Abstract
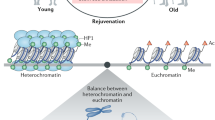
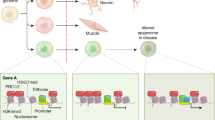
Cellular commitment to a specific lineage is controlled by differential silencing of genes, which in turn depends on epigenetic processes such as DNA methylation and histone modification. During early embryogenesis, the mammalian genome is 'wiped clean' of most epigenetic modifications, which are progressively re-established during embryonic development. Thus, the epigenome of each mature cellular lineage carries the record of its developmental history. The subsequent trajectory and pattern of development are also responsive to environmental influences, and such plasticity is likely to have an epigenetic basis. Epigenetic marks may be transmitted across generations, either directly by persisting through meiosis or indirectly through replication in the next generation of the conditions in which the epigenetic change occurred. Developmental plasticity evolved to match an organism to its environment, and a mismatch between the phenotypic outcome of adaptive plasticity and the current environment increases the risk of metabolic and cardiovascular disease. These considerations point to epigenetic processes as a key mechanism that underpins the developmental origins of chronic noncommunicable disease. Here, we review the evidence that environmental influences during mammalian development lead to stable changes in the epigenome that alter the individual's susceptibility to chronic metabolic and cardiovascular disease, and discuss the clinical implications.
Key Points
-
Developmental plasticity enables an organism to respond to environmental cues and adjust its phenotypic development to match its environment
-
Developmental plasticity is effected, at least in part, by epigenetic changes that are established in early life and modulate gene expression during development and maturity
-
In mammals, the window during which the epigenome is susceptible to nutritional cues extends from conception to at least weaning
-
Mismatch between the early and mature environments may result in inappropriate patterns of epigenetic changes and gene expression that increase subsequent susceptibility to metabolic and cardiovascular diseases
-
The available evidence suggests that interventions to prevent metabolic and cardiovascular diseases should focus on the prenatal and early postnatal periods
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
De Prins, F. A. & Van Assche, F. A. Intrauterine growth retardation and development of endocrine pancreas in the experimental rat. Biol. Neonat. 41, 16–21 (1982).
Godfrey, K. The 'developmental origins' hypothesis: epidemiology. In Developmental Origins of Health and Disease (Eds Gluckman, P. D. & Hanson, M. A.) 6–32 (Cambridge University Press, Cambridge, 2006).
Hales, C. N. & Barker, D. J. Type 2 (non-insulin-dependent) diabetes mellitus: the thrifty phenotype hypothesis. Diabetologia 35, 595–601 (1992).
Painter, R. C., Roseboom, T. J. & Bleker, O. P. Prenatal exposure to the Dutch famine and disease in later life: an overview. Reprod. Toxicol. 20, 345–352 (2005).
Osmond, C., Barker, D. J., Winter, P. D., Fall, C. H. & Simmonds, S. J. Early growth and death from cardiovascular disease in women. BMJ 307, 1519–1524 (1993).
Yajnik, C. S. Early life origins of insulin resistance and type 2 diabetes in India and other Asian countries. J. Nutr. 134, 205–210 (2004).
Kuzawa, C. W. & Adair, L. S. Lipid profiles in adolescent Filipinos: relation to birth weight and maternal energy status during pregnancy. Am. J. Clin. Nutr. 77, 960–966 (2003).
Singhal, A. Early nutrition and long-term cardiovascular health. Nutr. Rev. 64, S44–S49 (2006).
Boney, C. M., Verma, A., Tucker, R. & Vohr, B. R. Metabolic syndrome in childhood: association with birth weight, maternal obesity, and gestational diabetes mellitus. Pediatrics 115, 290–296 (2005).
Kuzawa, C. W. et al. Evolution, developmental plasticity, and metabolic disease. In Evolution in Health and Disease (Eds Stearns, S. C. and Koella, J. C.) 253–264 (Oxford University Press, Oxford, 2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Whincup, P. H. et al. Early evidence of ethnic differences in cardiovascular risk: cross sectional comparison of British South Asian and white children. BMJ 324, 635 (2002).
