Key Points
-
Messenger RNA (mRNA) is a pivotal molecule of life, involved in almost all aspects of cell biology.
-
As the subject of basic and applied research for more than 5 decades, mRNA has only recently come into the focus as a potentially powerful drug class able to deliver genetic information.
-
Synthetic mRNA can be engineered to resemble mature and processed mRNA molecules as they occur naturally in the cytoplasm of eukaryotic cells and to transiently deliver proteins.
-
Recent advances addressed challenges inherent to this drug class and provided the basis for a broad spectrum of applications
-
Besides cancer immunotherapies and infectious disease vaccines novel approaches such as in vivo delivery of mRNA to replace or supplement proteins, mRNA-based induction of pluripotent stem cells, or mRNA-assisted delivery of designer nucleases for genome engineering rapidly emerged and entered into pharmaceutical development.
-
This Review gives a comprehensive overview of the current state of mRNA drug technologies, their applications and crucial aspects relevant to mRNA based drug discovery and development.
Abstract
In vitro transcribed (IVT) mRNA has recently come into focus as a potential new drug class to deliver genetic information. Such synthetic mRNA can be engineered to transiently express proteins by structurally resembling natural mRNA. Advances in addressing the inherent challenges of this drug class, particularly related to controlling the translational efficacy and immunogenicity of the IVTmRNA, provide the basis for a broad range of potential applications. mRNA-based cancer immunotherapies and infectious disease vaccines have entered clinical development. Meanwhile, emerging novel approaches include in vivo delivery of IVT mRNA to replace or supplement proteins, IVT mRNA-based generation of pluripotent stem cells and genome engineering using IVT mRNA-encoded designer nucleases. This Review provides a comprehensive overview of the current state of mRNA-based drug technologies and their applications, and discusses the key challenges and opportunities in developing these into a new class of drugs.
Similar content being viewed by others
Main
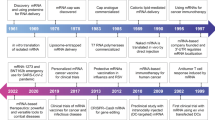
The concept of nucleic acid-encoded drugs was conceived over two decades ago when Wolff et al.1 demonstrated that direct injection of in vitro transcribed (IVT) mRNA or plasmid DNA (pDNA) into the skeletal muscle of mice led to the expression of the encoded protein in the injected muscle. At the time, mRNA was not pursued further as it is less stable than DNA, and the field focused on technologies based on pDNA and viral DNA. Nevertheless, since its discovery in 1961, mRNA has been the subject of consistent basic and applied research for various diseases (Fig. 1 (TIMELINE)). In the first decades after its discovery, the focus was on understanding the structural and functional aspects of mRNA and its metabolism in the eukaryotic cell. This is in addition to making tools for mRNA recombinant engineering more accessible to a broader research community. In the 1990s, preclinical exploration of IVT mRNA was initiated for diverse applications, including protein substitution and vaccination approaches for cancer and infectious diseases2,3,4,5,6,7,8,9,10,11. Consequently, accumulated knowledge enabled recent scientific and technological advances to overcome some of the obstacles associated with mRNA, such as its short half-life and unfavourable immunogenicity.
CAR, chimeric antigen receptor; Cas9, CRISPR-associated protein 9; CRISPR, clustered regularly interspaced short palindromic repeat; DC, dendritic cell; dsRNA, double-stranded RNA; iPSC, induced pluripotent stem cell; RSV, respiratory syncytial virus; ssRNA, singlestranded RNA; TALEN, transcription activator-like effector nuclease; TLR, Toll-like receptor.
Conceptually, there are several important differences between IVT mRNA-based therapeutic approaches and other nucleic acid-based therapies. IVT mRNA does not need to enter into the nucleus to be functional; once it has reached the cytoplasm the mRNA is translated instantly. By contrast, DNA therapeutics need to access the nucleus to be transcribed into RNA, and their functionality depends on nuclear envelope breakdown during cell division. In addition, IVT mRNA-based therapeutics, unlike plasmid DNA and viral vectors, do not integrate into the genome and therefore do not pose the risk of insertional mutagenesis. For most pharmaceutical applications it is also advantageous that IVT mRNA is only transiently active and is completely degraded via physiological metabolic pathways. Moreover, the production of IVT mRNA is relatively simple and inexpensive, and so the development of IVT mRNA-based therapeutics has garnered broad interest (Box 1).
In the field of therapeutic cancer vaccination, IVT mRNA has undergone extensive preclinical investigation and has reached Phase III clinical testing12,13,14,15,16,17,18. In other areas such as protein-replacement therapies in oncology11,19,20,21, cardiology22,23, endocrinology24, haematology25,26, pulmonary medicine25,27 or the treatment of other diseases6,28, the development of IVT mRNA-based therapeutics is at the preclinical stage. To advance protein replacement therapies, unresolved issues such as the targeted delivery of mRNA and its complex pharmacology need to be addressed.
Here, we provide a comprehensive overview of the current state of mRNA-based drug technologies and their various applications. We discuss the peculiarities of the biopharmaceutical development of mRNA-based therapeutics, as well as the strengths and the key challenges that might affect the progress of this drug class.
Principal concept of mRNA pharmacology
The concept behind using IVT mRNA as a drug is the transfer of a defined genetic message into the cells of a patient for the ultimate purpose of preventing or altering a particular disease state.
In principle, two approaches of using IVT mRNA are being pursued. One is to transfer it into the patient's cells ex vivo; these transfected cells are then adoptively administered back to the patient. This method is being investigated for genome engineering, genetic reprogramming, T cell- and dendritic cell (DC)-based immunotherapies to treat cancer and infectious diseases, and some protein-replacement approaches. The second approach is direct delivery of the IVT mRNA using various routes. This is being developed for applications in oncology and infectious diseases, tolerization regimens to treat allergies and for other protein-replacement therapies. For both ex vivo transfection and direct vaccination, the following general principles of mRNA pharmacology apply.
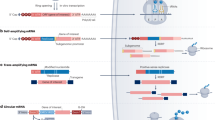
The machinery of the transfected cell is utilized for in vivo translation of the message to the corresponding protein, which is the pharmacologically active product. Thus, therapeutic protein-coding mRNA works in an opposite manner to small interfering RNA (siRNA), which inhibits the expression of proteins. IVT mRNA is engineered to structurally resemble naturally occurring mature and processed mRNA in the cytoplasm of eukaryotic cells. Hence, the IVT mRNA is single-stranded, has a 5′ cap and a 3′ poly(A) tail. The open reading frame (ORF) encoding the protein of interest is marked by start and stop codons and is flanked by untranslated regions (UTRs) (Fig. 2).
a | A linearized DNA plasmid template with the antigen-coding sequence is used for in vitro transcription. The in vitro transcribed mRNA contains the cap, 5′and 3′ untranslated regions (UTRs), the open reading frame (ORF) and the poly(A) tail, which determine the translational activity and stability of the mRNA molecule after its transfer into cells. b | Step 1: a fraction of exogenous mRNA escapes degradation by ubiquitous RNases and is spontaneously endocytosed by cell-specific mechanisms (for example, macropinocytosis in immature dendritic cells) and enters endosomal pathways. Step 2: release mechanisms of mRNA into the cytoplasm are not fully understood. Step 3: translation of mRNA uses the protein synthesis machinery of host cells. The rate-limiting step of mRNA translation is the binding of the eukaryotic translation initiation factor 4E (eIF4E) to the cap structure222,223. Binding of the mRNA to ribosomes, the eukaryotic initiation factors eIF4E and eIF4G, and poly(A)-binding protein, results in the formation of circular structures and active translation224. Step 4: termination of translation by degradation of mRNAs is catalysed by exonucleases225,226. The cap is hydrolysed by the scavenger decapping enzymes DCP1, DCP2 and DCPS32, followed by digestion of the residual mRNA by 5′–3′ exoribonuclease 1 (XRN1). Degradation may be delayed if the mRNA is silenced and resides in cytoplasmic processing bodies227. Alternatively, endonucleolytic cleavage of mRNA in the exosome may occur228,229,230. The catabolism of abberant mRNA (for example, mRNA with a premature stop codon) is controlled by various other mechanisms231. Step 5: the translated protein product undergoes post-translational modification, the nature of which depends on the properties of the host cell. The translated protein can then act in the cell in which it has been generated. Step 6: alternatively, the protein product is secreted and may act via autocrine, paracrine or endocrine mechanisms. Step 7: for immunotherapeutic use of mRNA, the protein product needs to be degraded into antigenic peptide epitopes. These peptide epitopes are loaded onto major histocompatibility complex (MHC) molecules, which ensure surface presentation of these antigens to immune effector cells. Cytoplasmic proteins are proteasomally degraded and routed to the endoplasmic reticulum where they are loaded on MHC class I molecules to be presented to CD8+ cytotoxic T lymphocytes. MHC class I molecules are expressed by almost all cells. Step 8: in antigen-presenting cells, to obtain cognate T cell help for a more potent and sustainable immune response, the protein product needs to be routed to MHC class II loading compartments. This can be accomplished by incorporating routing signal-encoding sequences into the mRNA. Moreover, exogenous antigens that are taken up by dendritic cells can also be processed and loaded onto MHC class I molecules by a mechanism that is known as cross-priming232. Step 9: protein-derived epitopes can then be presented on the cell surface by both MHC class I and MHC class II molecules.
The mRNA is synthesized in a cell-free system by in vitro transcription from a DNA template, such as a linearized plasmid or a PCR product. With the exception of the 5′ cap, this DNA template encodes all the structural elements of a functional mRNA. In vitro transcription is performed with T7 or SP6 RNA polymerase in the presence of nucleotides and thereafter the mRNA is capped enzymatically. The template DNA is then digested by DNases and the mRNA is purified by conventionally used methods for isolating nucleic acids.
The primary compartment of the pharmacodynamic activity of IVT mRNA is the cytoplasm. In contrast to natural mRNA that is produced in the nucleus and enters the cytoplasm through nuclear export, IVT mRNA has to enter the cytoplasm from the extracellular space.
Irrespective of whether the IVT mRNA is delivered to the cells in vitro or in vivo, two key factors determine its cytoplasmic bioavailability. One is rapid degradation by the highly active ubiquitous RNases, which are abundant in the extracellular space. The other is the cell membrane, which hampers the passive diffusion of the negatively charged large mRNA molecule into the cytoplasm. In principle, eukaryotic cells are capable of actively engulfing naked mRNA. However, in most cell types the uptake rate and cytoplasmic transfer is minimal (less than 1 in 10,000 molecules of the initial mRNA input). The transfection of cells can be improved by formulating the IVT mRNA with complexing agents, which protect the mRNA from degradation by RNases and also act as facilitators for its cellular uptake. Alternatively, techniques such as electroporation in RNase-free buffer can be used for efficient ex vivo mRNA transfer into cells.
Once IVT mRNA has entered the cytoplasm, its pharmacology is governed by the same complex cellular mechanisms that regulate the stability and translation of native mRNA.
The protein product translated from the IVT mRNA undergoes post-translational modification, and this protein is the bioactive compound. The half-lives of both the IVT mRNA template and the protein product are critical determinants of the pharmacokinetics of mRNA-based therapeutics.
Once the encoded protein has been generated, its destination is determined by signal peptides. These may be either intrinsic to the natural protein sequence or recombinantly engineered to direct the protein to the desired cell compartment within the host cell. Alternatively, the protein may be secreted to act on neighbouring cells or, if released into the bloodstream, to act on distant organs.
For immunotherapeutic approaches, the processing pathways of the encoded protein are crucial for determining its pharmacodynamics. Similar to the fate of endogenously generated protein, mRNA-encoded products are degraded by proteasomes and presented on major histocompatibility complex (MHC) class I molecules to CD8+ T cells. In general, intracellular proteins do not reach the MHC class II processing pathway to induce T helper cell responses. However, by introducing a secretion signal into the antigen-encoding sequence, T helper cell responses can be achieved as the secretion signal redirects the protein antigen to the extracellular space.
Improving the translation and stability of mRNA
The amount of IVT mRNA required for a therapeutic effect and the treatment duration depends on many factors. These include the intended biological function of the encoded protein and its mode of action, as well as the potency and the circulation half-life of the protein, which will vary by several orders of magnitude for different applications. Nanogram to microgram amounts of highly antigenic proteins may be sufficient for the efficient induction of immune responses in humans. By contrast, milligram or even gram amounts of proteins may be required for the delivery of systemically active growth factors, hormones or monoclonal antibodies. Through iterative optimization of the translational potency and intracellular stability of IVT mRNA, the protein amounts that can be generated per unit of mRNA have considerably increased.
Substantial efforts have been invested in modifying structural elements of the IVT mRNA — notably the 5′ cap, 5′- and 3′-UTRs, the coding region, and the poly(A) tail — to systematically improve its intracellular stability and translational efficiency (Fig. 3). These improvements ultimately lead to the production of significant levels of the encoded protein over a longer timeframe; from a range of a few minutes to longer than 1 week29,30,31. The range of opportunities available for the modulation of mRNA pharmacology is still not fully explored, and a deeper understanding of mRNA-binding factors and their binding sites is likely to open up further opportunities for engineering mRNA vectors with diverse pharmacokinetic properties.
a | Key structural elements of in vitro transcribed (IVT) mRNA and strategies for their modifications. b | Depending on which elements (for example, modification of caps, untranslated regions (UTRs) or poly(A) tails) are used alone or in combination, the duration and kinetic profile of expression of the protein product can be modulated and fine-tuned. eIF4E, eukaryotic translation initiation factor 4E; IRES, internal ribosome entry site; ORF, open reading frame.
