Key Points
-
Sustained ventricular tachyarrhythmias are the most-common cause of sudden cardiac death (SCD), and traditional risk-stratification methods could be improved with genetic testing
-
Multiple genes affecting SCD risk in young patients have been discovered, but our understanding of the variability in disease severity is incomplete, and interpreting genetic test results is difficult
-
Next-generation sequencing in families that have inherited patterns of SCD, but are mutation-negative for the known disease genes, is expected to discover novel genes for these rare diseases
-
Genome-wide association studies (GWAS) have identified genomic loci that affect electrocardiogram indices, SCD risk in the general population, and risk of rare cardiac diseases such as Brugada syndrome
-
The unavailability of large, comprehensively phenotyped SCD cohorts has precluded the identification of genetic causes of SCD risk in patients with acquired cardiac disease
-
The mechanisms underlying loci identified through GWAS could involve functional regulatory elements in the noncoding region of the genome, and understanding them is important to understanding SCD
Abstract
Sudden cardiac death (SCD) resulting from ventricular tachyarrhythmia is a major contributor to mortality. Clinical management of SCD, currently based on clinical markers of SCD risk, can be improved by integrating genetic information. The identification of multiple disease-causing gene variants has already improved patient management and increased our understanding of the rare Mendelian diseases associated with SCD risk in the young, but marked variability in disease severity suggests that additional genetic modifiers exist. Next-generation DNA sequencing could be crucial to the discovery of SCD-associated genes, but large data sets can be difficult to interpret. SCD usually occurs in patients with an average age of 65 years who have complex cardiac disease stemming from multiple, common, acquired disorders. Heritable factors are largely unknown, but are likely to have a role in determining the risk of SCD in these patients. Numerous genetic loci have been identified that affect electrocardiogram indices, which are regarded as intermediate phenotypes for tachyarrhythmia. These loci could help to identify new molecules and pathways affecting cardiac electrical function. These loci are often located in intergenic regions, so our evolving understanding of the noncoding regulatory regions of the genome are likely to aid in the identification of novel genes that are important for cardiac electrical function and possibly SCD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
European Heart Rhythm Association et al. ACC/AHA/ESC 2006 guidelines for management of patients with ventricular arrhythmias and the prevention of sudden cardiac death: a report of the American College of Cardiology/American Heart Association Task Force and the European Society of Cardiology committee for practice guidelines. J. Am. Coll. Cardiol. 48, e247–e346 (2006).
de Vreede-Swagemakers, J. J. et al. Out-of-hospital cardiac arrest in the 1990's: a population-based study in the Maastricht area on incidence, characteristics and survival. J. Am. Coll. Cardiol. 30, 1500–1505 (1997).
Hulleman, M. et al. Implantable cardioverter-defibrillators have reduced the incidence of resuscitation for out-of-hospital cardiac arrest caused by lethal arrhythmias. Circulation 126, 815–821 (2012).
Olde Nordkamp, L. R. et al. The ICD for primary prevention in patients with inherited cardiac diseases: indications, use, and outcome: a comparison with secondary prevention. Circ. Arrhythm. Electrophysiol. 6, 91–100 (2013).
Kong, M. H. et al. Systematic review of the incidence of sudden cardiac death in the United States. J. Am. Coll. Cardiol. 57, 794–801 (2011).
US Department of Commerce. US and world population clock [online], (2013).
Cobb, L. A., Fahrenbruch, C. E., Olsufka, M. & Copass, M. K. Changing incidence of out-of-hospital ventricular fibrillation, 1980–2000. JAMA 288, 3008–3013 (2002).
Albert, C. M. et al. Prospective study of sudden cardiac death among women in the United States. Circulation 107, 2096–2101 (2003).
Myerburg, R. J., Kessler, K. M. & Castellanos, A. Sudden cardiac death: epidemiology, transient risk, and intervention assessment. Ann. Intern. Med. 119, 1187–1197 (1993).