McKeigue, P. M. Metabolic consequences of obesity and body fat pattern: lessons from migrant studies. Ciba Found. Symp. 201, 54–64 (1996).
Dhandapany, P. S. et al. A common MYBPC3 (cardiac myosin binding protein C) variant associated with cardiomyopathies in South Asia. Nat. Genet. 41, 187–191 (2009).
Yajnik, C. S. et al. Neonatal anthropometry: the thin-fat Indian baby. The Pune maternal nutrition study. Int. J. Obes. Relat. Metab. Disord. 27, 173–180 (2003).
Dabelea, D., Knowler, W. C. & Pettitt, D. J. Effect of diabetes in pregnancy on offspring: follow-up research in the Pima Indians. J. Matern. Fetal Med. 9, 83–88 (2000).
West-Eberhard, M. J. Developmental Plasticity and Evolution (Oxford University Press, New York, 2003).
Gluckman, P. D., Hanson, M. A. & Beedle, A. S. Early life events and their consequences for later disease: a life history and evolutionary perspective. Am. J. Hum. Biol. 19, 1–19 (2007).
Gluckman, P. D. & Hanson, M. A. Maternal constraint of fetal growth and its consequences. Semin. Fetal Neonatal Med. 9, 419–425 (2004).
Gluckman, P. D. & Hanson, M. A. Developmental Origins of Health and Disease (Cambridge University Press, Cambridge, 2006).
Waddington, C. H. The Strategy of the Genes: a Discussion of Some Aspects of Theoretical Biology (George Allen & Unwin Ltd, London, 1957).
Goldberg, A. D., Allis, C. D. & Bernstein, E. Epigenetics: a landscape takes shape. Cell 128, 635–638 (2007).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet. 39, 457–466 (2007).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Nightingale, K. P., O'Neill, L. P. & Turner, B. M. Histone modifications: signalling receptors and potential elements of a heritable epigenetic code. Curr. Opin. Genet. Dev. 16, 125–136 (2006).
Amaral, P. P. & Mattick, J. S. Noncoding RNA in development. Mamm. Genome 19, 454–492 (2008).
Panning, B. X-chromosome inactivation: the molecular basis of silencing. J. Biol. 7, 1–4 (2008).
Branco, M. R., Oda, M. & Reik, W. Safeguarding parental identity: Dnmtl maintains imprints during epigenetic reprogramming in early embryogenesis. Genes Dev. 22, 1567–1571 (2008).
Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447, 425–432 (2007).
Matzke, M. A., Mette, M. F. & Matzke, A. J. Transgene silencing by the host genome defense: implications for the evolution of epigenetic control mechanisms in plants and vertebrates. Plant Mol. Biol. 43, 401–415 (2000).
Schlichting, C. D. Origins of differentiation via phenotypic plasticity. Evol. Dev. 5, 98–105 (2003).
Renfree, M. B., Ager, E. I., Shaw, G. & Pask, A. J. Genomic imprinting in marsupial placentation. Reproduction 136, 523–531 (2008).
Nafee, T. M., Farrell, W. E., Carroll, W. D., Fryer, A. A. & Ismail, K. M. Epigenetic control of fetal gene expression. BJOG 115, 158–168 (2007).
Ng, R. K. et al. Epigenetic restriction of embryonic cell lineage fate by methylation of Elf5 . Nat. Cell Biol. 10, 1280–1290 (2008).
Yagi, S. et al. DNA methylation profile of tissue-dependent and differentially methylated regions (T-DMRs) in mouse promoter regions demonstrating tissue-specific gene expression. Genome Res. 18, 1969–1978 (2008).
Song, F. et al. Tissue specific differentially methylated regions (TDMR): changes in DNA methylation during development. Genomics 93, 130–139 (2009).