The 5′ cap. Robust translation of mRNA requires a functional 5′ cap structure. Natural eukaryotic mRNA has a 7-methylguanosine (m7G) cap linked to the mRNA during the transcription process by a 5′-5′-triphosphate bridge (ppp) (m7GpppN structure). Binding of the 5′ cap to eukaryotic translation initiation factor 4E (EIF4E) is crucial for efficient translation, whereas its binding to the mRNA decapping enzymes DCP1, DCP2 or DCPS regulates mRNA decay32. One approach to cap IVT mRNA after its initial synthesis is to perform a second step with recombinant vaccinia virus-derived capping enzymes33. The resulting cap structure is identical to the most frequent naturally occurring eukaryotic cap structure. The other more commonly used approach is to add a synthetic cap analogue into the in vitro transcription reaction and perform capping and in vitro transcription in a single step. However, the main limitation of this approach is that the cap analogue and the GTP nucleotide required for in vitro transcription compete, resulting in some of the mRNA remaining uncapped and translationally inactive.
Early mRNA research was performed with IVT mRNAs generated with a m7GpppG cap analogue1,34, and most of the ongoing clinical trials still use this type of mRNA. However, a substantial proportion of the m7GpppG analogue is incorporated in reverse orientation into the mRNA and therefore not recognized by the translational machinery, resulting in lower translational activity. Hence, so-called anti-reverse cap analogues (ARCAs; m27,3′−OGpppG) were introduced35,36. ARCA-capped mRNAs exhibited superior translational efficiency in various cell types37,38. Recently, a phosphorothioate-containing ARCA cap analogue was developed39. This cap analogue confers resistance to decapping by DCP2, thus further extending the half-life of the mRNA40. Experiments in mice showed that IVT mRNA containing a phosphorothioate-modified cap induced potent immune responses against the encoded protein, and the responses were stronger than those induced by mRNAs with a control cap41. The impact of the cap analogues on the translation and stability of IVT mRNA appears to depend on the cell type and cell differentiation state39.
The poly(A) tail. The poly(A) tail regulates the stability and translational efficiency of mRNA in synergy with the 5′ cap, the internal ribosomal entry site and various other determinants42. IVT mRNA is tailed either by encoding the poly(A) stretch in the template vector from which it is transcribed or by a two-step reaction that extends the IVT mRNA enzymatically using recombinant poly(A)polymerase. Recombinant poly(A)polymerase enables the incorporation of modified nucleotides into the poly(A) tail to inhibit deadenylation by poly(A)-specific nucleases43. This approach has been explored for various nucleoside analogues, including cordycepin (3′-deoxyadenosine)44; however, cordycepin failed to increase the half-life of the mRNA38. This failure is most probably because cordycepin is a chain terminator and thus incorporates only at the ultimate 3′ position.
A limitation of enzymatic polyadenylation is that each RNA preparation consists of a mixture of RNA species differing in the length of the poly(A) tail. In vitro transcription of RNA from a DNA template, by contrast, yields RNA with a defined poly(A) tail length, and is therefore preferred, particularly if the mRNA is intended for clinical applications. Analyses in DCs demonstrated that the 3′ end of the poly(A) tail should not be masked by additional bases and that the optimal length of the poly(A) tail is between 120 and 150 nucleotides29,37.
5′- and 3′-UTRs. Another strategy to optimize the translation and stability of IVT mRNA in cells is to incorporate 5′- and 3′-UTRs containing regulatory sequence elements that have been identified to modulate the translation and stability of endogenous mRNA.
For example, many IVT mRNAs contain the 3′-UTRs of α- and β-globin mRNAs that harbour several sequence elements that increase the stability and translation of mRNA30,45. The stabilizing effect of human β-globin 3′-UTR sequences is further augmented by using two human β-globin 3′-UTRs arranged in a head-to-tail orientation29. In addition, various regions of cellular and viral 5′- and 3′-UTRs enhance the stability and translational efficiency of mRNA. The 3′-UTR of the eukaryotic elongation factor 1α (EEF1A1) mRNA46 and a 5′-UTR element present in many orthopoxvirus mRNAs, for example, inhibited both decapping and 3′–5′ exonucleolytic degradation47 (reviewed in Ref. 48). For some applications, destabilizing the mRNA might be desirable to limit the duration of protein production. This effect can be achieved by incorporating AU-rich elements into 3′-UTRs, thus ensuring rapid mRNA degradation and a short duration of protein expression49.
The coding region. Codon composition is known to affect translation efficiency. Replacing rare codons with synonymous frequent codons improves translational yield50 because reuse of the same tRNA accelerates translation owing to amino-acylation of tRNAs in the vicinity of the ribosomes51. Codon context (that is, neighbouring nucleotides and codons) also affects the translational elongation rate and translational efficiency52. Similar to recombinant DNA-based approaches (reviewed in Ref. 53), codon-optimized IVT mRNAs have been successfully used in vaccine studies against viral infections54 and for the expression of non-viral proteins26.
However, there may be valid reasons to refrain from using optimized codons. Some proteins require slow translation, which is ensured by rare codons, for their proper folding55. It may also be beneficial for some IVT mRNA-encoded vaccines to maintain the original ORF. Potent cryptic T cell epitopes may be generated when the IVT mRNA is translated in different frames owing to ribosomal frame-shifting or when translation is initiated either internally or from a CUG start codon56,57,58. Codon optimization should eliminate these important sources of antigenic peptides.
Immune-stimulatory activity of IVT mRNA
For vaccination, the strong immune-stimulatory effect and intrinsic adjuvant activity of IVT mRNA are added benefits59,60 and lead to potent antigen-specific cellular and humoral immune responses9,61. The type of immune response appears to depend on a range of factors, including the characteristics of the type and size of particles in which the IVT mRNA is incorporated62,63,64. In applications such as protein-replacement therapies, however, activation of the innate immune system by IVT mRNA is a major disadvantage. The recent progress in identifying RNA sensors in cells and the structural elements within mRNA that are involved in immune recognition provides opportunities to augment immune activation by IVT mRNA, or alternatively, to create 'de-immunized' mRNA as needed.
IVT mRNA induces immune stimulation by activating pattern recognition receptors, the natural role of which is to identify and respond to viral RNAs by inducing various downstream effects (Fig. 4).
In vitro transcribed (IVT) mRNA is recognized by various endosomal innate immune receptors (Toll-like receptor 3 (TLR3), TLR7 and TLR8) and cytoplasmic innate immune receptors (protein kinase RNA-activated (PKR), retinoic acid-inducible gene I protein (RIG-I), melanoma differentiation-associated protein 5 (MDA5) and 2′–5′-oligoadenylate synthase (OAS)). Signalling through these different pathways results in inflammation associated with type 1 interferon (IFN), tumour necrosis factor (TNF), interleukin-6 (IL-6), IL-12 and the activation of cascades of transcriptional programmes. Overall, these create a pro-inflammatory microenvironment poised for inducing specific immune responses. Moreover, downstream effects such as slow-down of translation by eukaryotic translation initiation factor 2α (eIF2α) phosphorylation, enhanced RNA degradation by ribonuclease L (RNASEL) overexpression and inhibition of replication of self-amplifying mRNA are of relevance for the pharmacokinetics and pharmacodynamics of the IVT mRNA. IRF, interferon regulatory factor; ISRE7, interferon-stimulated response element; MAVS, mitochondrial antiviral signalling protein; MDA5, melanoma differentiation-associated protein 5; MYD88, myeloid differentiation primary response protein 88; MX1, myxovirus (influenza) resistance 1; NF-κB, nuclear factor-κB; TRIF, Toll-IL-1 receptor domain-containing adapter protein inducing IFNβ.
In immune cells, the Toll-like receptors TLR3, TLR7 and TLR8, which reside in the endosomal compartment, are activated by endocytosed IVT mRNA and induce secretion of interferon. TLR3 recognizes double-stranded RNA (dsRNA)65, whereas TLR7 and TLR8 sense single-stranded RNA (ssRNA)66,67. Poly(U) is the most potent interferon inducer, and acts through TLR7 (Refs 67,68). By contrast, in non-immune cells, most of the interferon production is induced through the activation of the cytosolic receptors cytoplasmic retinoic acid-inducible gene I protein (RIG-I; also known as DDX58) and melanoma differentiation-associated protein 5 (MDA5; also known as IFIH1)69,70. RIG-1 is activated by short, double-stranded, 5′-triphosphate RNA71, whereas MDA5 responds to long dsRNA and viral mRNAs lacking 2′-O-methylation72,73.
Cytoplasmic RNA sensors mediate immune stimulation and affect the efficiency of mRNA translation, which eventually leads to stalled translation, RNA degradation and direct antiviral activity (Fig. 4). These effects are partly mediated by protein kinase RNA-activated (PKR; also known as EIF2AK2), which phosphorylates the eukaryotic translation initiation factor 2α (eIF2α) and ultimately inhibits mRNA translation74. De-immunized IVT mRNA can be created by incorporating naturally occurring modified nucleosides such as pseudouridine, 2-thiouridine, 5-methyluridine, 5-methylcytidine or N6-methyladenosine into the IVT mRNA; this has been shown to suppress both the intrinsic adjuvant activity of IVT mRNA as well as its inhibitory effects on translation75,245. Such modified IVT mRNAs appear to avoid the activation of TLR7 and TLR8 (Ref. 75), and some of them, including pseudouridine and 2-thiouridine, were shown to make IVT mRNA undetectable by RIG-I and PKR76,77,78. When dsRNA contaminants (and thus any remaining TLR3 activation) were eliminated through high-performance liquid chromatography purification, the modified mRNA no longer induced any immune-stimulatory effect63. Superior translation of pseudouridine-modified mRNA has been attributed to its increased stability and reduced binding to PKR78,185.
Progress in improving mRNA delivery
Many cell types can spontaneously take up naked mRNA. Naked mRNA internalized by scavenger-receptor-mediated endocytosis accumulates in the lysosome, from which only small amounts of mRNA leak into the cytoplasm79. The mechanisms of mRNA release into the cytoplasm are not well characterized and may differ between cell types. Moreover, mRNA is exchanged via exosomes between cells80. In most cells, active uptake of mRNA is inefficient and saturated at low mRNA doses. Immature DCs, which are specialized to constantly engulf extracellular fluid while sampling their environment, are an exception because they take up mRNA by macropinocytosis and thereby accumulate it with high efficiency in a linear non-saturable manner in concentration ranges over several orders of magnitudes81. As a consequence, for delivery into most cell types, suitable formulations are required to protect IVT mRNA against extracellular RNase-mediated degradation and facilitate its entry into cells.
There are two challenges associated with the delivery of IVT mRNA: one is to achieve a sufficiently high net level of the encoded protein and the other is to reach a high number of cells.
Most physiological and pathological processes, such as cell migration, cell growth, wound healing, inflammation and angiogenesis, are controlled by paracrine signalling. Accordingly, many intended clinical applications involve signalling molecules such as chemokines, cytokines and growth factors, which are secreted and exert their biological function in a paracrine manner. In these cases, the amount of total protein is crucial for their biological effect. IVT mRNA may be delivered to any cell type that is accessible via the bloodstream or other routes, as long as sufficient doses of the encoded protein are released and reach their target cells. For such applications, transfection with the synthetic mRNA is a suitable approach. For potent protein hormones such as erythropoietin, systemic delivery of mRNA appears to result in adequate plasma levels, as shown by preclinical studies in mice and primates26. Liver cells are accessible to many types of polymer and liposomal delivery platforms, so targeting the liver may be one approach to produce high amounts of recombinant protein.
The challenge is different if deficient or defective intracellular proteins are to be substituted through IVT mRNA. Here, the proportion of cells that are restored by IVT mRNA transfer has to be significant enough to have a biological impact. For such applications, the proportion of transfected target cells rather than the absolute protein dose is crucial. If cells are transfected ex vivo, transfection efficiencies of up to 80% of cells are achievable for most cell types. In vivo delivery of mRNA, by contrast, into a high fraction of defined target cell populations is challenging and depends on the accessibility of target cells.
In vitro transfection strategies. In vitro transfection strategies to obtain cells transfected with IVT mRNA for adoptive transfer have benefited from the development of formulations to protect mRNA against RNase-mediated degradation and to facilitate its entry into cells. In addition to various positively charged lipids, other transfection agents containing polymers, such as polyethylenimine, cationic polypeptides and dendrimers (reviewed in Ref. 82), are commercially available. The number of mRNA transfection kits for in vitro and in vivo mRNA delivery is rapidly growing. Clinical applications using ex vivo IVT mRNA-transfected cells have benefited from these advances.
Electroporation, which was first applied to gene transfer in 1982 (Ref. 83), is now established as a favoured method for in vitro mRNA transfection of haematopoietic cell types84. Immunotherapy with DCs electroporated with IVT or autologous tumour-derived mRNA was shown to be safe in patients with cancer15,18,85,86,87,88,89,90. In Phase IB trials, patients with advanced stage III and IV melanoma generated strong CD4+ and CD8+ immune responses against the antigens encoded by the IVT mRNAs18,117. In a Phase I trial, 19 of 20 patients with metastatic prostate cancer also responded to the treatment by developing antigen-specific CD8+ T cells15. The development of novel devices for gentle electroporation of large numbers of cells under sterile conditions has enabled the development of a rapid clinical-grade protocol for a broad range of IVT mRNA-based cell therapy applications91.