Bardai, A. et al. Epilepsy is a risk factor for sudden cardiac arrest in the general population. PLoS ONE 7, e42749 (2012).
Huikuri, H. V., Castellanos, A. & Myerburg, R. J. Sudden death due to cardiac arrhythmias. N. Engl. J. Med. 345, 1473–1482 (2001).
Zheng, Z. J., Croft, J. B., Giles, W. H. & Mensah, G. A. Sudden cardiac death in the United States, 1989 to 1998. Circulation 104, 2158–2163 (2001).
Meyer, L. et al. Incidence, causes, and survival trends from cardiovascular-related sudden cardiac arrest in children and young adults 0 to 35 years of age: a 30-year review. Circulation 126, 1363–1372 (2012).
Corrado, D., Basso, C. & Thiene, G. Sudden cardiac death in young people with apparently normal heart. Cardiovasc. Res. 50, 399–408 (2001).
Tan, H. L., Hofman, N., van Langen, I. M., van der Wal, A. C. & Wilde, A. A. Sudden unexplained death: heritability and diagnostic yield of cardiological and genetic examination in surviving relatives. Circulation 112, 207–213 (2005).
George, A. L. Jr. Molecular and genetic basis of sudden cardiac death. J. Clin. Invest. 123, 75–83 (2013).
CARDIoGRAMplusC4D Consortium et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat. Genet. 45, 25–33 (2013).
Friedlander, Y. et al. Sudden death and myocardial infarction in first degree relatives as predictors of primary cardiac arrest. Atherosclerosis 162, 211–216 (2002).
Jouven, X., Desnos, M., Guerot, C. & Ducimetière, P. Predicting sudden death in the population: the Paris Prospective Study I. Circulation 99, 1978–1983 (1999).
Friedlander, Y. et al. Family history as a risk factor for primary cardiac arrest. Circulation 97, 155–160 (1998).
Dekker, L. R. et al. Familial sudden death is an important risk factor for primary ventricular fibrillation: a case-control study in acute myocardial infarction patients. Circulation 114, 1140–1145 (2006).
Kaikkonen, K. S., Kortelainen, M. L., Linna, E. & Huikuri, H. V. Family history and the risk of sudden cardiac death as a manifestation of an acute coronary event. Circulation 114, 1462–1467 (2006).
Myers, R. H., Kiely, D. K., Cupples, L. A. & Kannel, W. B. Parental history is an independent risk factor for coronary artery disease: the Framingham Study. Am. Heart J. 120, 963–969 (1990).
Stamler, R., Stamler, J., Riedlinger, W. F., Algera, G. & Roberts, R. H. Family (parental) history and prevalence of hypertension. Results of a nationwide screening program. JAMA 241, 43–46 (1979).
Meigs, J. B., Cupples, L. A. & Wilson, P. W. Parental transmission of type 2 diabetes: the Framingham Offspring Study. Diabetes 49, 2201–2207 (2000).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Bezzina, C. R. et al. Genome-wide association study identifies a susceptibility locus at 21q21 for ventricular fibrillation in acute myocardial infarction. Nat. Genet. 42, 688–691 (2010).
Arking, D. E. et al. Identification of a sudden cardiac death susceptibility locus at 2q24.2 through genome-wide association in European ancestry individuals. PLoS Genet. 7, e1002158 (2011).
Rizzo, S., Pilichou, K., Thiene, G. & Basso, C. The changing spectrum of arrhythmogenic (right ventricular) cardiomyopathy. Cell Tissue Res. 348, 319–323 (2012).
Watkins, H., Ashrafian, H. & Redwood, C. Inherited cardiomyopathies. N. Engl. J. Med. 364, 1643–1656 (2011).
Basso, C., Corrado, D., Marcus, F. I., Nava, A. & Thiene, G. Arrhythmogenic right ventricular cardiomyopathy. Lancet 373, 1289–1300 (2009).