Shechter, D. et al. Analysis of histones in Xenopus laevis. I. A distinct index of enriched variants and modifications exists in each cell type and is remodeled during developmental transitions. J. Biol. Chem. 284, 1064–1074 (2009).
Nicklay, J. J. et al. Analysis of histones in Xenopus laevis. II. Mass spectrometry reveals an index of cell-type specific modifications on H3 and H4. J. Biol. Chem. 284, 1075–1085 (2009).
Farthing, C. R. et al. Global mapping of DNA methylation in mouse promoters reveals epigenetic reprogramming of pluripotency genes. PLoS Genet. 4, e1000116 (2008).
Waterland, R. A. & Jirtle, R. L. Early nutrition, epigenetic changes at transposons and imprinted genes, and enhanced susceptibility to adult chronic diseases. Nutrition 20, 63–68 (2004).
Dolinoy, D. C., Weidman, J. R., Waterland, R. A. & Jirtle, R. L. Maternal genistein alters coat color and protects Avy mouse offspring from obesity by modifying the fetal epigenome. Environ. Health Perspect. 114, 567–572 (2006).
Waterland, R. A. et al. Maternal methyl supplements increase offspring DNA methylation at Axin Fused . Genesis 44, 401–406 (2006).
Park, J. H., Stoffers, D. A., Nicholls, R. D. & Simmons, R. A. Development of type 2 diabetes following intrauterine growth retardation in rats is associated with progressive epigenetic silencing of Pdx1 . J. Clin. Invest. 118, 2316–2324 (2008).
Lal, G. et al. Epigenetic regulation of Foxp3 expression in regulatory T cells by DNA methylation. J. Immunol. 182, 259–273 (2009).
Lillycrop, K. A. et al. Induction of altered epigenetic regulation of the hepatic glucocorticoid receptor in the offspring of rats fed a protein-restricted diet during pregnancy suggests that reduced DNA methyltransferase-1 expression is involved in impaired DNA methylation and changes in histone modifications. Br. J. Nutr. 97, 1064–1073 (2007).
Lillycrop, K. A. et al. Feeding pregnant rats a protein-restricted diet persistently alters the methylation of specific cytosines in the hepatic PPARα promoter of the offspring. Br. J. Nutr. 100, 278–282 (2008).
Bogdarina, I. et al. Epigenetic modification of the renin–angiotensin system in the fetal programming of hypertension. Circ. Res. 100, 520–526 (2007).
Pham, T. D. et al. Uteroplacental insufficiency increases apoptosis and alters p53 gene methylation in the full-term IUGR rat kidney. Am. J. Physiol. Regul. Integr. Comp. Physiol. 285, R962–R970 (2003).
Fu, Q. et al. Uteroplacental insufficiency induces site-specific changes in histone H3 covalent modification and affects DNA-histone H3 positioning in day 0 IUGR rat liver. Physiol. Genomics 20, 108–116 (2004).
Raychaudhuri, N., Raychaudhuri, S., Thamotharan, M. & Devaskar, S. U. Histone code modifications repress glucose transporter 4 expression in the intrauterine growth-restricted offspring. J. Biol. Chem. 283, 13611–13626 (2008).
Aagaard-Tillery, K. M. et al. Developmental origins of disease and determinants of chromatin structure: maternal diet modifies the primate fetal epigenome. J. Mol. Endocrinol. 41, 91–102 (2008).
Sinclair, K. D. et al. DNA methylation, insulin resistance, and blood pressure in offspring determined by maternal periconceptional B vitamin and methionine status. Proc. Natl Acad. Sci. USA 104, 19351–19356 (2007).
Yajnik, C. S. et al. Vitamin B12 and folate concentrations during pregnancy and insulin resistance in the offspring: the Pune maternal nutrition study. Diabetologia 51, 29–38 (2008).
Villeneuve, L. M. et al. Epigenetic histone H3 lysine 9 methylation in metabolic memory and inflammatory phenotype of vascular smooth muscle cells in diabetes. Proc. Natl Acad. Sci. USA 105, 9047–9052 (2008).