In vivo transfection strategies. Clinical trials in which naked or protamine-complexed mRNA vaccines are delivered either intradermally16,17,92 (ClinicalTrials.gov identifiers: NCT01915524 and NCT01817738) or injected directly into lymph nodes (ClinicalTrials.gov identifier: NCT01684241) are ongoing. However, although the delivery of naked IVT mRNA appears to be sufficient to induce potent immune responses, it may not be sufficient for other clinical applications in which a substantially higher amount of protein is required and cell types other than DCs are targeted.
For systemic delivery, nucleic acid-based drugs are administered as nanosized drug formulations and reach organs via either fenestrated or non-fenestrated capillaries. Cells in the bloodstream, cells of the liver and reticuloendothelial cells in the spleen and bone marrow are accessible for intravenously delivered nanosized carriers. Systemic delivery of IVT mRNA to other tissues, however, is hindered by non-fenestrated, non-permeable vascular endothelia, intercellular junctions and dense extracellular fibril networks that limit the accessibility of target cells93 (Fig. 5). The extensive but moderately successful efforts and difficulties experienced with siRNA and pDNA deliveries are predictive of what to expect for similar efforts in the mRNA field. However, it is important to consider the conceptual differences among these three platforms. In particular, the lessons learnt and improvements made in the delivery of siRNA (for example, clinically validated lipid nanoparticle formulations with excellent safety and efficacy profiles)94,95 and pDNA need to be considered when deciding whether these improvements can be similarly adopted to advance mRNA delivery or whether novel mRNA-tailored approaches have to be explored.
All three nucleic acid-based drug modalities are applied as nanosized drug formulations for systemic delivery and reach organs via capillary systems with either non-fenestrated (a) or fenestrated (b) capillaries. The primary pharmacological effect of small interfering RNA (siRNA), namely the deletion of a defined protein function in situ, is restricted to those very cells it has entered. siRNA cannot act in cells that are not directly accessed owing to biological barriers such as non-fenestrated capillaries. In tissues with endothelial fenestration, siRNA may reach a few tissue layers adjacent to capillaries. Plasmid DNA (pDNA) is only incorporated and active in those cells undergoing mitosis at the time of exposure. This impairs its use for tissues with non-fenestrated capillaries and restricts the number of transfectable cells in tissues with endothelial fenestration to those undergoing mitosis at the time of exposure. In contrast to pDNA, mRNA enters and acts in endothelial cells of non-fenestrated tissues, and in fenestrated tissues it reaches both mitotic and non-mitotic cells in cell layers adjacent to the capillaries233. Non-target cells, such as vascular endothelial cells transfected with mRNA or pDNA, can express pharmacologically active proteins and, via paracrine secretion, can reach target cells that are located behind the mRNA delivery barriers234 (obviously siRNA cannot rely on such a function). Proteins produced in transfected cells are able to diffuse after secretion into the target tissue and mediate the intended biological effects via paracrine activity on adjacent cell populations. Such paracrine activity may be of particular value in tissues that have non-fenestrated capillaries.
It is particularly challenging to target cells such as neurons and myocardial or skeletal muscle cells, as they are not directly accessible to nanosized carriers delivered by systemic routes. Therefore, different routes of delivery are being tested for therapeutic applications of IVT mRNA in vivo, such as intramuscular, intradermal, intranodal, subcutaneous, intravenous, intratracheal and intrathecal delivery4,7,9,25,26,96,97,98,99,100,101,102. For drug delivery to the lung, IVT mRNA is either administered as aerosol or by intravenous targeting of the lung vasculature27,103. Cells in the central nervous system may be reached via intrathecal injection. For effective cancer targeting, the enhanced permeability and retention effect may be exploited, which is a unique phenomenon of solid tumours related to their anatomical and pathophysiological differences from normal tissues21,104.
Preclinical and clinical applications
The progress in mRNA technology motivated the exploration of IVT mRNA for a broad range of therapeutic applications in numerous preclinical studies and a smaller number of clinical trials (Fig. 6; Table 1). The three major therapeutic areas for which mRNA drugs are currently being explored are immunotherapeutics, protein-replacement therapies and regenerative medicine applications.
The therapeutic applications of in vitro transcribed (IVT) mRNA are summarized in detail in Table 1. The solid arrows pointing in the right hand column denote applications that are in the clinic, whereas stippled arrows refer to preclinical applications. Cas9, CRISPR-associated protein 9; CRISPR, clustered regularly interspaced short palindromic repeat; EPO, erythropoietin; FOXP3, forkhead box P3; IL-10, interleukin-10; MSC, mesenchymal stem cell; RSV, respiratory syncytial virus; SPB, surfactant protein B; TALEN, transcription activator-like effector nuclease; VEGFA, vascular endothelial growth factor A; ZNF, zinc finger nuclease.
Cancer immunotherapy. Cancer immunotherapy is the field in which mRNA-based technologies have the longest history of systematic exploration in integrated preclinical and clinical programmes.
In 1995, it was demonstrated that intramuscular injection of naked RNA encoding carcinoembryonic antigen elicited antigen-specific antibody responses4. A year later, it was demonstrated that DCs exposed to mRNA coding for specific antigens or to total mRNA extracted from tumour cells and subcutaneously administered into tumour-bearing mice induced T cell immune responses and inhibited the growth of established tumours5. These findings, together with the increasing availability of vaccine targets owing to cloning of novel tumour antigens105,106, accelerated the development and clinical translation of the mRNA-transfected DC approach12,107,108,109. Since then, numerous clinical trials using vaccines based on ex vivo IVT mRNA-transfected DCs were performed in patients with cancer and thus established the feasibility and safety of this approach (reviewed in Refs 110,111).
Treatment protocols with IVT mRNA-transfected DCs were further refined, for example, by optimizing ex vivo maturation of DCs or by co-delivering IVT mRNA that encoded immune response modulators such as cytokines and co-stimulators18,112,113,114,115,116, thus enhancing antitumoural activity in patients87,117. Meanwhile, Argos Therapeutics has initiated a Phase III clinical trial using DCs loaded with autologous tumour-derived amplified mRNA in patients with advanced renal cell cancer; data from this study are anticipated in 2015 (ClinicalTrials.gov identifier: NCT01582672).
Cell therapies are costly and complex. Therefore, direct in vivo administration of IVT mRNA that encodes the tumour antigen has been revisited by various groups. Different delivery strategies, such as cationic liposomes and bombardment using gene guns, were tested with varying success (reviewed in Ref. 118).
For example, clinical trials of intradermal delivery of naked or protamine-complexed, sequence-optimized IVT mRNA demonstrated that expression of the encoded antigen by skin cells occurred, and that this led to the induction of T cell and antibody responses9. In addition, intradermal application of naked mRNA alone led to the induction of a T helper 2 (TH2)-type antigen-specific immune response. By contrast, a strong shift towards a TH1-type response was accomplished by co-delivering adjuvants such as granulocyte–macrophage colony-stimulating factor (GM-CSF)119 or complexing the IVT mRNA with protamine120. Early clinical trials with protamine-complexed IVT mRNA as well as mRNA combined with GM-CSF revealed that intradermal vaccination with these compounds is feasible, safe and can lead to the induction of antigen-specific antibody and T cell immune responses17,92. This approach was further optimized, resulting in a pharmaceutical-grade two-component vaccine combining a naked antigen-encoding mRNA plus a protamine-complexed mRNA adjuvant (developed by CureVac). This vaccine showed potent activity in preclinical studies62 and is currently being tested in ongoing clinical trials in patients with prostate cancer (ClinicalTrials.gov identifiers: NCT00831467 and NCT01817738) and non-small-cell lung cancer (ClinicalTrials.gov identifiers: NCT00923312 and NCT01915524)121.
In another strategy, the IVT mRNA was optimized for in situ transfection of DCs in vivo. The objectives of this approach were to improve the translation and stability of the IVT mRNA and to enhance the presentation of the mRNA-encoded antigen on MHC class I and II molecules of murine and human DCs29,41,122,123,124. Direct injection of naked IVT mRNA into lymph nodes was identified as the most effective route to induce potent T cell responses. The IVT mRNA was selectively and effectively taken up by lymph node-resident DCs through macropinocytosis and mediated their maturation via TLR7 signalling. Presentation of the IVT mRNA-encoded antigen and the immunostimulatory intralymphatic milieu induced strong antigen-specific T cell responses of the TH1 pro-inflammatory type and profound antitumour immunity in animal models81,97,98,125. Immune responses of DCs following delivery of IVT mRNA can be further potentiated by either co-administering the DC-activating ligand FMS-related tyrosine kinase 3 (FLT3) as a recombinant protein98 or co-transfecting DCs with the so-called TriMix, consisting of IVT mRNAs encoding the immunomodulators CD40L, CD70 and truncated, constitutively active TLR4 (Ref. 99). First-in-human testing of the intranodal injection of naked IVT mRNA encoding cancer antigens (developed by BioNTech) has recently begun in patients with melanoma (ClinicalTrials.gov identifier: NCT01684241).
Personalization of cancer immunotherapy may be facilitated by in vivo administered mRNA technology owing to its versatility, robustness and relatively low cost126,127,128. The clinical testing of the first individualized vaccine for treating patients with cancer has just been initiated (ClinicalTrials.gov identifier: NCT02035956). Tumour specimens from each enrolled patient are subjected to next-generation sequencing and individual immunogenic mutations are selected to construct a personalized IVT mRNA vaccine encoding a polypeptide that consists of aligned epitopes with individual mutations. Thus, IVT mRNA may become the first drug that is engineered according to personal genome information.
In addition to active immunization and immunomodulation, IVT mRNA is being investigated as a multipurpose tool for the transient modulation of immune cells. For example, IVT mRNA encoding tumour antigen-specific T cell receptor (TCR) or chimeric antigen receptor (CAR) has been transfected into T cells or natural killer cells ex vivo by electroporation. Transfected cells carrying such mRNA-encoded receptors are able to recognize and kill tumour cells that express the target antigen. The transient nature of mRNA reduces the risk of unwanted side effects by the uncontrolled expansion of adoptively transferred immune cells. IVT mRNAs encoding various antigen-specific receptors have been evaluated and their antitumour activity was demonstrated in animal models129,130,131,132. Recently, cell therapy using T cells electroporated with IVT mRNA encoding CARs entered clinical testing (ClinicalTrials.gov identifier: NCT01897415).
Vaccines against infectious diseases. In 1993, it was demonstrated that liposome-entrapped IVT mRNA encoding an influenza nucleoprotein induced a virus-specific T cell response in mice3. Recently, intramuscularly delivered, self-amplifying IVT RNA formulated with synthetic lipid nanoparticles was shown to induce protective antibody responses against respiratory syncytial virus (RSV) and influenza virus in mice101,133.
Currently, three types of IVT mRNA-based vaccine approaches for infectious diseases have entered pharmaceutical development.
For the treatment of HIV infections, patients under highly active antiretroviral therapy were immunized with DCs transfected with IVT mRNA encoding HIV proteins. Three Phase I/II clinical trials using IVT mRNA encoding combinations of different viral proteins showed that the vaccines are safe and that antigen-specific CD8+ and CD4+ T cell responses can be induced115,134,135. In one of these studies, increased HIV inhibition by antigen-specific CD8+ T cells was documented ex vivo; however, no antiviral effects were observed in the clinical trial135.
Two different strategies using IVT mRNA as prophylactic influenza vaccines are currently undergoing preclinical investigation. The first is based on an intradermally injected two-component vaccine containing an mRNA adjuvant and naked IVT mRNA encoding influenza haemagglutinin antigen alone or in combination with neuraminidase-encoding IVT mRNA. Both regimens induced protective immune responses against the corresponding influenza strains in aged and newborn mice, as well as long-lasting protective immunity in ferrets and pigs100.
The second strategy uses self-amplifying IVT mRNA containing sequences of positive-stranded RNA viruses (Box 2). Initially, this strategy was developed for a flavivirus model and protective immunity against flavivirus infection was achievable with intradermal delivery of less than 1 ng of IVT genomic mRNA, which corresponded to the attenuated virus7. Subsequent studies of RNA-based vaccines for infectious diseases focused on recombinant RNA replicon systems derived from the alphavirus family136,137 (reviewed in Refs 138,139). RNA replicon vectors in which genes encoding structural proteins were replaced by genes encoding viral antigens have been used to elicit protective antibody responses in animal models of flavivirus, RSV, influenza and parainfluenza virus infection140,141,142.
Vaccines to alleviate allergy. Antigen-specific immunotherapy is the only treatment modality for immunoglobulin E (IgE)-mediated type I allergic diseases. Modulating the type of T cell response and inducing IgG antibodies that compete with IgE antibodies for their binding sites on allergens are the primary modes of action of an effective immunotherapy143.