Varnava, A. M. et al. Hypertrophic cardiomyopathy: histopathological features of sudden death in cardiac troponin T disease. Circulation 104, 1380–1384 (2001).
Schober, T. et al. Myofilament Ca sensitization increases cytosolic Ca binding affinity, alters intracellular Ca homeostasis, and causes pause-dependent Ca-triggered arrhythmia. Circ. Res. 111, 170–179 (2012).
Rizzo, S. et al. Intercalated disc abnormalities, reduced Na+ current density, and conduction slowing in desmoglein-2 mutant mice prior to cardiomyopathic changes. Cardiovasc. Res. 95, 409–418 (2012).
Cerrone, M. et al. Sodium current deficit and arrhythmogenesis in a murine model of plakophilin-2 haploinsufficiency. Cardiovasc. Res. 95, 460–468 (2012).
Wang, Q. et al. SCN5A mutations associated with an inherited cardiac arrhythmia, long QT syndrome. Cell 80, 805–811 (1995).
Curran, M. E. et al. A molecular basis for cardiac arrhythmia: HERG mutations cause long QT syndrome. Cell 80, 795–803 (1995).
Schwartz, P. J. et al. Long QT syndrome patients with mutations of the SCN5A and HERG genes have differential responses to Na+ channel blockade and to increases in heart rate: implications for gene-specific therapy. Circulation 92, 3381–3386 (1995).
Wang, D. W., Yazawa, K., Makita, N., George, A. L. & Bennett, P. B. Pharmacological targeting of long QT mutant sodium channels. J. Clin. Invest. 99, 1714–1720 (1997).
Vincent, G. M. et al. High efficacy of beta-blockers in long-QT syndrome type 1: contribution of noncompliance and QT-prolonging drugs to the occurrence of beta-blocker treatment “failures”. Circulation 119, 215–221 (2009).
Barsheshet, A. et al. Mutations in cytoplasmic loops of the KCNQ1 channel and the risk of life-threatening events: implications for mutation-specific response to β-blocker therapy in type 1 long-QT syndrome. Circulation 125, 1988–1996 (2012).
Heijman, J. et al. Dominant-negative control of cAMP-dependent IKs upregulation in human long-QT syndrome type 1. Circ. Res. 110, 211–219 (2012).
Choveau, F. S. et al. KCNQ1 channels voltage dependence through a voltage-dependent binding of the S4–S5 linker to the pore domain. J. Biol. Chem. 286, 707–716 (2011).
Watanabe, H. et al. Flecainide prevents catecholaminergic polymorphic ventricular tachycardia in mice and humans. Nat. Med. 15, 380–383 (2009).
van der Werf, C. et al. Flecainide therapy reduces exercise-induced ventricular arrhythmias in patients with catecholaminergic polymorphic ventricular tachycardia. J. Am. Coll. Cardiol. 57, 2244–2254 (2011).
Ackerman, M. J. et al. HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies: this document was developed as a partnership between the Heart Rhythm Society (HRS) and the European Heart Rhythm Association (EHRA). Europace 13, 1077–1109 (2011).
Wilde, A. A. & Behr, E. R. Genetic testing for inherited cardiac disease. Nat. Rev. Cardiol. 10, 571–583 (2013).
Genomes Project Consortium. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Genomes Project Consortium. 1000 Genomes [online], (2012).
Tennessen, J. A. et al. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 337, 64–69 (2012).
NHLBI GO Exome Sequencing Project (ESP). Exome Variant Server [online], (2013).
Jabbari, J. et al. New exome data question the pathogenicity of genetic variants previously associated with catecholaminergic polymorphic ventricular tachycardia. Circ. Cardiovasc. Genet. 6, 481–489 (2013).
Risgaard, B. et al. High prevalence of genetic variants previously associated with Brugada syndrome in new exome data. Clin. Genet. 84, 489–495 (2013).
Rehm, H. L. Disease-targeted sequencing: a cornerstone in the clinic. Nat. Rev. Genet. 14, 295–300 (2013).
