & Khalil, A. M. Emerging roles for long non-coding RNAs in cancer and neurological disorders. Front. Genet. 3, 25 (2012).
Yang, L., Froberg, J. E. & Lee, J. T. Long noncoding RNAs: fresh perspectives into the RNA world. Trends Biochem. Sci. 39, 35–43 (2014).
Tilgner, H. et al. Deep sequencing of subcellular RNA fractions shows splicing to be predominantly co-transcriptional in the human genome but inefficient for lncRNAs. Genome Res. 22, 1616–1625 (2012).
Gratten, J., Wray, N. R., Keller, M. C. & Visscher, P. M. Large-scale genomics unveils the genetic architecture of psychiatric disorders. Nat. Neurosci. 17, 782–790 (2014).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Conn, S. J. et al. The RNA binding protein quaking regulates formation of circRNAs. Cell 160, 1125–1134 (2015).
Xiong, H. Y. et al. The human splicing code reveals new insights into the genetic determinants of disease. Science 347, 1254806 (2015). Comprehensive study showing the utility of a machine learning technique that scores sequence variants for splicing impact and reveals thousands of disease-associated mutations.
Papasaikas, P., Tejedor, J. R., Vigevani, L. & Valcarcel, J. Functional splicing network reveals extensive regulatory potential of the core spliceosomal machinery. Mol. Cell 57, 7–22 (2015).
Turunen, J. J., Niemela, E. H., Verma, B. & Frilander, M. J. The significant other: splicing by the minor spliceosome. Wiley Interdiscip. Rev. RNA 4, 61–76 (2013).
Braunschweig, U. et al. Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res. 24, 1774–1786 (2014). Although intron retention is the most common form of splicing regulation in plants, this study uncovers widespread intron retention in mammals and proposes intron retention as a regulatory mechanism to control transcript levels by NMD and nuclear sequestration and turnover.
Ibrahim, E. C. et al. Weak definition of IKBKAP exon 20 leads to aberrant splicing in familial dysautonomia. Hum. Mutat. 28, 41–53 (2007).
Muntoni, F., Torelli, S. & Ferlini, A. Dystrophin and mutations: one gene, several proteins, multiple phenotypes. Lancet Neurol. 2, 731–740 (2003).
Disset, A. et al. An exon skipping-associated nonsense mutation in the dystrophin gene uncovers a complex interplay between multiple antagonistic splicing elements. Hum. Mol. Genet. 15, 999–1013 (2006).
Samaranch, L. et al. PINK1-linked parkinsonism is associated with Lewy body pathology. Brain 133, 1128–1142 (2010).
Iovino, M. et al. The novel MAPT mutation K298E: mechanisms of mutant tau toxicity, brain pathology and tau expression in induced fibroblast-derived neurons. Acta Neuropathol. 127, 283–295 (2014).
Korvatska, O. et al. Altered splicing of ATP6AP2 causes X-linked parkinsonism with spasticity (XPDS). Hum. Mol. Genet. 22, 3259–3268 (2013).
Tanackovic, G. et al. A missense mutation in PRPF6 causes impairment of pre-mRNA splicing and autosomal-dominant retinitis pigmentosa. Am. J. Hum. Genet. 88, 643–649 (2011).
Cvackova, Z., Mateju, D. & Stanek, D. Retinitis pigmentosa mutations of SNRNP200 enhance cryptic splice-site recognition. Hum. Mutat. 35, 308–317 (2014).
Lorson, C. L., Hahnen, E., Androphy, E. J. & Wirth, B. A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc. Natl Acad. Sci. USA 96, 6307–6311 (1999).
Lefebvre, S. et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80, 155–165 (1995).
Sun, S. et al. ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP. Nat. Commun. 6, 6171 (2015).
Guo, W. et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat. Med. 18, 766–773 (2012).
Vieira, N. M. et al. A defect in the RNA-processing protein HNRPDL causes limb-girdle muscular dystrophy 1G (LGMD1G). Hum. Mol. Genet. 23, 4103–4110 (2014).
Bartoletti-Stella, A. et al. Messenger RNA processing is altered in autosomal dominant leukodystrophy. Hum. Mol. Genet. 24, 2746–2756 (2015).