The molecular identification of the most common hypersensitivity target antigens set the stage for recombinant vaccine approaches. In preclinical models, DNA-based genetic vaccines were shown to antagonize allergy mechanisms by inducing TH1-type T cell immune responses that suppressed allergen-specific IgE production144,145. However, the clinical translation of DNA-based allergy vaccines is hampered by safety considerations. It was shown that the injected DNA persisted for 2 weeks and could spread from the injection site to immune and non-immune tissues throughout the body, thus posing the potential risk of inducing severe anaphylactic side effects146. In this respect, IVT mRNA-based approaches may be advantageous because IVT mRNA undergoes rapid degradation in the extracellular space and can be engineered to have a short intracellular half-life. Combined with the strong TH1 immunostimulatory capacity of mRNA, it may be better suited than DNA for the development as an allergy vaccine. In a murine model of allergic rhinitis, intradermal injection of IVT mRNA before antigen sensitization induced long-lasting allergen-specific TH1-type immune responses147,148. These responses protected the mice against the induction of allergen-specific IgE and inhibited lung inflammation mediated by allergen exposure147,148.
Protein-replacement therapies. The supplementation of proteins that are not expressed or are not functional, as well as the substitution of foreign proteins that activate or inhibit cellular pathways (for example, therapeutic antibodies), are the most obvious applications for IVT mRNA-based drugs. Several diseases are being studied in which the malfunctioning proteins are being replaced by the in vivo production of the therapeutic intracellular and secreted proteins from transfected IVT mRNA. All such endeavours are at the preclinical stages of development.
The first preclinical application of an IVT mRNA for replacing a deficient, physiologically relevant protein was reported in 1992 and it remained the only such work for almost two decades2. The discovery that modified nucleosides can reduce the immune-stimulatory activity of RNA was pivotal in advancing this field of application75. Preclinical experiments showed that the use of nucleoside-modified IVT mRNA together with improved mRNA purification protocols eliminates immune activation by mRNA and increases its translation63, thereby opening the therapeutic application of IVT mRNA for the field of protein replacement.
IVT mRNA containing modified nucleosides (2-thiouridine and 5-methylcytidine) and encoding surfactant protein B (SPB) was tested in a mouse model of congenital lethal lung disease caused by the lack of the SPB protein. Aerosol delivery of SPB mRNA into the lung twice a week protected mice from respiratory failure and prolonged their average lifespan25. In experiments with mice and macaques, pseudouridine-modified IVT mRNA purified by high-performance liquid chromatography and encoding erythropoietin was administered, and therapeutically relevant levels of erythropoietin were detected26. In an asthma disease model, intratracheal delivery of a nucleoside-modified mRNA encoding the regulatory T cell transcription factor forkhead box protein P3 (FOXP3) rebalanced pulmonary TH cell responses and protected the mice from allergen-induced tissue inflammation and airway hyperresponsiveness27. It was also demonstrated that direct intramyocardial injection of IVT mRNA containing pseudouridine and 5-methylcytidine, and encoding vascular endothelial growth factor A (VEGFA), improved heart function and enhanced long-term survival in a mouse model of myocardial infarction23. In another study, mouse mesenchymal stem cells were transfected ex vivo with pseudouridine-containing IVT mRNA encoding the immunosuppressive cytokine interleukin-10 (IL-10) and the tissue homing factors P-selectin glycoprotein ligand 1 (PSGL1) and Sialyl-Lewis(x) (SLeX). Upon re-injection, these cells homed to inflamed tissues and promoted rapid healing28.
Despite these achievements, the development of IVT mRNA drugs for the purpose of protein replacement is still associated with technical challenges. For IVT mRNA-based protein delivery, cell type-specific differences in post-translational modification have to be considered. For example, for glycoproteins, the composition of the glycoconjugate is not encoded in the mRNA and depends on the tissue type in which the protein is generated. Not every cell has the capability to glycosylate each protein properly, particularly if highly complex glycosylation is required149. Another type of post-translational modification is proteolytic processing. Processing by endoproteases is an integral part of the maturation of various functional polypeptides, including growth factors, cytokines, receptors, neuropeptides, enzymes, hormones and plasma proteins150. Other proteins require well-defined cleavage by protein convertases to their biologically active form, which occurs intracellularly in the Golgi apparatus and the secretory granules151. Several convertase subtypes have been identified with different specificities and tissue distribution152. Cells that are transfected with IVT mRNA need to have the required convertase or endoprotease to process the encoded precursors to functional products.
When secreted proteins are expressed in heterologous tissues, their secretory signal peptides may be poorly recognized and most of the protein may remain within the cells. The relative secretory signal strengths differ153; thus, exchanging the natural signal peptides could lead to increased protein secretion. For example, in an animal model of plasmid-mediated expression of erythropoietin in muscles, significantly more erythropoietin was secreted when the natural signal peptide of erythropoietin was replaced with that of tissue plasminogen activator154. To achieve maximal effect, IVT mRNA should ideally be transfected into cells that naturally secrete the encoded protein, otherwise signal peptide optimization might be required155.
It is of considerable interest to utilize IVT mRNA for a broad range of protein-replacement applications, including those that are currently being addressed with recombinant proteins as well as those for which recombinant protein technology cannot be used. Given the diversity of proteins that may be potential candidates for the IVT mRNA approach and are currently being explored as such, it is difficult to predict which of these will be the first to be advanced into clinical development. The developmental hurdles may be lower for proteins with a broad therapeutic window, activity at low doses and for which there is already an established pharmacokinetic and pharmacodynamic understanding in humans. It may also be easier to develop IVT mRNA-based protein replacement approaches for proteins that are expressed in easily accessible organs such as the liver, as well as for proteins that are fully inactive in the respective patient population and therefore the substituted counterpart can be instantaneously and easily quantified for better control of its bioavailability.
Reprogramming of cell fates. Cell phenotype and function can be modulated in vitro using nucleoside-modified IVT mRNA. In 2010, it was demonstrated that IVT mRNA containing pseudouridine and 5-methylcytydine and encoding the Yamanaka stem cell factors156 could be used as a safe strategy for efficiently reprogramming cells to pluripotency without leaving residual traces of transgenes157. The IVT mRNA was not only used to induce pluripotency but also to differentiate induced pluripotent stem cells (iPSCs). The introduction of nucleoside-modified IVT mRNA encoding myoblast determination protein into iPSCs resulted in their direct differentiation into myocyte-like cells.
Since then, several variants of the original approach have been described, which claim more effective induction of either pluripotent stem cells or cell fate conversion158,159,204 (reviewed in Ref. 160). The reprogramming and direct differentiation of cells with IVT mRNA profits from its high in vitro transfection efficiency, transient expression with no genomic integration and the ability to transfer complex mixtures. The lack of residual expression of transgenes in IVT mRNA-induced iPSCs not only facilitates their utilization for disease modelling161 and toxicology testing162 but also forms the basis for their application in regenerative medicine163. Similarly, IVT mRNA transfer may be utilized to generate differentiated cells of clinical value.
Genome editing with IVT mRNA-encoded engineered nucleases. In the past decade, genome editing has emerged as a potential alternative to gene therapy. Custom-designed zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs) and CRISPR–Cas9 (clustered regularly spaced short palindromic repeat–CRISPR-associated protein 9) provide powerful tools for site-specific modification of genomes164,165,166. These approaches, however, have the risk of nonspecific editing. A prolonged presence of editing enzymes translated from DNA-based vectors resulted in off-target effects167. As the nucleases are only required for a short duration, their transient expression from IVT mRNA would minimize this nonspecific effect. IVT mRNAs encoding ZFNs, TALENs and Cas9 have been applied successfully to edit genomes by disrupting or integrating sequences ex vivo in embryonic cells from different species (for example, mice, rats and rabbits) and in vivo in rodents168,169,170,171,172,173,174,175. Most recently, cynomolgus monkeys with site-specific gene modifications were generated by injecting IVT mRNA encoding Cas9 at the one-cell embryonic stage, thus opening the opportunity to create primate models of human diseases176.
Transposase-encoding IVT mRNA has also been utilized for transposon-mediated stable gene transfer both in vitro and in vivo. For example, the expression of transposases of the Sleeping Beauty, piggyBac or Tol2 transposon systems by injection of their mRNAs resulted in stable genomic transposition in mammalian cells177 and in vivo in rodents178,179,180,181.
The use of IVT mRNA rather than plasmids for the expression of the transposases increased the survival rate of the injected cells because injection in the cytoplasm is more gentle than pronuclear injection179. By narrowing the duration of peak translation, the biosafety of the approach is increased because the probability of remobilization of the transgenes is reduced182.
Clinical translation of IVT mRNA
By relying on the patient's body to make the desired protein, IVT mRNA drugs provide an approach in which the robust and tunable production of a therapeutic protein is possible, bypassing the need for costly manufacturing of proteins in fermentation tanks. Associated with these unique features is the vision that utilizing IVT mRNA will help address challenges in newly emerging technologies such as targeted genome engineering, generation and reprograming of stem cells as well as production of on-demand personalized vaccines.
To enable the full potential of mRNA as a therapeutic modality to be realized, regulatory (Box 3) and scientific issues concerning clinical and product development require diligent consideration.
So far, clinical experience of IVT mRNA drugs is limited to immunotherapeutic applications. Of the clinical programmes in the field of vaccine development with IVT mRNA alone or IVT mRNA-transfected DCs, few are advanced enough to provide a sufficiently broad knowledge base for other applications. For each application, the well-established systematic exploration of the variables of treatment protocols, such as dosing, treatment schedule and route of administration, have to be delineated to identify the appropriate regimen.
Common objectives of early clinical trials are to explore pharmacokinetic characteristics of the drug and to conduct dose finding. However, the pharmacology of mRNA drugs is complex because the IVT mRNA is not the final pharmacologically active agent. So far, it has not been fully investigated whether the bioavailability of the protein it encodes can be robustly and precisely controlled under clinical conditions, which are particularly challenging because of high inter- and intra-individual variability. Accompanying medication also requires consideration, particularly when IVT mRNA therapies are combined with other drugs that affect mRNA metabolism and translation, such as certain antibiotics and anticancer drugs.
Other key challenges related to the complex pharmacology of mRNA, and to its delivery in particular, concern applications in which precise targeting of a particular cell type and organ is required.
Safety considerations
Clinical experience with IVT mRNA for immunotherapy has demonstrated an excellent tolerability and safety profile and showed that mRNA drugs have no platform-inherent major risks.
However, for most of the other applications of mRNA-based therapeutics, including mRNA-based protein-replacement therapies, there is no clinical experience yet, leaving developers and regulators uncertain about the nature and frequency of safety problems that might occur.
Various safety risks associated with other drug classes do not apply to mRNA-based therapeutics (Box 3). IVT mRNA manufacturing is relatively simple and the manufacturing process as well as product quality is uniform and easy to control (Box 4). With no cellular and animal components involved, process-related risks are considerably lower for IVT mRNA as compared to recombinant proteins.
Nevertheless, various risk factors have to be considered.
IVT mRNA-mediated activation of immune mechanisms. The immune-activating property of IVT mRNA is an important feature to be considered from a safety perspective, particularly for systemically administered IVT mRNA. The underlying mechanisms are being extensively investigated. As discussed above, several signalling receptors of the innate immune system, including TLR3, TLR7 and TLR8, have been shown to mediate mRNA-induced immune activation and cytokine secretion65,66,67,183,184. In preclinical studies, interferon-α, IL-6, tumour necrosis factor-α and interferon-γ-induced protein 10 (IP-10; also known as CXCL10) were determined as key cytokines that are upregulated by systemic IVT mRNA delivery. The immune activation and profiles of secreted cytokines depend on the formulation of the IVT mRNA, including the particle size64.
Safety studies in animals are desirable but may not be fully conclusive owing to species-specific differences. To complement animal studies, IVT mRNA formulations should be tested in vitro for their pharmacodynamic effects on human leukocytes. As immune activation is dose-dependent, conservative dose-escalation protocols with low starting doses and close monitoring of patients are advised. Future studies will show whether nucleoside-modified IVT mRNA will avoid the activation of human TLRs in the clinical setting.
For applications of IVT mRNA as vaccines, transient immune activation is desirable. However, it is important to dissect the exact nature of the immune-modifying effect of each individual mRNA drug as part of the clinical research programme and to assess whether it is indeed desired. For example, induction of interferon-α, which slows down the translation machinery, should be avoided186.
The current data do not indicate that there is any induction of immunogenicity against IVT mRNA itself. However, mounting evidence suggests that patients with systemic lupus erythematosus and other autoimmune diseases can develop anti-self RNA autoantibodies that have a role in the induction and progression of autoimmunity187. Thus, under certain circumstances, such as long-term repetitive systemic application of mRNAs, anti-RNA antibodies may potentially form and mediate immune pathology. One might consider screening mRNA sequences to avoid conformations that are prone to inducing mRNA-specific antibodies188. Clinical monitoring of autoimmune phenomena and laboratory tests for antinuclear antibodies are therefore advised.
Immunogenicity of the IVT mRNA-encoded proteins. For recombinant proteins it is well established that unintended immunogenicity may result in adverse events such as anaphylaxis, cytokine release syndrome and infusion reactions. Moreover, immune responses may neutralize the biological activity of the protein drug as well as the endogenous protein counterpart189,190,191. A prominent example is the induction of neutralizing antibodies to therapeutic erythropoietin that caused red cell aplasia in monkeys and humans by crossreacting with endogenous erythropoietin192,193,194.
In principle, antiprotein antibodies can develop against proteins expressed from any IVT mRNA, in particular if repeat administration regimens are pursued.