Cooper, G. M. & Shendure, J. Needles in stacks of needles: finding disease-causal variants in a wealth of genomic data. Nat. Rev. Genet. 12, 628–640 (2011).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Ware, J. S., Walsh, R., Cunningham, F., Birney, E. & Cook, S. A. Paralogous annotation of disease-causing variants in long QT syndrome genes. Hum. Mutat. 33, 1188–1191 (2012).
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Katsanis, S. H. & Katsanis, N. Molecular genetic testing and the future of clinical genomics. Nat. Rev. Genet. 14, 415–426 (2013).
Boycott, K. M., Vanstone, M. R., Bulman, D. E. & Mackenzie, A. E. Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat. Rev. Genet. 14, 681–691 (2013).
Crotti, L. et al. Calmodulin mutations associated with recurrent cardiac arrest in infants. Circulation 127, 1009–1017 (2013).
Lynch, M. Rate, molecular spectrum, and consequences of human mutation. Proc. Natl. Acad. Sci. USA 107, 961–968 (2010).
Marsman, R. F. et al. A mutation in CALM1 encoding calmodulin in familial idiopathic ventricular fibrillation in childhood and adolescence. J. Am. Coll. Cardiol. http://dx.doi.org/10.1016/j.jacc.2013.07.091.
Boczek, N. J. et al. Exome sequencing and systems biology converge to identify novel mutations in the L-type calcium channel, CACNA1C, linked to autosomal dominant long QT syndrome. Circ. Cardiovasc. Genet. 6, 279–289 (2013).
Schwartz, P. J. in Sudden death (eds Kulbertus, H. E. & Wellens, H. J. J.) 358–378 (Martinus Nijhoff Publishers, 1980).
Bezzina, C. et al. A single Na+ channel mutation causing both long-QT and Brugada syndromes. Circ. Res. 85, 1206–1213 (1999).
Crotti, L. et al. NOS1AP is a genetic modifier of the long-QT syndrome. Circulation 120, 1657–1663 (2009).
Priori, S. G., Napolitano, C. & Schwartz, P. J. Low penetrance in the long-QT syndrome: clinical impact. Circulation 99, 529–533 (1999).
Priori, S. G. et al. Clinical and genetic heterogeneity of right bundle branch block and ST-segment elevation syndrome: a prospective evaluation of 52 families. Circulation 102, 2509–2515 (2000).
Brink, P. A. et al. Phenotypic variability and unusual clinical severity of congenital long-QT syndrome in a founder population. Circulation 112, 2602–2610 (2005).
Priori, S. G. et al. Risk stratification in the long-QT syndrome. N. Engl. J. Med. 348, 1866–1874 (2003).
Postema, P. G. et al. Founder mutations in the Netherlands: familial idiopathic ventricular fibrillation and DPP6. Neth. Heart J. 19, 290–296 (2011).
Schwartz, P. J. et al. Neural control of heart rate is an arrhythmia risk modifier in long QT syndrome. J. Am. Coll. Cardiol. 51, 920–929 (2008).
Roden, D. M. Drug-induced prolongation of the QT interval. N. Engl. J. Med. 350, 1013–1022 (2004).
Schwartz, P. J., Priori, S. G. & Napolitano, C. How really rare are rare diseases? The intriguing case of independent compound mutations in the long QT syndrome. J. Cardiovasc. Electrophysiol. 14, 1120–1121 (2003).
Westenskow, P., Splawski, I., Timothy, K. W., Keating, M. T. & Sanguinetti, M. C. Compound mutations: a common cause of severe long-QT syndrome. Circulation 109, 1834–1841 (2004).
Giudicessi, J. R. & Ackerman, M. J. Prevalence and potential genetic determinants of sensorineural deafness in KCNQ1 homozygosity and compound heterozygosity. Circ. Cardiovasc. Genet. 6, 193–200 (2013).