Acknowledgements
The authors regret that many important studies were not cited owing to space limitations. Work in the authors' laboratories is funded by grants to M.S.S. from the US National Institutes of Health (NIH AR046799, NS058901), the Muscular Dystrophy Association (MDA276063), the W.M. Keck Foundation (F013635) and the Marigold Foundation. M.M.S. is the recipient of an NIH pre-doctoral traineeship (NIAMS T32 AR7605-15).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Long non-coding RNAs
-
(lncRNAs). RNAs of >200 nucleotides in length that generally do not encode proteins.
- Pseudogenes
-
Non-functional versions of genes that are generated either by duplication and mutation or by retrotransposition.
- Splicing factors
-
Proteins that participate in splicing regulation but are not stable constituents of small nuclear ribonucleoprotein particles (snRNPs).
- Single-nucleotide polymorphisms
-
(SNPs). Variations in individual nucleotides that are common in the human genome and can influence splicing regulation.
- Nonsense-mediated decay
-
(NMD). A process of enhanced RNA turnover induced by a premature termination codon (PTC) which is designed to block the synthesis of truncated proteins and modulate the appearance of full-length proteins during development.
- Penetrance
-
The percentage of individuals carrying a disease mutation who show clinical symptoms. Incomplete, or reduced, penetrance occurs when not all individuals with a particular genetic mutation develop the associated disease.
- Expressivity
-
The degree to which a mutant gene is phenotypically expressed. Variable expressivity refers to the symptomatic range that is displayed by different individuals with the same mutation.
- Core consensus sequences
-
Conserved RNA sequence motifs, including the 5′ and 3′ splice sites and the branch point region, which are required for spliceosome recruitment.
- Branch point
-
(BP). A partially conserved sequence, generally <50 nucleotides upstream of the 3′ splice site (see Box 1), that reacts with the 5′ splice site during the first step of the splicing reaction.
- Spliceosome
-
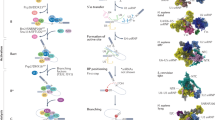
The large RNA–protein complex that catalyses splicing and is composed of multiple small nuclear RNAs (snRNAs) and many associated protein factors. Whereas the major and minor spliceosomes both contain U5, the other snRNA components differ (for the major spliceosome, U1, U2, U4, U6; for the minor spliceosome, U11, U12, U4atac, U6atac) (see Fig. 2).
- Tri-snRNP
-
A preassembled complex of U4 snRNA hybridized to U6 (U4/U6 or U4atac/U6atac) that also contains U5 (U4/U6.U5) together with associated proteins (see Fig. 2).
- Haploinsufficiency
-
A condition due to inactivating mutations in one copy of a gene when expression from the remaining copy is insufficient to produce an unaffected phenotype.
- HITS-CLIP
-
(High-throughput sequencing of RNA isolated by crosslinking immunoprecipitation; also known as CLIP–seq). A technique to map the binding sites of splicing, and other, factors on target RNAs. Related techniques include photoactivatable-ribonucleoside-enhanced-CLIP (PAR-CLIP) and individual-nucleotide resolution CLIP (iCLIP).
- Cryptic splice sites
-
Splice sites that are not normally recognized by the spliceosome but can be activated either by mutations in cis-acting elements or trans-acting factors.
- Splicing regulatory elements
-
RNA sequence motifs in either exons or introns that modulate splicing primarily by binding trans-acting splicing regulatory factors.
- Morpholino
-
An antisense oligomer with standard nucleic acid bases but instead of deoxyribose contains a six-member morpholine ring linked with phosphorodiamidate (PMO). PMOs function by steric blocking and vivo-morpholinos, composed of a morpholino oligomer covalently attached to an octa-guanidine dendrimer, are optimized for in vivo delivery.
- Human transcriptome
-
All of the RNAs transcribed from the human genome.
Rights and permissions
About this article
Cite this article
Scotti, M., Swanson, M. RNA mis-splicing in disease. Nat Rev Genet 17, 19–32 (2016). https://doi.org/10.1038/nrg.2015.3
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2015.3
This article is cited by
-
High-throughput RNA isoform sequencing using programmed cDNA concatenation
Nature Biotechnology (2024)
-
Whole genome sequencing enables new genetic diagnosis for inherited retinal diseases by identifying pathogenic variants
npj Genomic Medicine (2024)
-
Intronic position +9 and −9 are potentially splicing sites boundary from intronic variants analysis of whole exome sequencing data
BMC Medical Genomics (2023)
-
Identified eleven exon variants in PKD1 and PKD2 genes that altered RNA splicing by minigene assay
BMC Genomics (2023)
-
Benchmarking splice variant prediction algorithms using massively parallel splicing assays
Genome Biology (2023)