However, in contrast to recombinant protein drugs, in vivo-generated protein therapeutics are autologous, produced in human cells and are likely to undergo the correct post-translational modifications and folding. Moreover, risk factors for immunogenicity related to the protein manufacturing process, such as protein aggregation or impurities derived from cells or medium in which the protein was produced, do not occur with IVT mRNA.
Since most of the immune-mediated adverse effects against a therapeutic protein product are mediated by humoral mechanisms, circulating antibodies to the therapeutic protein product have been the main criterion for defining an immune response. These should be screened for in clinical studies of IVT mRNA-mediated protein-replacement approaches.
It is also conceivable that the expression of a foreign protein together with the pro-inflammatory effects mediated by the mRNA backbone may result in immunopathology on the tissue level. For immunotherapeutic approaches, this may be of minor relevance as antigen-presenting cells are the target cells of mRNA delivery and these are short-lived once they have transitioned into the mature state. If, however, other organs such as the liver, kidney, lungs or myocardium are targeted, this risk needs to be addressed. Various applications that are being pursued use the liver as the target organ, as it has been shown (at least for various siRNA delivery platforms) that nucleic-acid-based drugs are routed to the liver by default, and therefore liver targeting can be accomplished without further optimizing delivery195. As hepatic toxicity may be life threatening, particular caution is warranted and liver enzymes such as transaminases need to be measured. However, given the unique immunological properties of this organ, it may even be de-risking to use the liver as a depot organ for protein expression of the first-generation of mRNA-based therapeutics for protein replacement, as its capacity to induce antigen-specific tolerance may counteract immunogenecity196.
Risks associated with non-natural nucleotides. The highly abundant extracellular RNases have evolved as a powerful control mechanism of RNA levels in the extracellular space197. No significant risks are anticipated to be associated with the absorption, distribution, metabolism and excretion profile of IVT mRNA drugs that are composed of natural nucleotides because the human body breaks down much higher amounts of natural mRNA every day. However, this may not apply to investigational mRNA drugs containing unnatural modified nucleotides. Mechanisms of catabolism and excretion and potential unwanted cross-effects on other toxicity-relevant pathways of unnatural nucleotides in a polynucleotide structure or their metabolites and potential risks associated with these are still unknown.
This caveat is supported by observations from unnatural nucleoside analogues used as antiviral and anticancer drugs that interfere with viral and tumour cell replication. Many of these nucleoside analogues demonstrated unexpected mitochondrial toxicities198,199 that are associated with functions of nucleoside transporters200. Nucleoside reverse transcriptase inhibitors used for treating HIV-infected patients caused severe clinical toxicities (for example, myopathy, polyneuropathy, lactic acidosis, liver steatosis, pancreatitis and lipodystrophy) “including fatal complications due to mitochondrial dysfunction199. These adverse effects of unnatural modified nucleosides were caused by the inhibition of DNA polymerase γ, an enzyme solely responsible for mitochondrial DNA replication, blocking de novo mitochondrial DNA synthesis201. These risks were not identified in preclinical studies performed in mice and rats owing to interspecies differences in the subcellular localization of nucleoside transporter 1 (Refs 202,203).
In clinical trial design, the potential toxicity of nucleoside analogues should be addressed diligently by conservative dose-escalation regimens and close assessment of risk organs. Safety monitoring has to consider that adverse effects may only occur after prolonged treatment with nucleoside analogues.
Safety considerations regarding the encoded protein. In addition to the risks described above, 'content'-specific risks determined by the nature of the encoded protein and by the type of application have to be considered. The number of genes and modes of action executed by these genes is highly diverse. Therefore, no general exhaustive risk assessment can be provided; instead, risks have to be evaluated with due diligence on a case-by-case basis.
Depending on the specific application, it may be an important safety measure to ensure that in vivo transferred RNA only enters the cell type it is intended for.
Another caveat concerns proteins that are challenging with regard to dosing, such as proteins with a narrow therapeutic window or with a steep dose–response relationship. The key challenge for such protein targets is to control the robustness and fidelity of their bioavailability and address potential inter-individual variations by closely monitored, individually adjusted dosing schedules.
Conclusions and perspectives
As outlined in this Review, cancer immunotherapy is the only field in which clinical testing and industrialization of the manufacturing of mRNA drugs is at an advanced stage. For vaccination against infectious diseases, IVT mRNA is in early clinical testing, whereas in all other medical applications, such as protein replacement, it is at the preclinical stage.
The instability of mRNA (originally perceived as the primary hurdle in RNA drug development) has been efficiently addressed. Intracellular stability can be achieved and the half-life of mRNA translational activity tuned from minutes to days by structural elements that modulate mRNA translation and mRNA metabolism, and that are now used in the design of IVT mRNAs. Extracellular stability is being addressed through the development of formulations — for example, protamine and nanoparticle carriers. Progress in de-immunization techniques of IVT mRNA has facilitated the control of the inflammatory activity of mRNA in animal models. Moreover, for the biopharmaceutical development of mRNA drugs, the initial foundations regarding patent and intellectual property issues have been laid (Box 5). IVT mRNA can be manufactured at relatively low costs and within a few hours, and the production and purification processes are robust, enabling the generation of mRNA ranging from a few hundred to more than 10,000 nucleotides in length. The robustness and ease of the production process facilitates the implementation of high-throughput approaches for drug discovery and iterative drug optimization. Once the clinical mRNA drug candidate is identified, process optimization and clinical-grade good manufacturing practice (GMP) production can be carried out within several months. In our experience, production costs for GMP batches for clinical studies are on average five- to tenfold lower for IVT mRNA than for recombinant protein therapeutics produced in eukaryotic cells.
The major challenges for which satisfactory solutions are still pending, in particular for non-immunotherapy-related in vivo applications, are targeting to the desired organ or cell type in vivo and the complex pharmacology of IVT mRNA. This means that the question of consistent dosing across tissues and patients can become a significant roadblock for the clinical development of in vivo administered IVT mRNA. As discussed above, it is still unclear how to accurately deliver the IVT mRNA to the target cell type and how to achieve the right therapeutic dose level. Moreover, it has not been thoroughly investigated whether mRNA dose–protein-effect relationships vary inter-individually or even intra-individually when comparing independent routes of administration.
In this respect, the use of cells that are transfected ex vivo with IVT mRNA, particularly for immunotherapeutic approaches, can be viewed as the 'low-hanging fruit'. For immunotherapy, relatively small amounts of IVT mRNA encoding the corresponding antigen are sufficient to obtain robust efficacy signals, which is further supported by the intrinsic adjuvant activity of mRNA. Moreover, professional antigen-presenting cells, which are the targets for mRNA-based vaccine delivery, are constitutively equipped with a specialized mechanism for mRNA uptake. Beyond applications for cancer immunotherapy, mRNA-based vaccine development may also create opportunities to manage newly emerging pandemics. Recent progress in synthetic DNA technology has enabled the rapid and accurate synthesis of genes encoding any potential target antigen205. The large-scale manufacturing of rapidly assembled synthetic genes that are suitable DNA templates for in vitro transcription could accelerate the overall process of mRNA-based vaccine production.
When extending IVT mRNA therapeutics into applications such as protein-replacement therapies, delivery, dosability and robustness of dosing, as well as the tissue selectivity of the in vivo administered mRNA drug need to be carefully addressed. Moreover, immune stimulation is unwanted. As a consequence, the hurdle for advancing non-immunotherapy applications is higher and its acceleration through spill-over effects from the immunotherapy field is limited. For many protein-replacement therapy applications, IVT mRNA delivery may be successfully realized by optimizing existing delivery tools. The most reasonable approach would be to select diseases in which the target tissues are easily accessible and the encoded proteins are active even at low doses and have broad therapeutic windows.
For the development of mRNA as a biopharmaceutical, the mRNA technology platform has to become an industry-compatible process. For IVT mRNA that is used ex vivo in cell therapeutic applications, this will be limited by the challenging industrialization hurdles faced by cell therapy206. IVT mRNA for in vivo use, by contrast, follows 'platform'-specific patterns in terms of general pharmaceutical properties, and its manufacturing is straightforward, cost-effective and does not carry specific challenges. Progress will also depend on how process automation will evolve and whether specialized companies can deliver standard or customized equipment for that purpose. For product-based companies seeking to outsource manufacturing, the low number of service providers can impede project planning and timelines. The development of a diversified landscape of a service and supply industry around the core mRNA drug product has begun and will facilitate industrialization.
Under the shadow of disappointments and failures in the neighbouring fields of gene therapy and siRNA, the mRNA field has been advanced with due caution. Cardinal faults such as premature adoption of new technology, clinical trials with unnecessary safety risks, as well as unrealistic expectations of industry leaders and investors, have been avoided. Ongoing clinical testing programmes have been initiated based on thorough preclinical exploration and understanding of underlying mechanisms. It is advisable that this prudence is further maintained.
References
Wolff, J. A. et al. Direct gene transfer into mouse muscle in vivo. Science 247, 1465–1468 (1990).
Jirikowski, G. F. et al. Reversal of diabetes insipidus in Brattleboro rats: intrahypothalamic injection of vasopressin mRNA. Science 255, 996–998 (1992).
Martinon, F. et al. Induction of virus-specific cytotoxic T lymphocytes in vivo by liposome-entrapped mRNA. Eur. J. Immunol. 23, 1719–1722 (1993).
Conry, R. M. et al. Characterization of a messenger RNA polynucleotide vaccine vector. Cancer Res. 55, 1397–1400 (1995).
Boczkowski, D. et al. Dendritic cells pulsed with RNA are potent antigen-presenting cells in vitro and in vivo. J. Exp. Med. 184, 465–472 (1996).
Qiu, P. et al. Gene gun delivery of mRNA in situ results in efficient transgene expression and genetic immunization. Gene Ther. 3, 262–268 (1996).
Mandl, C. W. et al. In vitro-synthesized infectious RNA as an attenuated live vaccine in a flavivirus model. Nature Med. 4, 1438–1440 (1998).
Zhou, W. Z. et al. RNA melanoma vaccine: induction of antitumor immunity by human glycoprotein 100 mRNA immunization. Hum. Gene Ther. 10, 2719–2724 (1999).
Hoerr, I. et al. In vivo application of RNA leads to induction of specific cytotoxic T lymphocytes and antibodies. Eur. J. Immunol. 30, 1–7 (2000).
Koido, S. et al. Induction of antitumor immunity by vaccination of dendritic cells transfected with MUC1 RNA. J. Immunol. 165, 5713–5719 (2000).
Schirmacher, V. et al. Intra-pinna anti-tumor vaccination with self-replicating infectious RNA or with DNA encoding a model tumor antigen and a cytokine. Gene Ther. 7, 1137–1147 (2000).
Heiser, A. et al. Autologous dendritic cells transfected with prostate-specific antigen RNA stimulate CTL responses against metastatic prostate tumors. J. Clin. Invest. 109, 409–417 (2002).
Morse, M. A. et al. The feasibility and safety of immunotherapy with dendritic cells loaded with CEA mRNA following neoadjuvant chemoradiotherapy and resection of pancreatic cancer. Int. J. Gastrointest Cancer 32, 1–6 (2002).
Morse, M. A. et al. Immunotherapy with autologous, human dendritic cells transfected with carcinoembryonic antigen mRNA. Cancer Invest. 21, 341–349 (2003).
Su, Z. et al. Telomerase mRNA-transfected dendritic cells stimulate antigen-specific CD8+ and CD4+ T cell responses in patients with metastatic prostate cancer. J. Immunol. 174, 3798–3807 (2005).
Weide, B. et al. Direct injection of protamine-protected mRNA: results of a phase 1/2 vaccination trial in metastatic melanoma patients. J. Immunother. 32, 498–507 (2009).
Rittig, S. M. et al. Intradermal vaccinations with RNA coding for TAA generate CD8+ and CD4+ immune responses and induce clinical benefit in vaccinated patients. Mol. Ther. 19, 990–999 (2011).
Wilgenhof, S. et al. A phase IB study on intravenous synthetic mRNA electroporated dendritic cell immunotherapy in pretreated advanced melanoma patients. Ann. Oncol. 24, 2686–2693 (2013).
Okumura, K. et al. Bax mRNA therapy using cationic liposomes for human malignant melanoma. J. Gene Med. 10, 910–917 (2008).
Mitchell, D. A. et al. Selective modification of antigen-specific T cells by RNA electroporation. Hum. Gene Ther. 19, 511–521 (2008).
Wang, Y. et al. Systemic delivery of modified mRNA encoding herpes simplex virus 1 thymidine kinase for targeted cancer gene therapy. Mol. Ther. 21, 358–367 (2013).
Zimmermann, O. et al. Successful use of mRNA-nucleofection for overexpression of interleukin-10 in murine monocytes/macrophages for anti-inflammatory therapy in a murine model of autoimmune myocarditis. J. Am. Heart Assoc. 1, e003293 (2012).
Zangi, L. et al. Modified mRNA directs the fate of heart progenitor cells and induces vascular regeneration after myocardial infarction. Nature Biotech. 31, 898–907 (2013).
Creusot, R. J. et al. A short pulse of IL-4 delivered by DCs electroporated with modified mRNA can both prevent and treat autoimmune diabetes in NOD mice. Mol. Ther. 18, 2112–2120 (2010).