El Osta, A. et al. Transient high glucose causes persistent epigenetic changes and altered gene expression during subsequent normoglycemia. J. Exp. Med. 205, 2409–2417 (2008).
Ling, C. et al. Genetic and epigenetic factors are associated with expression of respiratory chain component NDUFB6 in human skeletal muscle. J. Clin. Invest. 117, 3427–3435 (2007).
Rönn, T. et al. Age influences DNA methylation and gene expression of COX7A1 in human skeletal muscle. Diabetologia 51, 1159–1168 (2008).
Jiang, M. H. et al. Hypermethylation of hepatic Gck promoter in ageing rats contributes to diabetogenic potential. Diabetologia 51, 1525–1533 (2008).
Heijmans, B. T. et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc. Natl Acad. Sci. USA 105, 17046–17049 (2008).
Amor, D. J. & Halliday, J. A review of known imprinting syndromes and their association with assisted reproduction technologies. Hum. Reprod. 23, 2826–2834 (2008).
Waterland, R. A., Lin, J. R., Smith, C. A. & Jirtle, R. L. Post-weaning diet affects genomic imprinting at the insulin-like growth factor 2 (Igf2) locus. Hum. Mol. Genet. 15, 705–716 (2006).
Weaver, I. C. et al. Epigenetic programming by maternal behavior. Nat. Neurosci. 7, 847–854 (2004).
Champagne, F. A. & Curley, J. P. Maternal regulation of estrogen receptor α methylation. Curr. Opin. Pharmacol. 8, 1–5 (2008).
Weksberg, R. et al. Discordant KCNQ1OT1 imprinting in sets of monozygotic twins discordant for Beckwith–Wiedemann syndrome. Hum. Mol. Genet. 11, 1317–1325 (2002).
Rosa, A. et al. Differential methylation of the X-chromosome is a possible source of discordance for bipolar disorder female monozygotic twins. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 147, 459–462 (2008).
Fraga, M. F. et al. Epigenetic differences arise during the lifetime of monozygotic twins. Proc. Natl Acad. Sci. USA 102, 10604–10609 (2005).
Drake, A. J., Walker, B. R. & Seckl, J. R. Intergenerational consequences of fetal programming by in utero exposure to glucocorticoids in rats. Am. J. Physiol. Regul. Integr. Comp. Physiol. 288, R34–R38 (2005).
Burdge, G. C. et al. Dietary protein restriction of pregnant rats in the F0 generation induces altered methylation of hepatic gene promoters in the adult male offspring in the F1 and F2 generations. Br. J. Nutr. 97, 435–439 (2007).
Gluckman, P. D., Hanson, M. A. & Beedle, A. S. Non-genomic transgenerational inheritance of disease risk. Bioessays 29, 145–154 (2007).
Jimenez-Chillaron, J. C. et al. Intergenerational transmission of glucose intolerance and obesity by in utero undernutrition in mice. Diabetes 58, 460–468 (2009).
Rakyan, V. K. et al. Transgenerational inheritance of epigenetic states at the murine Axin (Fu) allele occurs after maternal and paternal transmission. Proc. Natl Acad. Sci. USA 100, 2538–2543 (2003).
Waterland, R. A., Travisano, M. & Tahiliani, K. G. Diet-induced hypermethylation at agouti viable yellow is not inherited transgenerationally through the female. FASEB J. 21, 3380–3385 (2007).
Painter, R. C. et al. Transgenerational effects of prenatal exposure to the Dutch famine on neonatal adiposity and health in later life. BJOG 115, 1243–1249 (2008).
Hitchins, M. et al. Inheritance of a cancer-associated MLH1 germ-line epimutation. N. Engl. J. Med. 356, 697–705 (2007).
Rassoulzadegan, M. et al. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006).
