Key Points
-
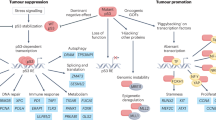
The tumour suppressor p53 induces cell death by apoptosis in response to various stress conditions, such as oncogene activation or DNA damage.
-
The loss of p53 tumour-suppressor activity — either by mutation/deletion of the TP53 gene or by inhibition of the p53 protein — favours the development of cancer.
-
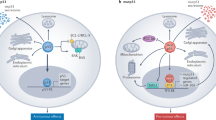
The MDM2 protein is a negative regulator of p53. After binding to p53, it inhibits its transcriptional activity, favours its nuclear export and stimulates its degradation.
-
The overexpression of MDM2 in various tumours inhibits p53, therefore favouring uncontrolled cell proliferation.
-
The inhibition of the p53–MDM2 interaction is an attractive strategy to activate p53-mediated apoptosis in tumours with overexpressed MDM2, but wild-type p53.
-
Several low-molecular-weight compounds and peptides that inhibit the p53–MDM2 interaction have been obtained. The peptidic inhibitors show an antiproliferative effect in tumour cells overexpressing MDM2.
Abstract
p53 is an attractive therapeutic target in oncology because its tumour-suppressor activity can be stimulated to eradicate tumour cells. Inhibiting the p53–MDM2 interaction is a promising approach for activating p53, because this association is well characterized at the structural and biological levels. MDM2 inhibits p53 transcriptional activity, favours its nuclear export and stimulates its degradation, so inhibiting the p53–MDM2 interaction with synthetic molecules should lead to p53-mediated cell-cycle arrest or apoptosis in p53-positive stressed cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Oren, M. Regulation of the p53 tumor suppressor protein. J. Biol. Chem. 274, 36031–36034 (1999).
Vousden, K. H. & Lu, X. Live or let die: the cell's response to p53. Nature Rev. Cancer 2, 594–604 (2002).
Donehower, L. A. et al. Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 356, 215–221 (1992).
Hainaut, P. & Hollstein, M. p53 and human cancer: the first ten thousand mutations. Adv. Cancer Res. 77, 81–137 (2000).
Kirsch, D. G. & Kastan, M. B. Tumor-suppressor p53: implications for tumor development and prognosis. J. Clin. Oncol. 16, 3158–3168 (1998).
Blagosklonny, M. V. p53: an ubiquitous target for anticancer drugs. Int. J. Cancer 98, 161–166 (2002).
Chene, P. Targeting p53 in cancer. Curr. Med. Chem. Anticancer Agents 1, 151–161 (2001).
Komarova, E. A. & Gudkov, A. V. Suppression of p53: a new approach to overcome side effects of antitumor therapy. Biochemistry 65, 41–48 (2000).
Picksley, S. M. & Lane, D. P. The p53–MDM2 autoregulatory feedback loop: a paradigm for the regulation of growth control by p53? Bioessays 15, 689–690 (1993).
Wu, X., Bayle, J. H., Olson, D. & Levine, A. J. The p53–MDM2 autoregulatory feedback loop. Genes Dev. 7, 1126–1132 (1993).
Barak, Y., Juven, T., Haffner, R. & Oren, M. Mdm2 expression is induced by wild type p53 activity. EMBO J. 12, 461–468 (1993).
Chen, J., Marechal, V. & Levine, A. J. Mapping of the p53 and Mdm-2 interaction domains. Mol. Cell. Biol. 13, 4107–4114 (1993).
Momand, J., Zambetti, G. P., Olson, D. C., George, D. & Levine, A. J. The Mdm-2 oncogene product forms a complex with the p53 protein and inhibits p53-mediated transactivation. Cell 69, 1237–1245 (1992).
Roth, J., Dobbelstein, M., Freedman, D. A., Shenk, T. & Levine, A. J. Nucleo-cytoplasmic shuttling of the HDM2 oncoprotein regulates the levels of the p53 protein via a pathway used by the human immunodeficiency virus rev protein. EMBO J. 17, 554–564 (1998).