Kormann, M. S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nature Biotech. 29, 154–157 (2011).
Kariko, K. et al. Increased erythropoiesis in mice injected with submicrogram quantities of pseudouridine-containing mRNA encoding erythropoietin. Mol. Ther. 20, 948–953 (2012).
Mays, L. E. et al. Modified Foxp3 mRNA protects against asthma through an IL-10-dependent mechanism. J. Clin. Invest. 123, 1216–1228 (2013).
Levy, O. et al. mRNA-engineered mesenchymal stem cells for targeted delivery of interleukin-10 to sites of inflammation. Blood 122, e23–e32 (2013).
Holtkamp, S. et al. Modification of antigen-encoding RNA increases stability, translational efficacy, and T-cell stimulatory capacity of dendritic cells. Blood 108, 4009–4017 (2006).
Karikó, K., Kuo, A. & Barnathan, E. Overexpression of urokinase receptor in mammalian cells following administration of the in vitro transcribed encoding mRNA. Gene Ther. 6, 1092–1100 (1999).
Kallen, K.-J. & Theß, A. A development that may evolve into a revolution in medicine: mRNA as the basis for novel, nucleotide-based vaccines and drugs. Ther. Adv. Vaccines 2, 10–31 (2014).
Li, Y. & Kiledjian, M. Regulation of mRNA decapping. Wiley Interdiscip. Rev. RNA 1, 253–265 (2010).
Martin, S. A., Paoletti, E. & Moss, B. Purification of mRNA guanylyltransferase and mRNA (guanine-7-) methyltransferase from vaccinia virions. J. Biol. Chem. 250, 9322–9329 (1975).
Malone, R. W., Felgner, P. L. & Verma, I. M. Cationic liposome-mediated RNA transfection. Proc. Natl Acad. Sci. USA 86, 6077–6081 (1989).
Stepinski, J. et al. Synthesis and properties of mRNAs containing the novel “anti-reverse” cap analogs 7-methyl(3′-O-methyl)GpppG and 7-methyl (3′-deoxy)GpppG. RNA 7, 1486–1495 (2001).
Jemielity, J. et al. Novel “anti-reverse” cap analogs with superior translational properties. RNA 9, 1108–1122 (2003).
Mockey, M. et al. mRNA transfection of dendritic cells: synergistic effect of ARCA mRNA capping with poly(A) chains in cis and in trans for a high protein expression level. Biochem. Biophys. Res. Commun. 340, 1062–1068 (2006).
Rabinovich, P. M. et al. Synthetic messenger RNA as a tool for gene therapy. Hum. Gene Ther. 17, 1027–1035 (2006).
Grudzien-Nogalska, E. et al. Phosphorothioate cap analogs stabilize mRNA and increase translational efficiency in mammalian cells. RNA 13, 1745–1755 (2007).
Kowalska, J. et al. Synthesis and characterization of mRNA cap analogs containing phosphorothioate substitutions that bind tightly to eIF4E and are resistant to the decapping pyrophosphatase DcpS. RNA 14, 1119–1131 (2008).
Kuhn, A. N. et al. Phosphorothioate cap analogs increase stability and translational efficiency of RNA vaccines in immature dendritic cells and induce superior immune responses in vivo. Gene Ther. 17, 961–971 (2010).
Gallie, D. R. The cap and poly(A) tail function synergistically to regulate mRNA translational efficiency. Genes Dev. 5, 2108–2116 (1991).
Korner, C. G. & Wahle, E. Poly(A) tail shortening by a mammalian poly(A)-specific 3′-exoribonuclease. J. Biol. Chem. 272, 10448–10456 (1997).
Martin, G. & Keller, W. Tailing and 3′-end labeling of RNA with yeast poly(A) polymerase and various nucleotides. RNA 4, 226–230 (1998).
Ross, J. & Sullivan, T. Half-lives of β and γ globin messenger RNAs and of protein synthetic capacity in cultured human reticulocytes. Blood 66, 1149–1154 (1985).
Zinckgraf, J. W. & Silbart, L. K. Modulating gene expression using DNA vaccines with different 3′-UTRs influences antibody titer, seroconversion and cytokine profiles. Vaccine 21, 1640–1649 (2003).
Bergman, N. et al. Lsm proteins bind and stabilize RNAs containing 5′ poly(A) tracts. Nature Struct. Mol. Biol. 14, 824–831 (2007).
Kuhn, A. N. et al. mRNA as a versatile tool for exogenous protein expression. Curr. Gene Ther. 12, 347–361 (2012).
Chen, C. Y. & Shyu, A. B. AU-rich elements: characterization and importance in mRNA degradation. Trends Biochem. Sci. 20, 465–470 (1995).
Gustafsson, C., Govindarajan, S. & Minshull, J. Codon bias and heterologous protein expression. Trends Biotechnol. 22, 346–353 (2004).
Cannarozzi, G. et al. A role for codon order in translation dynamics. Cell 141, 355–367 (2010).
Bossi, L. & Ruth, J. R. The influence of codon context on genetic code translation. Nature 286, 123–127 (1980).
Gustafsson, C. et al. Engineering genes for predictable protein expression. Protein Expr. Purif. 83, 37–46 (2012).
Van Gulck, E. R. A. et al. Efficient stimulation of HIV-1-specific T cells using dendritic cells electroporated with mRNA encoding autologous HIV-1 Gag and Env proteins. Blood 107, 1818–1827 (2006).
Kimchi-Sarfaty, C. et al. A “silent” polymorphism in the MDR1 gene changes substrate specificity. Science 315, 525–528 (2007).
Malarkannan, S. et al. Presentation of out-of-frame peptide/MHC class I complexes by a novel translation initiation mechanism. Immunity 10, 681–690 (1999).
Saulquin, X. et al. +1 frameshifting as a novel mechanism to generate a cryptic cytotoxic T lymphocyte epitope derived from human interleukin 10. J. Exp. Med. 195, 353–358 (2002).
Schwab, S. R. et al. Constitutive display of cryptic translation products by MHC class I molecules. Science 301, 1367–1371 (2003).
Bourquin, C. et al. Immunostimulatory RNA oligonucleotides trigger an antigen-specific cytotoxic T-cell and IgG2a response. Blood 109, 2953–2960 (2007).
Sander, L. E. et al. Detection of prokaryotic mRNA signifies microbial viability and promotes immunity. Nature 474, 385–389 (2011).
Weissman, D. et al. HIV gag mRNA transfection of dendritic cells (DC) delivers encoded antigen to MHC class I and II molecules, causes DC maturation, and induces a potent human in vitro primary immune response. J. Immunol. 165, 4710–4717 (2000).
Fotin-Mleczek, M. et al. Messenger RNA-based vaccines with dual activity induce balanced TLR-7 dependent adaptive immune responses and provide antitumor activity. J. Immunother. 34, 1–15 (2011).
Karikó, K. et al. Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res. 39, e142 (2011).
Rettig, L. et al. Particle size and activation threshold: a new dimension of danger signaling. Blood 115, 4533–4541 (2010).
Alexopoulou, L. et al. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Diebold, S. S. et al. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Heil, F. et al. Species-specific recognition of single-stranded RNA via Toll-like receptor 7 and 8. Science 303, 1526–1529 (2004).
Diebold, S. S. et al. Nucleic acid agonists for Toll-like receptor 7 are defined by the presence of uridine ribonucleotides. Eur. J. Immunol. 36, 3256–3267 (2006).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nature Immunol. 5, 730–737 (2004).
Yoneyama, M. et al. Shared and unique functions of the DExD/H-Box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity. J. Immunol. 175, 2851–2858 (2005).
Schlee, M. et al. Recognition of 5′ triphosphate by RIG-I helicase requires short blunt double-stranded RNA as contained in panhandle of negative-strand virus. Immunity 31, 25–34 (2009).
Pichlmair, A. et al. Activation of MDA5 requires higher-order RNA structures generated during virus infection. J. Virol. 83, 10761–10769 (2009).
Zust, R. et al. Ribose 2′-O-methylation provides a molecular signature for the distinction of self and non-self mRNA dependent on the RNA sensor Mda5. Nature Immunol. 12, 137–143 (2011).
Balachandran, S. et al. Essential role for the dsRNA-dependent protein kinase PKR in innate immunity to viral infection. Immunity 13, 129–141 (2000).
Karikó, K. et al. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Hornung, V. et al. 5′-triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Nallagatla, S. R. & Bevilacqua, P. C. Nucleoside modifications modulate activation of the protein kinase PKR in an RNA structure-specific manner. RNA 14, 1201–1213 (2008).
Anderson, B. R. et al. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic Acids Res. 38, 5884–5892 (2010).
Lorenz, C. et al. Protein expression from exogenous mRNA: uptake by receptor-mediated endocytosis and trafficking via the lysosomal pathway. RNA Biol. 8, 627–636 (2011).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biol. 9, 654–659 (2007).
Diken, M. et al. Selective uptake of naked vaccine RNA by dendritic cells is driven by macropinocytosis and abrogated upon DC maturation. Gene Ther. 18, 702–708 (2011).
Wang, W. et al. Non-viral gene delivery methods. Curr. Pharm. Biotechnol. 14, 46–60 (2013).
Neumann, E. et al. Gene transfer into mouse lyoma cells by electroporation in high electric fields. EMBO J. 1, 841–845 (1982).
Van Tendeloo, V. F. et al. Nonviral transfection of distinct types of human dendritic cells: high-efficiency gene transfer by electroporation into hematopoietic progenitor- but not monocyte-derived dendritic cells. Gene Ther. 5, 700–707 (1998).
Kyte, J. A. et al. Phase I/II trial of melanoma therapy with dendritic cells transfected with autologous tumor-mRNA. Cancer Gene Ther. 13, 905–918 (2006).
Van Driessche, A. et al. Clinical-grade manufacturing of autologous mature mRNA-electroporated dendritic cells and safety testing in acute myeloid leukemia patients in a phase I dose-escalation clinical trial. Cytotherapy 11, 653–668 (2009).
Van Tendeloo, V. F. et al. Induction of complete and molecular remissions in acute myeloid leukemia by Wilms' tumor 1 antigen-targeted dendritic cell vaccination. Proc. Natl Acad. Sci. USA 107, 13824–13829 (2010).
Wilgenhof, S. et al. Therapeutic vaccination with an autologous mRNA electroporated dendritic cell vaccine in patients with advanced melanoma. J. Immunother. 34, 448–456 (2011).
Van Nuffel, A. M. et al. Intravenous and intradermal TriMix-dendritic cell therapy results in a broad T-cell response and durable tumor response in a chemorefractory stage IV-M1c melanoma patient. Cancer Immunol. Immunother. 61, 1033–1043 (2012).
Van Nuffel, A. M. et al. Dendritic cells loaded with mRNA encoding full-length tumor antigens prime CD4+ and CD8+ T cells in melanoma patients. Mol. Ther. 20, 1063–1074 (2012).
Geng, T. et al. Transfection of cells using flow-through electroporation based on constant voltage. Nature Protoc. 6, 1192–1208 (2011).
Weide, B. et al. Results of the first phase I/II clinical vaccination trial with direct injection of mRNA. J. Immunother. 31, 180–188 (2008).
Wang, T., Upponi, J. R. & Torchilin, V. P. Design of multifunctional non-viral gene vectors to overcome physiological barriers: dilemmas and strategies. Int. J. Pharm. 427, 3–20 (2012).
Semple, S. C. et al. Rational design of cationic lipids for siRNA delivery. Nature Biotech. 28, 172–176 (2010).
Coelho, T. et al. Safety and efficacy of RNAi therapy for transthyretin amyloidosis. N. Engl. J. Med. 369, 819–829 (2013).
Granstein, R. D., Ding, W. & Ozawa, H. Induction of anti-tumor immunity with epidermal cells pulsed with tumor-derived RNA or intradermal administration of RNA. J. Invest. Dermatol. 114, 632–636 (2000).
Kreiter, S. et al. Intranodal vaccination with naked antigen-encoding RNA elicits potent prophylactic and therapeutic antitumoral immunity. Cancer Res. 70, 9031–9040 (2010).
Kreiter, S. et al. FLT3 ligand enhances the cancer therapeutic potency of naked RNA vaccines. Cancer Res. 71, 6132–6142 (2011).
Van Lint, S. et al. Preclinical evaluation of TriMix and antigen mRNA-based antitumor therapy. Cancer Res. 72, 1661–1671 (2012).
Petsch, B. et al. Protective efficacy of in vitro synthesized, specific mRNA vaccines against influenza A virus infection. Nature Biotech. 30, 1210–1216 (2012).
Geall, A. J. et al. Nonviral delivery of self-amplifying RNA vaccines. Proc. Natl Acad. Sci. USA 109, 14604–14609 (2012).
Uchida, S. et al. In vivo messenger RNA introduction into the central nervous system using polyplex nanomicelle. PLoS ONE 8, e56220 (2013).
Azarmi, S., Roa, W. H. & Lobenberg, R. Targeted delivery of nanoparticles for the treatment of lung diseases. Adv. Drug Deliv. Rev. 60, 863–875 (2008).
Torchilin, V. Tumor delivery of macromolecular drugs based on the EPR effect. Adv. Drug Deliv. Rev. 63, 131–135 (2011).
van der Bruggen, P. et al. A gene encoding an antigen recognized by cytolytic T lymphocytes on a human melanoma. Science 254, 1643–1647 (1991).
