Abstract
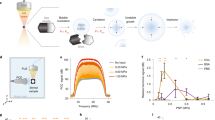
Gas vesicles (GVs) are a unique class of gas-filled protein nanostructures that are detectable at subnanomolar concentrations and whose physical properties allow them to serve as highly sensitive imaging agents for ultrasound and MRI. Here we provide a protocol for isolating GVs from native and heterologous host organisms, functionalizing these nanostructures with moieties for targeting and fluorescence, characterizing their biophysical properties and imaging them using ultrasound and MRI. GVs can be isolated from natural cyanobacterial and haloarchaeal host organisms or from Escherichia coli expressing a heterologous GV gene cluster and purified using buoyancy-assisted techniques. They can then be modified by replacing surface-bound proteins with engineered, heterologously expressed variants or through chemical conjugation, resulting in altered mechanical, surface and targeting properties. Pressurized absorbance spectroscopy is used to characterize their mechanical properties, whereas dynamic light scattering (DLS)and transmission electron microscopy (TEM) are used to determine nanoparticle size and morphology, respectively. GVs can then be imaged with ultrasound in vitro and in vivo using pulse sequences optimized for their detection versus background. They can also be imaged with hyperpolarized xenon MRI using chemical exchange saturation transfer between GV-bound and dissolved xenon—a technique currently implemented in vitro. Taking 3–8 d to prepare, these genetically encodable nanostructures enable multimodal, noninvasive biological imaging with high sensitivity and potential for molecular targeting.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Ferrara, K., Pollard, R. & Borden, M. Ultrasound microbubble contrast agents: fundamentals and application to gene and drug delivery. Annu. Rev. Biomed. Eng. 9, 415–447 (2007).
Kaufmann, B.A. & Lindner, J.R. Molecular imaging with targeted contrast ultrasound. Curr. Opin. Biotechnol. 18, 11–16 (2007).
Weissleder, R. et al. Ultrasmall superparamagnetic iron oxide: characterization of a new class of contrast agents for MR imaging. Radiology 175, 489–493 (1990).
Lee, J.-H. et al. Artificially engineered magnetic nanoparticles for ultra-sensitive molecular imaging. Nat. Med. 13, 95–99 (2007).
Caravan, P., Ellison, J.J., McMurry, T.J. & Lauffer, R.B. Gadolinium (III) chelates as MRI contrast agents: structure, dynamics, and applications. Chem. Rev. 99, 2293–2352 (1999).
Walsby, A.E. Gas vesicles. Microbiol. Rev. 58, 94–144 (1994).
Pfeifer, F. Distribution, formation and regulation of gas vesicles. Nat. Rev. Microbiol. 10, 705–715 (2012).
Shapiro, M.G. et al. Biogenic gas nanostructures as ultrasonic molecular reporters. Nat. Nanotechnol. 9, 311–316 (2014).
Maresca, D. et al. Nonlinear ultrasound imaging of nanoscale acoustic biomolecules. Appl. Phys. Lett. 110, 073704 (2017).
Cherin, E. et al. Acoustic behavior of Halobacterium salinarum gas vesicles in the high-frequency range: experiments and modeling. Ultrasound Med. Biol. 43, 1016–1030 (2017).
Shapiro, M.G. et al. Genetically encoded reporters for hyperpolarized xenon magnetic resonance imaging. Nat. Chem. 6, 629–634 (2014).
Hane, F.T. et al. In vivo detection of cucurbit[6]uril, a hyperpolarized xenon contrast agent for a xenon magnetic resonance imaging biosensor. Sci. Rep. 7, 41027 (2017).
Barskiy, D.A. et al. NMR hyperpolarization techniques of gases. Chem. Eur. J. 23, 725–751 (2017).
Lakshmanan, A. et al. Molecular engineering of acoustic protein nanostructures. ACS Nano 10, 7314–7322 (2016).
Li, N. & Cannon, M.C. Gas vesicle genes identified in Bacillus megaterium and functional expression in Escherichia coli. J. Bacteriol. 180, 2450–2458 (1998).
Gilad, A.A. & Shapiro, M.G. Molecular imaging in synthetic biology, and synthetic biology in molecular imaging. Mol. Imaging Biol. 19, 373–378 (2017).
Zakeri, B. et al. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. PNAS 109, E690–E697 (2012).
Schröder, L. Chapter 8 HyperCEST Imaging. In Chemical Exchange Saturation Transfer Imaging: Advances and Applications. pp 121–158 (Pan Stanford Publishing, 2017).
Schröder, L., Chapter 17 - Xenon Biosensor HyperCEST MRI A2 - Albert, Mitchell S. In Hyperpolarized xenon for NMR and MRI (ed. Hane, F.T.) pp 263–277 (Academic Press, Boston, 2017).
Witte, C. et al. Hyperpolarized xenon for NMR and MRI applications. JoVE 67, E4268 (2012).
Witte, C., Kunth, M., Rossella, F. & Schröder, L. Observing and preventing rubidium runaway in a direct-infusion xenon-spin hyperpolarizer optimized for high-resolution hyper-CEST (chemical exchange saturation transfer using hyperpolarized nuclei) NMR. J. Chem. Phys. 140, 084203 (2014).