Kapplinger, J. D. et al. Spectrum and prevalence of mutations from the first 2,500 consecutive unrelated patients referred for the FAMILION long QT syndrome genetic test. Heart Rhythm 6, 1297–1303 (2009).
Moss, A. J. et al. Clinical aspects of type-1 long-QT syndrome by location, coding type, and biophysical function of mutations involving the KCNQ1 gene. Circulation 115, 2481–2489 (2007).
Shimizu, W. et al. Genotype-phenotype aspects of type 2 long QT syndrome. J. Am. Coll. Cardiol. 54, 2052–2062 (2009).
Tomás, M. et al. Polymorphisms in the NOS1AP gene modulate QT interval duration and risk of arrhythmias in the long QT syndrome. J. Am. Coll. Cardiol. 55, 2745–2752 (2010).
Bezzina, C. R. et al. Common variants at SCN5A-SCN10A and HEY2 are associated with Brugada syndrome, a rare disease with high risk of sudden cardiac death. Nat. Genet. 9, 1044–1049 (2013).
Duchatelet, S. et al. Identification of a KCNQ1 polymorphism acting as a protective modifier against arrhythmic risk in long-QT syndrome. Circ. Cardiovasc. Genet. 6, 354–361 (2013).
Arking, D. E. et al. A common genetic variant in the NOS1 regulator NOS1AP modulates cardiac repolarization. Nat. Genet. 38, 644–651 (2006).
Pfeufer, A. et al. Common variants at ten loci modulate the QT interval duration in the QTSCD study. Nat. Genet. 41, 407–414 (2009).
Newton-Cheh, C. et al. Common variants at ten loci influence QT interval duration in the QTGEN study. Nat. Genet. 41, 399–406 (2009).
Amin, A. S. et al. Variants in the 3′ untranslated region of the KCNQ1-encoded Kv7.1 potassium channel modify disease severity in patients with type 1 long QT syndrome in an allele-specific manner. Eur. Heart J. 33, 714–723 (2012).
Serre, D. et al. Differential allelic expression in the human genome: a robust approach to identify genetic and epigenetic cis-acting mechanisms regulating gene expression. PLoS Genet. 4, e1000006 (2008).
Mizusawa, Y. & Wilde, A. A. Brugada syndrome. Circ. Arrhythm. Electrophysiol. 5, 606–616 (2012).
Crotti, L. et al. Spectrum and prevalence of mutations involving BrS1- through BrS12-susceptibility genes in a cohort of unrelated patients referred for Brugada syndrome genetic testing: implications for genetic testing. J. Am. Coll. Cardiol. 60, 1410–1418 (2012).
Chambers, J. C. et al. Genetic variation in SCN10A influences cardiac conduction. Nat. Genet. 42, 149–152 (2010).
Holm, H. et al. Several common variants modulate heart rate, PR interval and QRS duration. Nat. Genet. 42, 117–122 (2010).
Pfeufer, A. et al. Genome-wide association study of PR interval. Nat. Genet. 42, 153–159 (2010).
Sotoodehnia, N. et al. Common variants in 22 loci are associated with QRS duration and cardiac ventricular conduction. Nat. Genet. 42, 1068–1076 (2010).
Fischer, A. & Gessler, M. Hey genes in cardiovascular development. Trends Cardiovasc. Med. 13, 221–226 (2003).
Albert, C. M. et al. Common variants in cardiac ion channel genes are associated with sudden cardiac death. Circ. Arrhythm. Electrophysiol. 3, 222–229 (2010).
Westaway, S. K. et al. Common variants in CASQ2, GPD1L, and NOS1AP are significantly associated with risk of sudden death in patients with coronary artery disease. Circ. Cardiovasc. Genet. 4, 397–402 (2011).
Stecker, E. C. et al. Allelic variants of SCN5A and risk of sudden cardiac arrest in patients with coronary artery disease. Heart Rhythm 3, 697–700 (2006).
Burke, A. et al. Role of SCN5A Y1102 polymorphism in sudden cardiac death in blacks. Circulation 112, 798–802 (2005).