Tao, W. & Levine, A. J. Nucleocytoplasmic shuttling of oncoprotein Hdm2 is required for Hdm2-mediated degradation of p53. Proc. Natl Acad. Sci. USA 96, 3077–3080 (1999).
Honda, R., Tanaka, H. & Yasuda, H. Oncoprotein Mdm2 is ubiquitin ligase E3 for tumour suppressor p53. FEBS Lett. 420, 25–27 (1997).
Haupt, Y., Maya, R., Kazaz, A. & Oren, M. Mdm2 promotes the rapid degradation of p53. Nature 387, 296–299 (1997).
Kubbutat, M. H., Jones, S. N. & Vousden, K. H. Regulation of p53 stability by Mdm2. Nature 387, 299–303 (1997).
Jones, S. N., Roe, A. E., Donehower, L. A. & Bradley, A. Rescue of embryonic lethality in Mdm2-deficient mice by absence of p53. Nature 378, 206–208 (1995).
Montes de Oca Luna, R., Wagner, D. S. & Lozano, G. Rescue of early embryonic lethality in Mdm2-deficient mice by deletion of p53. Nature 378, 203–206 (1995).
Chen, J., Wu, X., Lin, J. & Levine, A. J. Mdm-2 inhibits the G1 arrest and apoptosis function of the p53 tumor suppressor protein. Mol. Cell. Biol. 16, 2445–2452 (1996).
Schon, O., Friedler, A., Bycroft, M., Freund, S. M. V. & Fersht, A. R. Molecular mechanism of the interaction between Mdm2 and p53. J. Mol. Biol. 323, 491–501 (2002).
Jabbur, J. R. et al. Mdm-2 binding at TAFII31 recruitment is regulated by hydrogen bond disruption between the p53 residues Thr18 and Asp21. Oncogene 21, 7100–7113 (2002).
Shieh, S. Y., Ahn, J., Tamai, K., Taya, Y. & Prives, C. The human homologs of checkpoint kinases Chk1 and Cds1 (Chk2) phosphorylate p53 at multiple DNA damage-inducible sites. Genes Dev. 14, 289–300 (2000).
Sakaguchi, K. et al. Damage-mediated phosphorylation of human p53 threonine 18 through a cascade mediated by a casein 1-like kinase. Effect on Mdm2 binding. J. Biol. Chem. 275, 9278–9283 (2000).
Hirao, A. et al. DNA damage-induced activation of p53 by the checkpoint kinase Chk2. Science 287, 1824–1827 (2000).
Chehab, N. H., Malikzay, A., Stavridi, E. S. & Halazonetis, T. D. Phosphorylation of Ser-20 mediates stabilization of human p53 in response to DNA damage. Proc. Natl Acad. Sci. USA 96, 13777–13782 (1999).
Eischen, M. D., Weber, J. D., Roussel, M. F., Sherr, C. J. & Cleveland, J. L. Disruption of the ARF–Mdm2–p53 tumor suppressor pathway in Myc-induced lymphomagenesis. Genes Dev. 13, 2658–2669 (1999).
Palmero, I., Pantoja, C. & Serrano, M. p19ARF links the tumour suppressor p53 to Ras. Nature 395, 125–126 (1998).
Momand, J., Jung, D., Wilczynski, S. & Niland, J. The MDM2 gene amplification database. Nucleic Acids Res. 26, 3453–3459 (1998).
Eymin, B., Gazzeri, S., Brambilla, C. & Brambilla, E. Mdm2 overexpression and p14ARF inactivation are two mutually exclusive events in primary human lung tumors. Oncogene 21, 2750–2761 (2002).
Polsky, D. et al. Hdm2 protein overexpression, but not amplification, is related to tumorigenesis of cutaneous melanoma. Cancer Res. 61, 7642–7646 (2001).