Ibáñez, L., Potau, N., Enriquez, G., Marcos, M. V. & de Zegher, F. Hypergonadotrophinaemia with reduced uterine and ovarian size in women born small-for-gestational-age. Hum. Reprod. 18, 1565–1569 (2003).
Kwong, W. Y. et al. Imprinted gene expression in the rat embryo–fetal axis is altered in response to periconceptional maternal low protein diet. Reproduction 132, 265–277 (2006).
Watkins, A. J. et al. Low protein diet fed exclusively during mouse oocyte maturation leads to behavioural and cardiovascular abnormalities in offspring. J. Physiol. 586, 2231–2244 (2008).
Vickers, M. H., Breier, B. H., Cutfield, W. S., Hofman, P. L. & Gluckman, P. D. Fetal origins of hyperphagia, obesity, and hypertension and postnatal amplification by hypercaloric nutrition. Am. J. Physiol. Endocrinol. Metab. 279, E83–E87 (2000).
Schaefer-Graf, U. M. et al. Birth weight and parental BMI predict overweight in children from mothers with gestational diabetes. Diabetes Care 28, 1745–1750 (2005).
Gluckman, P. D. et al. Metabolic plasticity during mammalian development is directionally dependent on early nutritional status. Proc. Natl Acad. Sci. USA 104, 12796–12800 (2007).
Robinson, S. M. et al. Impact of educational attainment on the quality of young women's diets. Eur. J. Clin. Nutr. 58, 1174–1180 (2004).
Meigs, J. B. et al. Genotype score in addition to common risk factors for prediction of type 2 diabetes. N. Engl. J. Med. 359, 2208–2219 (2008).
Lyssenko, V. et al. Clinical risk factors, DNA variants, and the development of type 2 diabetes. N. Engl. J. Med. 359, 2220–2232 (2008).
Lillycrop, K. A. et al. Dietary protein restriction of pregnant rats induces and folic acid supplementation prevents epigenetic modification of hepatic gene expression in the offspring. J. Nutr. 135, 1382–1386 (2005).
Vickers, M. H. et al. Neonatal leptin treatment reverses developmental programming. Endocrinology 146, 4211–4216 (2005).
Lawlor, D. A. et al. Epidemiologic evidence for the fetal overnutrition hypothesis: findings from the mater-university study of pregnancy and its outcomes. Am. J. Epidemiol. 165, 418–424 (2007).
Waterland, R. A., Travisano, M., Tahiliani, K. G., Rached, M. T. & Mirza, S. Methyl donor supplementation prevents transgenerational amplification of obesity. Int. J. Obes (Lond.) 32, 1373–1379 (2008).
Acknowledgements
P. D. Gluckman, T. Buklijas, F. M. Low and A. S. Beedle are funded by the National Research Centre for Growth and Development (New Zealand). M. A. Hanson is funded by the British Heart Foundation.
Désirée Lie, University of California, Irvine, CA, is the author of and is solely responsible for the content of the learning objectives, questions and answers of the Medscape-accredited continuing medical education activity associated with this article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Gluckman, P., Hanson, M., Buklijas, T. et al. Epigenetic mechanisms that underpin metabolic and cardiovascular diseases. Nat Rev Endocrinol 5, 401–408 (2009). https://doi.org/10.1038/nrendo.2009.102
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrendo.2009.102
This article is cited by
-
Lessons and Applications of Omics Research in Diabetes Epidemiology
Current Diabetes Reports (2024)
-
The contribution of the exposome to the burden of cardiovascular disease
Nature Reviews Cardiology (2023)
-
Dietary supplements and vascular function in hypertensive disorders of pregnancy
Pflügers Archiv - European Journal of Physiology (2023)
-
Lower miR-21/ROS/HNE levels associate with lower glycemia after habit-intervention: DIAPASON study 1-year later
Cardiovascular Diabetology (2022)
-
Beneficial effects of voluntary wheel running on activity rhythms, metabolic state, and affect in a diurnal model of circadian disruption
Scientific Reports (2022)