Splawski, I. et al. Variant of SCN5A sodium channel implicated in risk of cardiac arrhythmia. Science 297, 1333–1336 (2002).
Albert, C. M. et al. Cardiac sodium channel gene variants and sudden cardiac death in women. Circulation 117, 16–23 (2008).
Crotti, L. et al. Torsades de pointes following acute myocardial infarction: evidence for a deadly link with a common genetic variant. Heart Rhythm 9, 1104–1112 (2012).
Sotoodehnia, N. et al. β2-Adrenergic receptor genetic variants and risk of sudden cardiac death. Circulation 113, 1842–1848 (2006).
Tseng, Z. H. et al. Common β-adrenergic receptor polymorphisms are not associated with risk of sudden cardiac death in patients with coronary artery disease. Heart Rhythm 5, 814–821 (2008).
Gavin, M. C. et al. A common variant in the β2-adrenergic receptor and risk of sudden cardiac death. Heart Rhythm 8, 704–710 (2011).
Mikkelsson, J., Perola, M., Penttilä, A. & Karhunen, P. J. Platelet glycoprotein Ibα HPA-2 Met/VNTR B haplotype as a genetic predictor of myocardial infarction and sudden cardiac death. Circulation 104, 876–880 (2001).
Mikkelsson, J., Perola, M., Laippala, P., Penttilä, A. & Karhunen, P. J. Glycoprotein IIIa Pl(A1/A2) polymorphism and sudden cardiac death. J. Am. Coll. Cardiol. 36, 1317–1323 (2000).
Sotoodehnia, N. et al. Genetic variation in angiotensin-converting enzyme-related pathways associated with sudden cardiac arrest risk. Heart Rhythm 6, 1306–1314 (2009).
Huertas-Vazquez, A. et al. A common missense variant in the neuregulin1 gene is associated with both schizophrenia and sudden cardiac death. Heart Rhythm 10, 994–998 (2013).
Newton-Cheh, C. et al. A common variant at 9p21 is associated with sudden and arrhythmic cardiac death. Circulation 120, 2062–2068 (2009).
Voight, B. F. et al. The metabochip, a custom genotyping array for genetic studies of metabolic, cardiovascular, and anthropometric traits. PLoS Genet. 8, e1002793 (2012).
Huertas-Vazquez, A. et al. Novel loci associated with increased risk of sudden cardiac death in the context of coronary artery disease. PLoS ONE 8, e59905 (2013).
Arking, D. E. et al. Genome-wide association study identifies GPC5 as a novel genetic locus protective against sudden cardiac arrest. PLoS ONE 5, e9879 (2010).
Aouizerat, B. E. et al. GWAS for discovery and replication of genetic loci associated with sudden cardiac arrest in patients with coronary artery disease. BMC Cardiovasc. Disord. 11, 29 (2011).
Noutsias, M. et al. Human coxsackie-adenovirus receptor is colocalized with integrins αvβ3 and αvβ5 on the cardiomyocyte sarcolemma and upregulated in dilated cardiomyopathy: implications for cardiotropic viral infections. Circulation 104, 275–280 (2001).
Lim, B. et al. Coxsackievirus and adenovirus receptor (CAR) mediates atrioventricular-node function and connexin 45 localization in the murine heart. J. Clin. Invest. 118, 2758–2770 (2008).
Lisewski, U. et al. The tight junction protein CAR regulates cardiac conduction and cell-cell communication. J. Exp. Med. 205, 2369–2379 (2008).
Bugert, P. et al. No evidence for an association between the rs2824292 variant at chromosome 21q21 and ventricular fibrillation during acute myocardial infarction in a German population. Clin. Chem. Lab. Med. 49, 1237–1239 (2011).
Park, J. H. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat. Genet. 42, 570–575 (2010).
Kolder, I. C., Tanck, M. W. & Bezzina, C. R. Common genetic variation modulating cardiac ECG parameters and susceptibility to sudden cardiac death. J. Mol. Cell. Cardiol. 52, 620–629 (2012).