Leite, K. R. et al. Abnormal expression of Mdm2 in prostate carcinoma. Mod. Pathol. 14, 428–436 (2001).
Wang, H. et al. MDM2 oncogene as a target for cancer therapy: an antisense approach. Int. J. Oncol. 15, 653–660 (1999).
Oliner, J. D. et al. Oncoprotein Mdm2 conceals the activation domain of tumour suppressor p53. Nature 362, 857–860 (1993).
Lin, J., Chen, J., Elenbaas, B. & Levine, A. J. Several hydrophobic amino acids in the p53 amino-terminal domain are required for transcriptional activation, binding to Mdm-2 and the adenovirus 5 E1B 55-kD protein. Genes Dev. 8, 1235–1246 (1994).
Picksley, S. M., Vojtesek, B., Sparks, A. & Lane, D. P. Immunochemical analysis of the interaction of p53 with Mdm2: fine mapping of the mdm2 binding site on p53 using synthetic peptides. Oncogene 9, 2523–2529 (1994).
Lai, Z., Auger, K. R., Manubay, C. M. & Copeland, R. A. Thermodynamics of p53 binding to Hdm2(1-126): effects of phosphorylation and p53 peptide length. Arch. Biochem. Biophys. 381, 278–284 (2000).
Kussie, P. H. et al. Structure of the Mdm2 oncoprotein bound to the p53 tumor suppressor transactivation domain. Science 274, 948–953 (1996). Structure of the p53–MDM2 complex.
Uesugi, M. & Verdine, G. L. The α-helical FXXFF motif in p53: TAF interaction and discrimination by Mdm2. Proc. Natl Acad. Sci. USA 96, 14801–14806 (1999).
Blommers, M. J. J., Fendrich, G., Garcia-Echeverria, C. & Chene, P. On the interaction between p53 and Mdm2: transfer NOE study of p53-derived peptide ligated to Mdm2. J. Am. Chem. Soc. 119, 3425–3426 (1997).
Botuyan, M. V. E., Momand, J. & Chen, Y. Solution conformation of an essential region of the p53 transactivation domain. Fold Des. 2, 331–342 (1997).
Massova, I. & Kollman, P. A. Computational alanine scanning to probe protein–protein interactions: a novel approach to evaluate binding free energies. J. Am. Chem. Soc. 121, 8133–8143 (1999).
Jones, S. & Thornton, J. M. Principles of protein–protein interactions. Proc. Natl Acad. Sci. USA 93, 13–20 (1996).
Yang, A. et al. p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol. Cell 2, 305–316 (1998).
Kaghad, M. et al. Monoallelically expressed gene related to p53 at 1p36, a region frequently deleted in neuroblastoma and other human cancers. Cell 90, 809–819 (1997).
Shvarts, A. et al. Mdmx: a novel p53-binding protein with some functional properties of Mdm2. EMBO J. 15, 5349–5357 (1996).
Bartel, F., Taubert, H. & Harris, L. C. Alternative and aberrant splicing of Mdm2 mRNA in human cancer. Cancer Cell 2, 9–15 (2002).
Strano, S. et al. From p63 to p53 across p73. FEBS Lett. 490, 163–170 (2001).
Marin, C. & Kaelin, W. G. p63 and p73: old members of a new family. Biochim. Biophys. Acta 1470, M93–M100 (2000).
Michael, D. & Oren, M. The p53 and Mdm2 families. Curr Opin Genet Dev 12, 53–59 (2002).
Levrero, M. et al. The p53/p63/p73 family of transcription factors: overlapping and distinct functions. J. Cell Sci. 113, 1661–1670 (2000).
Yang, A. & McKeon, F. p63 and p73: p53 menaces and more. Nature Rev. Mol. Cell Biol. 1, 199–207 (2000).
Bottger, V. et al. Comparative study of the p53–Mdm2 and p53–MDMX interfaces. Oncogene 18, 189–199 (1999).