Sahin, U. et al. Human neoplasms elicit multiple specific immune responses in the autologous host. Proc. Natl Acad. Sci. USA 92, 11810–11813 (1995).
Nair, S. K. et al. Induction of cytotoxic T cell responses and tumor immunity against unrelated tumors using telomerase reverse transcriptase RNA transfected dendritic cells. Nature Med. 6, 1011–1017 (2000).
Nair, S. K. et al. Induction of primary carcinoembryonic antigen (CEA)-specific cytotoxic T lymphocytes in vitro using human dendritic cells transfected with RNA. Nature Biotech. 16, 364–369 (1998).
Morse, M. A. et al. Optimization of the sequence of antigen loading and CD40-ligand-induced maturation of dendritic cells. Cancer Res. 58, 2965–2968 (1998).
Van Lint, S. et al. mRNA: From a chemical blueprint for protein production to an off-the-shelf therapeutic. Hum. Vaccin. Immunother. 9, 265–274 (2013).
Kreiter, S. et al. Tumor vaccination using messenger RNA: prospects of a future therapy. Curr. Opin. Immunol. 23, 399–406 (2011).
Cisco, R. M. et al. Induction of human dendritic cell maturation using transfection with RNA encoding a dominant positive Toll-like receptor 4. J. Immunol. 172, 7162–7168 (2004).
Bonehill, A. et al. Enhancing the T-cell stimulatory capacity of human dendritic cells by co-electroporation with CD40L, CD70 and constitutively active TLR4 encoding mRNA. Mol. Ther. 16, 1170–1180 (2008).
Calderhead, D. M. et al. Cytokine maturation followed by CD40L mRNA electroporation results in a clinically relevant dendritic cell product capable of inducing a potent proinflammatory CTL response. J. Immunother. 31, 731–741 (2008).
Routy, J.-P. et al. Immunologic activity and safety of autologous HIV RNA-electroporated dendritic cells in HIV-1 infected patients receiving antiretroviral therapy. Clin. Immunol. 134, 140–147 (2010).
Bontkes, H. J. et al. Dendritic cells transfected with interleukin-12 and tumor-associated antigen messenger RNA induce high avidity cytotoxic T cells. Gene Ther. 14, 366–375 (2007).
Aarntzen, E. H. J. G. et al. Vaccination with mRNA-electroporated dendritic cells induces robust tumor antigen-specific CD4+ and CD8+ T cells responses in stage III and IV melanoma patients. Clin. Cancer Res. 18, 5460–5470 (2012).
Pascolo, S. Vaccination with messenger RNA (mRNA). Handb Exp. Pharmacol. 183, 221–235 (2008).
Carralot, J. P. et al. Polarization of immunity induced by direct injection of naked sequence-stabilized mRNA vaccines. Cell. Mol. Life Sci. 61, 2418–2424 (2004).
Scheel, B. et al. Toll-like receptor-dependent activation of several human blood cell types by protamine-condensed mRNA. Eur. J. Immunol. 35, 1557–1566 (2005).
Fotin-Mleczek, M. et al. Highly potent mRNA based cancer vaccines represent an attractive platform for combination therapies supporting an improved therapeutic effect. J. Gene Med. 14, 428–439 (2012).
Kreiter, S. et al. Simultaneous ex vivo quantification of antigen-specific CD4+ and CD8+ T cell responses using in vitro transcribed RNA. Cancer Immunol. Immunother. 56, 1577–1587 (2007).
Kreiter, S. et al. Increased antigen presentation efficiency by coupling antigens to MHC class I trafficking signals. J. Immunol. 180, 309–318 (2008).
Diken, M. et al. Antitumor vaccination with synthetic mRNA: strategies for in vitro and in vivo preclinical studies. Methods Mol. Biol. 969, 235–246 (2013).
Diken, M. et al. mTOR inhibition improves antitumor effects of vaccination with antigen-encoding RNA. Cancer Immunol. Res. 1, 386–392 (2013).
Castle, J. C. et al. Exploiting the mutanome for tumor vaccination. Cancer Res. 72, 1081–1091 (2012).
Kreiter, S. et al. Targeting the tumor mutanome for personalized vaccination therapy. Oncoimmunology 1, 768–769 (2012).
Britten, C. M. et al. The regulatory landscape for actively personalized cancer immunotherapies. Nature Biotech. 31, 880–882 (2013).
Zhao, Y. et al. Multiple injections of electroporated autologous T cells expressing a chimeric antigen receptor mediate regression of human disseminated tumor. Cancer Res. 70, 9053–9061 (2010).
Almasbak, H. et al. Transiently redirected T cells for adoptive transfer. Cytotherapy 13, 629–640 (2011).
Barrett, D. M. et al. Treatment of advanced leukemia in mice with mRNA engineered T cells. Hum. Gene Ther. 22, 1575–1586 (2011).
Barrett, D. M. et al. Regimen specific effects of RNA-modified chimeric antigen receptor T cells in mice with advanced leukemia. Hum. Gene Ther. 24, 717–727 (2013).
Hekele, A. et al. Rapidly produced SAM® vaccine against H7N9 influenza is immunogenic in mice. Emerg. Microbes Infect. 2, e52 (2013).
Allard, S. D. et al. A phase I/IIa immunotherapy trial of HIV-1-infected patients with Tat, Rev and Nef expressing dendritic cells followed by treatment interruption. Clin. Immunol. 142, 252–268 (2012).
Van Gulck, E. et al. mRNA-based dendritic cell vaccination induces potent antiviral T-cell responses in HIV-1-infected patients. AIDS 26, F1–F12 (2012).
Liljestrom, P. & Garoff, H. A new generation of animal cell expression vectors based on the Semliki Forest virus replicon. Biotechnology 9, 1356–1361 (1991).
Zhou, X. et al. Self-replicating Semliki Forest virus RNA as recombinant vaccine. Vaccine 12, 1510–1514 (1994).
Ulmer, J. B. et al. RNA-based vaccines. Vaccine 30, 4414–4418 (2012).
Geall, A. J., Mandl, C. W. & Ulmer, J. B. RNA: the new revolution in nucleic acid vaccines. Semin. Immunol. 25, 152–159 (2013).
Fleeton, M. N. et al. Self-replicative RNA vaccines elicit protection against influenza A virus, respiratory syncytial virus, and a tickborne encephalitis virus. J. Infect. Dis. 183, 1395–1398 (2001).
Anraku, I. et al. Kunjin virus replicon vaccine vectors induce protective CD8+ T-cell immunity. J. Virol. 76, 3791–3799 (2002).
Greer, C. E. et al. A chimeric alphavirus RNA replicon gene-based vaccine for human parainfluenza virus type 3 induces protective immunity against intranasal virus challenge. Vaccine 25, 481–489 (2007).
Valenta, R. et al. From allergen genes to allergy vaccines. Annu. Rev. Immunol. 28, 211–241 (2010).
Raz, E. et al. Preferential induction of a Th1 immune response and inhibition of specific IgE antibody formation by plasmid DNA immunization. Proc. Natl Acad. Sci. USA 93, 5141–5145 (1996).
Chua, K. Y., Kuo, I. C. & Huang, C. H. DNA vaccines for the prevention and treatment of allergy. Curr. Opin. Allergy Clin. Immunol. 9, 50–54 (2009).
Slater, J. E. et al. The latex allergen Hev b 5 transcript is widely distributed after subcutaneous injection in BALB/c mice of its DNA vaccine. J. Allergy Clin. Immunol. 102, 469–475 (1998).
Roesler, E. et al. Immunize and disappear-safety-optimized mRNA vaccination with a panel of 29 allergens. J. Allergy Clin. Immunol, 124, 1070–1077.e11 (2009).
Weiss, R. et al. Prophylactic mRNA vaccination against allergy. Curr. Opin. Allergy Clin. Immunol. 10, 567–574 (2010).
Kolarich, D. et al. Comprehensive glyco-proteomic analysis of human α1-antitrypsin and its charge isoforms. Proteomics 6, 3369–3380 (2006).
Seidah, N. G. & Chrétien, M. Proprotein and prohormone convertases: a family of subtilases generating diverse bioactive polypeptides. Brain Res. 848, 45–62 (1999).
Nakayama, K. Furin: a mammalian subtilisin/Kex2p-like endoprotease involved in processing of a wide variety of precursor proteins. Biochem. J. 327, 625–635 (1997).
Seidah, N. G. et al. Precursor convertases: an evolutionary ancient, cell-specific, combinatorial mechanism yielding diverse bioactive peptides and proteins. Ann. NY Acad. Sci. 839, 9–24 (1998).
Barash, S., Wang, W. & Shi, Y. Human secretory signal peptide description by hidden Markov model and generation of a strong artificial signal peptide for secreted protein expression. Biochem. Biophys. Res. Commun. 294, 835–842 (2002).
Fattori, E. et al. Gene electro-transfer of an improved erythropoietin plasmid in mice and non-human primates. J. Gene Med. 7, 228–236 (2005).
Roberts, A. A. et al. Engineering factor Viii for hemophilia gene therapy. J. Genet. Syndr. Gene Ther. 1, S1–006 (2011).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Warren, L. et al. Feeder-free derivation of human induced pluripotent stem cells with messenger RNA. Sci. Rep. 2, 657 (2012).
Mandal, P. K. & Rossi, D. J. Reprogramming human fibroblasts to pluripotency using modified mRNA. Nature Protoc. 8, 568–582 (2013).
Bernal, J. A. RNA-based tools for nuclear reprogramming and lineage-conversion: towards clinical applications. J. Cardiovasc. Transl. Res. 6, 956–968 (2013).
Miller, J. D. et al. Human iPSC-based modeling of late-onset disease via progerin-induced aging. Cell Stem Cell 13, 691–705 (2013).
Scott, C. W., Peters, M. F. & Dragan, Y. P. Human induced pluripotent stem cells and their use in drug discovery for toxicity testing. Toxicol. Lett. 219, 49–58 (2013).
Okano, H. et al. Steps toward safe cell therapy using induced pluripotent stem cells. Circ. Res. 112, 523–533 (2013).
Miller, J. C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nature Biotech. 25, 778–785 (2007).
Hockemeyer, D. et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nature Biotech. 29, 731–734 (2011).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nature Biotech. 31, 822–826 (2013).
Doyon, Y. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nature Biotech. 26, 702–708 (2008).
Tesson, L. et al. Knockout rats generated by embryo microinjection of TALENs. Nature Biotech. 29, 695–696 (2011).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Wefers, B. et al. Generation of targeted mouse mutants by embryo microinjection of TALEN mRNA. Nature Protocols 8, 2355–2379 (2013).
Geurts, A. M. et al. Knockout rats via embryo microinjection of zinc-finger nucleases. Science 325, 433 (2009).
Shen, B. et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res. 23, 720–723 (2013).
Yang, D. et al. Effective gene targeting in rabbits using RNA-guided Cas9 nucleases. J. Mol. Cell Biol. 6, 97–99 (2014).
Ma, Y. et al. Generating rats with conditional alleles using CRISPR/Cas9. Cell Res. 24, 122–125 (2014).
Niu, Y. et al. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156, 836–843 (2014).
Dupuy, A. J. et al. Mammalian germ-line transgenesis by transposition. Proc. Natl Acad. Sci. USA 99, 4495–4499 (2002).
Wilber, A. et al. RNA as a source of transposase for sleeping beauty-mediated gene insertion and expression in somatic cells and tissues. Mol. Ther. 13, 625–630 (2006).
Sumiyama, K., Kawakami, K. & Yagita, K. A simple and highly efficient transgenesis method in mice with the Tol2 transposon system and cytoplasmic microinjection. Genomics 95, 306–311 (2010).
Suster, M. L., Sumiyama, K. & Kawakami, K. Transposon-mediated BAC transgenesis in zebrafish and mice. BMC Genomics 10, 477 (2009).
Furushima, K. et al. Insertional mutagenesis by a hybrid piggyBac and sleeping beauty transposon in the rat. Genetics 192, 1235–1248 (2012).
Bire, S. et al. Exogenous mRNA delivery and bioavailability in gene transfer mediated by piggyBac transposition. BMC Biotechnol. 13, 75 (2013).
Isaacs, A., Cox, R. A. & Rotem, Z. Foreign nucleic acids as the stimulus to make interferon. Lancet 282, 113–116 (1963).
Tytell, A. A. et al. Inducers of interferon and host resistance. 3. Double-stranded RNA from reovirus type 3 virions (reo 3-RNA). Proc. Natl Acad. Sci. USA 58, 1719–1722 (1967).
Anderson, B. R. et al. Nucleoside modifications in RNA limit activation of 2′-5′-oligoadenylate synthetase and increase resistance to cleavage by RNase L. Nuc. Acids Res. 39, 9329–9338 (2011).
Pollard, C. et al. Type I IFN counteracts the induction of antigen-specific immune responses by lipid-based delivery of mRNA vaccines. Mol. Ther. 21, 251–259 (2013).
Hwang, S. H. et al. B cell TLR7 expression drives anti-RNA autoantibody production and exacerbates disease in systemic lupus erythematosus-prone mice. J. Immunol. 189, 5786–5796 (2012).
Lipes, B. D. & Keene, J. D. Autoimmune epitopes in messenger RNA. RNA 8, 762–771 (2002).