Straus, S. M. et al. Prolonged QTc interval and risk of sudden cardiac death in a population of older adults. J. Am. Coll. Cardiol. 47, 362–367 (2006).
Schwartz, P. J. & Wolf, S. QT interval prolongation as predictor of sudden death in patients with myocardial infarction. Circulation 57, 1074–1077 (1978).
Chugh, S. S. et al. Determinants of prolonged QT interval and their contribution to sudden death risk in coronary artery disease: the Oregon Sudden Unexpected Death Study. Circulation 119, 663–670 (2009).
Kashani, A. & Barold, S. S. Significance of QRS complex duration in patients with heart failure. J. Am. Coll. Cardiol. 46, 2183–2192 (2005).
Zimetbaum, P. J. et al. Electrocardiographic predictors of arrhythmic death and total mortality in the Multicenter Unsustained Tachycardia Trial. Circulation 110, 766–769 (2004).
Singh, J. P. et al. Heritability of heart rate variability: the Framingham Heart Study. Circulation 99, 2251–2254 (1999).
Friedlander, Y., Lapidos, T., Sinnreich, R. & Kark, J. D. Genetic and environmental sources of QT interval variability in Israeli families: the Kibbutz Settlements Family Study. Clin. Genet. 56, 200–209 (1999).
den Hoed, M. et al. Identification of heart rate-associated loci and their effects on cardiac conduction and rhythm disorders. Nat. Genet. 45, 621–631 (2013).
Cho, Y. S. et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat. Genet. 41, 527–534 (2009).
Eijgelsheim, M. et al. Genome-wide association analysis identifies multiple loci related to resting heart rate. Hum. Mol. Genet. 19, 3885–3894 (2010).
Nolte, I. M. et al. Common genetic variation near the phospholamban gene is associated with cardiac repolarisation: meta-analysis of three genome-wide association studies. PLoS ONE 4, e6138 (2009).
Smith, J. G. et al. Genome-wide association studies of the PR interval in African Americans. PLoS Genet. 7, e1001304 (2011).
Marroni, F. et al. A genome-wide association scan of RR and QT interval duration in 3 European genetically isolated populations: the EUROSPAN project. Circ. Cardiovasc. Genet. 2, 322–328 (2009).
Kao, W. H. et al. Genetic variations in nitric oxide synthase 1 adaptor protein are associated with sudden cardiac death in US white community-based populations. Circulation 119, 940–951 (2009).
Noseworthy, P. A. et al. Common genetic variants, QT interval, and sudden cardiac death in a Finnish population-based study. Circ. Cardiovasc. Genet. 4, 305–311 (2011).
Pazoki, R. et al. SNPs identified as modulators of ECG traits in the general population do not markedly affect ECG traits during acute myocardial infarction nor ventricular fibrillation risk in this condition. PLoS ONE 8, e57216 (2013).
Ritchie, M. D. et al. Genome- and phenome-wide analyses of cardiac conduction identifies markers of arrhythmia risk. Circulation 127, 1377–1385 (2013).
Janse, M. J. & Wit, A. L. Electrophysiological mechanisms of ventricular arrhythmias resulting from myocardial ischemia and infarction. Physiol. Rev. 69, 1049–1169 (1989).
Nattel, S., Maguy, A., Le Bouter, S. & Yeh, Y. H. Arrhythmogenic ion-channel remodeling in the heart: heart failure, myocardial infarction, and atrial fibrillation. Physiol. Rev. 87, 425–456 (2007).
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Freedman, M. et al. Principles for the post-GWAS functional characterisation of cancer risk loci. Nat. Genet. 43, 513–518 (2011).
Kathiresan, S. & Srivastava, D. Genetics of human cardiovascular disease. Cell 148, 1242–1257 (2012).
ENCODE Project Consortium. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
May, D. et al. Large-scale discovery of enhancers from human heart tissue. Nat. Genet. 44, 89–93 (2012).