Kadakia, M., Slader, C. & Berberich, S. J. Regulation of p63 function by Mdm2 and Mdmx. DNA Cell Biol. 20, 321–330 (2001).
Ongkeko, W. M. et al. Mdm2 and Mdmx bind and stabilize the p53-related protein p73. Curr. Biol. 9, 829–832 (1999).
Kane, S. A. et al. Development of binding assay for p53/Hdm2 by using homogeneous time-resolved fluorescence. Anal. Biochem. 278, 29–38 (2000).
Zeng, X. et al. MDM2 suppresses p73 function without promoting p73 degradation. Mol. Cell. Biol. 19, 3257–3266 (1999).
Bottger, V. et al. Identification of novel Mdm2 binding peptides by phage display. Oncogene 13, 2141–2147 (1996).
Bottger, A. et al. Molecular characterization of the Hdm2–p53 interaction. J. Mol. Biol. 269, 744–756 (1997).
Banerjee, R., Basu, G., Chene, P. & Roy, S. Aib-based peptide backbone as scaffolds for helical peptide mimics. J. Pept. Res 60, 88–94 (2002).
Garcia-Echeverria, C., Chene, P., Blommers, M. J. & Furet, P. Discovery of potent antagonists of the interaction between human double minute 2 and tumour suppressor p53. J. Med. Chem. 43, 3205–3208 (2000). Design of the most potent peptidic inhibitors of the p53–MDM2 interaction.
Majeux, N., Scarsi, M. & Caflisch, A. Efficient electrostatic model for protein-fragment docking. Proteins 42, 256–268 (2001).
Duncan, S. J. et al. Isolation and structure elucidation of chlorofusin, a novel p53–Mdm2 antagonist from a Fusarium sp. J. Am. Chem. Soc. 123, 554–560 (2001).
Zhao, J. et al. The initial evaluation of non-peptidic small-molecule HDM2 inhibitors based on p53–HDM2 complex structure. Cancer Lett. 183, 69–77 (2002).
Stoll, R. et al. Chalcone derivatives antagonize interactions between the human oncoprotein MDM2 and p53. Biochemistry 40, 336–344 (2001). Identification and characterization of non-peptidic inhibitors of the p53–MDM2 interaction.
Bottger, A. et al. Design of a synthetic Mdm2-binding mini protein that activates the p53 response in vivo. Curr. Biol. 7, 860–869 (1997). Inhibition of the p53–MDM2 interaction in tumour cells with the TIP inhibitors.
Wasylyk, C. et al. p53 mediated death of cells overexpressing MDM2 by an inhibitor of MDM2 interaction with p53. Oncogene 18, 1921–1934 (1999). Inhibition of the p53–MDM2 interaction in tumour cells with the GST inhibitors.
Kanovsky, M. et al. Peptides from the amino terminal Mdm-2-binding domain of p53, designed from conformational analysis, are selectively cytotoxic to transformed cells. Proc. Natl Acad. Sci. USA 98, 12438–12443 (2001).
Chene, P., Fuchs, J., Carena, I., Furet, P. & Garcia Echeverria, C. Study of the cytotoxic effect of a peptidic inhibitor of the p53–Hdm2 interaction in tumour cells. FEBS Lett. 529, 293–297 (2002). Inhibition of the p53–MDM2 interaction in tumour cells with a peptide.
Chene, P. et al. A small synthetic peptide, which inhibits the p53–Hdm2 interaction, stimulates the p53 pathway in tumour cell lines. J. Mol. Biol. 299, 245–253 (2000).
Polyak, K., Waldman, T., He, T. C., Kinzler, K. W. & Vogelstein, B. Genetic determinants of p53-induced apoptosis and growth arrest. Genes Dev. 10, 1945–1952 (1996).
Blaydes, J. P. & Wynford-Thomas, D. The proliferation of normal human fibroblasts is dependent upon negative regulation of p53 function by Mdm2. Oncogene 16, 3317–3322 (1998).