Murphy, K. (ed) in Janeway's Immunobiology 367–408 (Garland Science Publishing, 2011).
Worobec, A. & Rosenberg, A. S. A risk-based approach to immunogenicity concerns of therapeutic protein products, part 1: considering consequences of the immune response to a protein. BioPharm International 22–26 (2004).
Koren, E. et al. Recommendations on risk-based strategies for detection and characterization of antibodies against biotechnology products. J. Immunol. Methods 333, 1–9 (2008).
Casadevall, N. et al. Pure red-cell aplasia and antierythropoietin antibodies in patients treated with recombinant erythropoietin. N. Engl. J. Med. 346, 469–475 (2002).
Gao, G. et al. Erythropoietin gene therapy leads to autoimmune anemia in macaques. Blood 103, 3300–3302 (2004).
Kromminga, A. & Schellekens, H. Antibodies against erythropoietin and other protein-based therapeutics: an overview. Ann, NY Acad. Sci. 1050, 257–265 (2005).
Czech, M. P., Aouadi, M. & Tesz, G. J. RNAi-based therapeutic strategies for metabolic disease. Nature Rev. Endocrinol. 7, 473–484 (2011).
Racanelli, V. & Rehermann, B. The liver as an immunological organ. Hepatology 43 (Suppl. 1), S54–S62 (2006).
Dyer, K. D. & Rosenberg, H. F. The RNase a superfamily: generation of diversity and innate host defense. Mol. Divers. 13, 13 (2006).
McKenzie, R. et al. Hepatic failure and lactic acidosis due to fialuridine (FIAU), an investigational nucleoside analogue for chronic hepatitis B. N. Engl. J. Med. 333, 1099–1105 (1995).
Lewis, W. Defective mitochondrial DNA replication and NRTIs: pathophysiological implications in AIDS cardiomyopathy. Am. J. Physiol. Heart Circ. Physiol. 284, H1–H9 (2003).
Griffiths, M. et al. Cloning of a human nucleoside transporter implicated in the cellular uptake of adenosine and chemotherapeutic drugs. Nature Med. 3, 89–93 (1997).
Lewis, W. et al. Fialuridine and its metabolites inhibit DNA polymerase γ at sites of multiple adjacent analog incorporation, decrease mtDNA abundance, and cause mitochondrial structural defects in cultured hepatoblasts. Proc. Natl Acad. Sci. USA 93, 3592–3597 (1996).
Lai, Y., Tse, C.-M. & Unadkat, J. D. Mitochondrial expression of the human equilibrative nucleoside transporter 1 (hENT1) results in enhanced mitochondrial toxicity of antiviral drugs. J. Biol. Chem. 279, 4490–4497 (2004).
Lee, E.-W. et al. Identification of the mitochondrial targeting signal of the human equilibrative nucleoside transporter 1 (hENT1): implications for interspecies differences in mitochondrial toxicity of fialuridine. J. Biol. Chem. 281, 16700–16706 (2006).
Yoshioka, N. et al. Efficient generation of human iPSCs by a synthetic self-replicative RNA. Cell Stem Cell 13, 246–254 (2013).
Dormitzer, P. R. et al. Synthetic generation of influenza vaccine viruses for rapid response to pandemics. Sci. Transl. Med. 5, 185ra68 (2013).
Prieels, J.-P. et al. Mastering industrialization of cell therapy products. BioProcess Int. 10, S12–S15 (2012).
Dahm, R. Friedrich Miescher and the discovery of DNA. Dev. Biol. 278, 274–288 (2005).
Brenner, S., Jacob, F. & Meselson, M. An unstable intermediate carrying information from genes to ribosomes for protein synthesis. Nature 190, 576–581 (1961).
Racaniello, V. R. & Baltimore, D. Cloned poliovirus complementary DNA is infectious in mammalian cells. Science 214, 916–919 (1981).
Rice, C. M. et al. Production of infectious RNA transcripts from Sindbis virus cDNA clones: mapping of lethal mutations, rescue of a temperature-sensitive marker, and in vitro mutagenesis to generate defined mutants. J. Virol. 61, 3809–3819 (1987).
Etchison, D. & Ehrenfeld, E. Comparison of replication complexes synthesizing poliovirus RNA. Virology 111, 33–46 (1981).
Mizutani, S. & Colonno, R. J. In vitro synthesis of an infectious RNA from cDNA clones of human rhinovirus type 14. J. Virol. 56, 628–632 (1985).
van der Werf, S. et al. Synthesis of infectious poliovirus RNA by purified T7 RNA polymerase. Proc. Natl Acad. Sci. USA 83, 2330–2334 (1986).
Khromykh, A. A. & Westaway, E. G. Subgenomic replicons of the flavivirus Kunjin: construction and applications. J. Virol. 71, 1497–1505 (1997).
Perri, S. et al. An alphavirus replicon particle chimera derived from venezuelan equine encephalitis and sindbis viruses is a potent gene-based vaccine delivery vector. J. Virol. 77, 10394–10403 (2003).
Rolls, M. M. et al. Novel infectious particles generated by expression of the vesicular stomatitis virus glycoprotein from a self-replicating RNA. Cell 79, 497–506 (1994).
Xiong, C. et al. Sindbis virus: an efficient, broad host range vector for gene expression in animal cells. Science 243, 1188–1191 (1989).
Ying, H. et al. Cancer therapy using a self-replicating RNA vaccine. Nature Med. 5, 823–827 (1999).
Lundstrom, K. Alphaviruses in gene therapy. Viruses 1, 13–25 (2009).
Hewson, R. RNA viruses: emerging vectors for vaccination and gene therapy. Mol. Med. Today 6, 28–35 (2000).
Lundin, P. Is silence still golden? Mapping the RNAi patent landscape. Nature Biotech. 29, 493–497 (2011).
Modrak-Wojcik, A. et al. Eukaryotic translation initiation is controlled by cooperativity effects within ternary complexes of 4E-BP1, eIF4E, and the mRNA 5′ cap. FEBS Lett. 587, 3928–3934 (2013).
Rau, M. et al. A reevaluation of the cap-binding protein, eIF4E, as a rate-limiting factor for initiation of translation in reticulocyte lysate. J. Biol. Chem. 271, 8983–8990 (1996).
Wells, S. E. et al. Circularization of mRNA by eukaryotic translation initiation factors. Mol. Cell 2, 135–140 (1998).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Balagopal, V., Fluch, L. & Nissan, T. Ways and means of eukaryotic mRNA decay. Biochim. Biophys. Acta 1819, 593–603 (2012).
Shyu, A. B., Wilkinson, M. F. & van Hoof, A. Messenger RNA regulation: to translate or to degrade. EMBO J. 27, 471–481 (2008).
Tomecki, R. & Dziembowski, A. Novel endoribonucleases as central players in various pathways of eukaryotic RNA metabolism. RNA 16, 1692–1724 (2010).
Li, W. M., Barnes, T. & Lee, C. H. Endoribonucleases — enzymes gaining spotlight in mRNA metabolism. FEBS J. 277, 627–641 (2010).
Wilusz, J. RNA stability: is it the endo' the world as we know it? Nature Struct. Mol. Biol. 16, 9–10 (2009).
Garneau, N. L., Wilusz, J. & Wilusz, C. J. The highways and byways of mRNA decay. Nature Rev. Mol. Cell Biol. 8, 113–126 (2007).
Bevan, M. J. Cross-priming. Nature Immunol. 7, 363–365 (2006).
Thomsen, L. B. et al. Nanoparticle-derived non-viral genetic transfection at the blood–brain barrier to enable neuronal growth factor delivery by secretion from brain endothelium. Curr. Med. Chem. 18, 3330–3334 (2011).
Hayashi, S. et al. Autocrine-paracrine effects of overexpression of hepatocyte growth factor gene on growth of endothelial cells. Biochem. Biophys. Res. Commun. 220, 539–545 (1996).
Zeis, M. et al. Generation of cytotoxic responses in mice and human individuals against hematological malignancies using survivin-RNA-transfected dendritic cells. J. Immunol. 170, 5391–5397 (2003).
Siegel, S. et al. Induction of cytotoxic T-cell responses against the oncofetal antigen-immature laminin receptor for the treatment of hematologic malignancies. Blood 102, 4416–4423 (2003).
Yoon, S. H. et al. Adoptive immunotherapy using human peripheral blood lymphocytes transferred with RNA encoding Her-2/neu-specific chimeric immune receptor in ovarian cancer xenograft model. Cancer Gene Ther. 16, 489–497 (2009).
Rabinovich, P. M. et al. Chimeric receptor mRNA transfection as a tool to generate antineoplastic lymphocytes. Hum. Gene Ther. 20, 51–61 (2009).
Bonehill, A. et al. Single-step antigen loading and activation of dendritic cells by mRNA electroporation for the purpose of therapeutic vaccination in melanoma patients. Clin. Cancer Res. 15, 3366–3375 (2009).
Maus, M. V. et al. T cells expressing chimeric antigen receptors can cause anaphylaxis in humans. Cancer Immunol. Res. 1, 26–31 (2013).
Benteyn, D. et al. Characterization of CD8+ T-cell responses in the peripheral blood and skin injection sites of melanoma patients treated with mRNA electroporated autologous dendritic cells (TriMixDC-MEL). Biomed Res Int http://dx.doi.org/10.1155/2013/976383 (2013).
Lorenzi, J. C. et al. Intranasal vaccination with messenger RNA as a new approach in gene therapy: use against tuberculosis. BMC Biotechnol. 10, 77 (2010).
Wood, A. J. et al. Targeted genome editing across species using ZFNs and TALENs. Science 333, 307 (2011).
Davies, B. et al. Site specific mutation of the Zic2 locus by microinjection of TALEN mRNA in mouse CD1, C3H and C57BL/6J oocytes. PLoS ONE 8, e60216 (2013).
Karikó, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol Ther. 16, 1833–1840 (2008).
Angel, M. & Yanik, M. F. Innate immune suppression enables frequent transfection with RNA encoding reprogramming proteins. PLoS ONE 5, e11756 (2010).
Yakubov, E. et al. Reprogramming of human fibroblasts to pluripotent stem cells using mRNA of four transcription factors. Biochem. Biophys. Res. Commun. 394, 189–193 (2010).
Smull, C. E., Mallette, M. F. & Ludwig, E. H. The use of basic proteins to increase the infectivity of enterovirus ribonucleic acid. Biochem. Biophys. Res. Commun. 5, 247–249 (1961).
Papahadjopoulos, D. et al. Cochleate lipid cylinders: formation by fusion of unilamellar lipid vesicles. Biochim. Biophys. Acta 394, 483–491 (1975).
Dimitriadis, G. J. Translation of rabbit globin mRNA introduced by liposomes into mouse lymphocytes. Nature 274, 923–924 (1978).
Muthukrishnan, S., Both, G. W., Furuichi, Y. & Shatkin, A. J. 5′-Terminal 7-methylguanosine in eukaryotic mRNA is required for translation. Nature 255, 33–37 (1975).
Furuichi, Y. & Miura, K. A blocked structure at the 5′ terminus of mRNA from cytoplasmic polyhedrosis virus. Nature 253, 374–375 (1975).
Lockard, R. E. & Lingrel, J. B. The synthesis of mouse hemoglobin β-chains in a rabbit reticulocyte cell-free system programmed with mouse reticulocyte 9S RNA. Biochem. Biophys. Res. Commun. 37, 204–212 (1969).
Gurdon, J. B. et al. Use of frog eggs and oocytes for the study of messenger RNA and its translation in living cells. Nature. 233, 177–182 (1971).
Krieg, P. A. & Melton, D. A. Functional messenger RNAs are produced by SP6 in vitro transcription of cloned cDNAs. Nucl. Acids Res. 12, 7057–7070 (1984).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nature Biotech. 31, 227–229 (2013).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
U.S. is an inventor of multiple patents related to mRNA therapeutics. He is co-founder and CEO of the biotechnology company BioNTech (Mainz, Germany), which focuses on the development of mRNA therapies.
K.K. is an inventor of several patents related to mRNA technologies. She is an employee and the vice president of mRNA drug development in the BioNTech Incorporation (Mainz, Germany).
O.T. is an inventor of several patents related to mRNA technologies.
Related links
FURTHER INFORMATION
Rights and permissions
About this article
Cite this article
Sahin, U., Karikó, K. & Türeci, Ö. mRNA-based therapeutics — developing a new class of drugs. Nat Rev Drug Discov 13, 759–780 (2014). https://doi.org/10.1038/nrd4278
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd4278
This article is cited by
-
High-throughput barcoding of nanoparticles identifies cationic, degradable lipid-like materials for mRNA delivery to the lungs in female preclinical models
Nature Communications (2024)
-
Bioinformatic and literature assessment of toxicity and allergenicity of a CRISPR-Cas9 engineered gene drive to control Anopheles gambiae the mosquito vector of human malaria
Malaria Journal (2023)
-
Enhancement of bone regeneration by coadministration of angiogenic and osteogenic factors using messenger RNA
Inflammation and Regeneration (2023)
-
mRNA nanodelivery systems: targeting strategies and administration routes
Biomaterials Research (2023)
-
Oligonucleotide mapping via mass spectrometry to enable comprehensive primary structure characterization of an mRNA vaccine against SARS-CoV-2
Scientific Reports (2023)