He, A., Kong, S. W., Ma, Q. & Pu, W. T. Co-occupancy by multiple cardiac transcription factors identifies transcriptional enhancers active in heart. Proc. Natl. Acad. Sci. USA 108, 5632–5637 (2011).
van den Boogaard, M. et al. Genetic variation in T-box binding element functionally affects SCN5A/SCN10A enhancer. J. Clin. Invest. 122, 2519–2530 (2012).
Arnolds, D. E. et al. TBX5 drives Scn5a expression to regulate cardiac conduction system function. J. Clin. Invest. 122, 2509–2518 (2012).
Akopian, A. N., Sivilotti, L. & Wood, J. N. A tetrodotoxin-resistant voltage-gated sodium channel expressed by sensory neurons. Nature 379, 257–262 (1996).
Verkerk, A. O. et al. Functional Nav1.8 channels in intracardiac neurons: the link between SCN10A and cardiac electrophysiology. Circ. Res. 111, 333–343 (2012).
Yang, T. et al. Blocking Scn10a channels in heart reduces late sodium current and is antiarrhythmic. Circ. Res. 111, 322–332 (2012).
Pallante, B. A. et al. Contactin-2 expression in the cardiac Purkinje fiber network. Circ. Arrhythm. Electrophysiol. 3, 186–194 (2010).
Holm, H. et al. A rare variant in MYH6 is associated with high risk of sick sinus syndrome. Nat. Genet. 43, 316–320 (2011).
Nature Publishing Group. ENCODE [online], (2013).
Genomes Project Consortium. 1000 Genomes Browser [online], (2013).
Berdowski, J. et al. Impact of onsite or dispatched automated external defibrillator use on survival after out-of-hospital cardiac arrest. Circulation 124, 2225–2232 (2011).
Kleinjan, D. & van Heyningen, V. Long-range control of gene expression: emerging mechanisms and disruption in disease. Am. J. Hum. Genet. 76, 8–32 (2005).
Wang, Q. et al. Positional cloning of a novel potassium channel gene: KVLQT1 mutations cause cardiac arrhythmias. Nat. Genet. 12, 17–23 (1996).
Acknowledgements
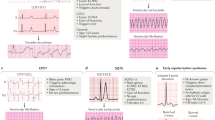
The authors are supported by grants from the Netherlands Organization for Scientific Research (NWO, grant ZonMW Vici 918.86.616 to H. L. Tan), the Netherlands Heart Foundation (NHS2007B202 and NHS2007B010 to C. R. Bezzina), the Center for Translational Molecular Medicine (CTMM-COHFAR project, C. R. Bezzina) and the CardioVascular Onderzoek Nederland and Netherlands Heart Foundation project PREDICT (C. R. Bezzina and H. L. Tan). We thank Yuka Mizusawa for the providing the electrocardiograms in Figure 3.
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, discussed its content, and wrote, edited, and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Marsman, R., Tan, H. & Bezzina, C. Genetics of sudden cardiac death caused by ventricular arrhythmias. Nat Rev Cardiol 11, 96–111 (2014). https://doi.org/10.1038/nrcardio.2013.186
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrcardio.2013.186
This article is cited by
-
Epidemiologie des Kreislaufstillstands in Europa
Notfall + Rettungsmedizin (2021)
-
Barbaloin inhibits ventricular arrhythmias in rabbits by modulating voltage-gated ion channels
Acta Pharmacologica Sinica (2018)
-
Using iPSC Models to Probe Regulation of Cardiac Ion Channel Function
Current Cardiology Reports (2018)
-
Association of common genetic variants related to atrial fibrillation and the risk of ventricular fibrillation in the setting of first ST-elevation myocardial infarction
BMC Medical Genetics (2017)
-
Common lipid features of lethal ventricular tarchyarrhythmias (LVTAs) induced by myocardial infarction and myocardial ion channel diseases
Scientific Reports (2017)