Toogood, P. L. Inhibition of protein–protein association by small molecules: approaches and progress. J. Med. Chem. 45, 1543–1558 (2002).
Mecozzi, S. & Rebek, J. The 55% solution: a formula for molecular recognition in the liquid state. Chem. Eur. J. 4, 1016–1022 (1998).
Leveillard, T. & Wasylyk, B. The Mdm2 C-terminal region binds to TAFII250 and is required for Mdm2 regulation of cyclin A promoter. J. Biol. Chem. 272, 30651–30661 (1997).
Juven-Gershon, T. et al. The Mdm2 oncoprotein interacts with the cell fate regulator Numb. Mol. Cell. Biol. 18, 3974–3982 (1998).
Boyd, M. T., Vlatkovic, N. & Haines, D. S. A novel cellular protein (MTBP) binds to Mdm2 and induces a G1 arrest that is suppressed by Mdm2. J. Biol. Chem. 275, 31883–31890 (2000).
Author information
Authors and Affiliations
Related links
Related links
DATABASES
LocusLink
FURTHER INFORMATION
Glossary
- NUCLEAR MAGNETIC RESONANCE
-
(NMR). A technique that uses the magnetic properties of certain atomic nuclei (such as 1H, 13C and 15N) to determine the structure of the proteins.
- X-RAY CRYSTALLOGRAPHY
-
A technique that uses the diffraction of the X-rays to determine the structure of the proteins.
- PHAGE DISPLAY
-
A technology that is used for displaying a protein (or peptide) on the surface of a bacteriophage, which contains the gene(s) that encodes the displayed protein(s), thereby physically linking the genotype and phenotype.
- IC50
-
The concentration of an inhibitor that is required to inhibit 50% of the p53–MDM2 interaction.
- AIB
-
(α-amino isobutyric acid). A non-natural amino acid that is used to favour helical conformations in peptides.
- AC3C
-
(1-amino-cyclopropanecarboxylic acid). A non-natural amino acid that is used to stabilize 310-helix conformations in peptides.
- PMP
-
(Phosphonomethylphenylalanine). A tyrosine substituted at its hydroxyl group by a phosphonomethyl moiety.
- 6CLTRP
-
(6-chloro-tryptophan). Tryptophan with a chlorine at position-6 (corresponds to Cη2).
- ELISA
-
(Enzyme-linked immunosorbent assay). A solid-phase immunoassay that detects the interaction between proteins and specific antibodies.
- FACS
-
(Fluorescence-activated cell sorting). A technique that is used in flow cytometry to detect cells that are labelled with fluorescent dyes.
- TUNEL
-
(TdT-mediated dUTP-X nick-end labelling). A method that is used to measure DNA strand breaks during apoptosis.
- E6 PROTEIN
-
A viral oncoprotein that is derived from certain human papillomavirus types that are associated with increased risk of cervical cancer. E6 binds to and targets p53 for ubiquitin-mediated degradation.
Rights and permissions
About this article
Cite this article
Chène, P. Inhibiting the p53–MDM2 interaction: an important target for cancer therapy. Nat Rev Cancer 3, 102–109 (2003). https://doi.org/10.1038/nrc991
Issue Date:
DOI: https://doi.org/10.1038/nrc991
This article is cited by
-
p53 suppresses MHC class II presentation by intestinal epithelium to protect against radiation-induced gastrointestinal syndrome
Nature Communications (2024)
-
Association of TP53 rs1042522 G > C, MDM2 rs2279744 T > G, and miR-34b/c rs4938723 T > C polymorphisms with aneuploidy pregnancy susceptibility
BMC Pregnancy and Childbirth (2023)
-
Visualizing single-molecule conformational transition and binding dynamics of intrinsically disordered proteins
Nature Communications (2023)
-
Permission to pass: on the role of p53 as a gatekeeper for aneuploidy
Chromosome Research (2023)
-
Steady-State Analysis of p53 Protein Pathway Using DTMC and CTMC
Transactions of the Indian National Academy of Engineering (2023)