Nikolaou, P. et al. Near-unity nuclear polarization with an open-source 129Xe hyperpolarizer for NMR and MRI. PNAS 110, 14150–14155 (2013).
Kunth, M., Witte, C., Hennig, A. & Schröder, L. Identification, classification, and signal amplification capabilities of high-turnover gas binding hosts in ultra-sensitive NMR. Chem. Sci. 6, 6069–6075 (2015).
Kunth, M., Witte, C. & Schröder, L. Continuous-wave saturation considerations for efficient xenon depolarization. NMR Biomed. 28, 601–606 (2015).
Smith, R. & Peat, A. Growth and gas-vacuole development in vegetative cells of Anabaena flos-aquae. Arch. Microbiol. 58, 117–126 (1967).
Buchholz, B., Hayes, P. & Walsby, A. The distribution of the outer gas vesicle protein, GvpC, on the Anabaena gas vesicle, and its ratio to GvpA. Microbiology 139, 2353–2363 (1993).
Simon, R.D. Morphology and protein composition of gas vesicles from wild type and gas vacuole defective strains of Halobacterium salinarium strain 5. Microbiology 125, 103–111 (1981).
Kunth, M., Witte, C. & Schröder, L. Quantitative chemical exchange saturation transfer with hyperpolarized nuclei (qHyper-CEST): sensing xenon-host exchange dynamics and binding affinities by NMR. J. Chem. Phys. 141, 194202 (2014).
Zaiss, M., Schnurr, M. & Bachert, P. Analytical solution for the depolarization of hyperpolarized nuclei by chemical exchange saturation transfer between free and encapsulated xenon (HyperCEST). J. Chem. Phys. 136, 144106 (2012).
Acknowledgements
This research was supported by the National Institutes of Health (R01-EB018975 to M.G.S.), DARPA (W911NF-14-1-0111 to M.G.S.), CIHR (grant no. FDN148367 to F.S.F.) and HFSP (RGP0050/2016 to M.G.S. and L.S.). A.L. is supported by an NSF graduate research fellowship (no. 1144469). A.F. is supported by an NSERC graduate fellowship. S.P.N. is supported by a Caltech Summer Undergraduate Research Fellowship. D. Maresca is supported by an HFSP Cross-Disciplinary Postdoctoral Fellowship. Research in the Shapiro laboratory is also supported by the Heritage Medical Research Institute, the Burroughs Wellcome Fund, the Pew Charitable Trust, the Sontag Foundation and the David and Lucile Packard Foundation. Research in the Schröder laboratory is also supported by the Michael J. Fox Foundation for Parkinson's Research (no. 12549) and a Koselleck Grant by the German Research Foundation (DFG; project SCHR 995/5-1). We acknowledge the Jensen Lab and the Beckman Resource Center for Electron Microscopy (EM) at the California Institute of Technology for technical guidance and for allowing us to use their resources. The EM facility is funded by the Beckman Foundation, the Gordon and Betty Moore Foundation, the Agouron Institute and the HHMI.
Author information
Authors and Affiliations
Contributions
A.L., G.J.L., A.F., S.P.N. and M.G.S. conceived the manuscript layout and coordinated its writing. A.L., G.J.L., A.F., S.P.N., A.L.-G., D. Maresca, D. Malounda, R.W.B. and M.G.S developed methods for GV production, functionalization, and characterization, as well as for ultrasound imaging. M.Y., J.Y. and F.S.F. contributed methods for in vivo ultrasound imaging of GVs. M.K., C.W., L.S. and M.G.S. contributed methods for hyperpolarized xenon MRI of GVs. All authors contributed to writing the manuscript. A.L. compiled and edited the protocol and G.J.L. assembled the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Ultrasound imaging of Mega GVs
B-mode ultrasound images of purified, wild-type Ana GVs (OD500,ps: 2.2) versus a purified and unclustered batch of Mega GVs at (a) equal molarity, (b) equal protein concentration and (c) equal gas fraction. Scale bars are 1 mm. Images were acquired using the Verasonics L22-14v transducer and the ray-lines script with the following parameters: transmit frequency: 18MHz, number of cycles of the transmitted pulse: 6, F number: 2, imaging voltage: 3V, with the transducer focus (8 mm depth) aligned close to the center of the sample well. Images were processed and analyzed using MATLAB. Images are shown before (top panel) and after collapse (bottom panel) using a high-power burst from the transducer at 25V for 10 s. (d) Quantification of ultrasound signal was performed by selecting a region of interest (ROI) of defined size within the sample well and calculating the mean intensity per pixel for the selected ROI, after post-collapse background subtraction (n=12 for Ana GVs, n=4 for Mega GVs at each condition shown in a, b and c; error bars are SEM).
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1. (PDF 306 kb)
Supplementary Data
MATLAB scripts. (ZIP 4 kb)
Rights and permissions
About this article
Cite this article
Lakshmanan, A., Lu, G., Farhadi, A. et al. Preparation of biogenic gas vesicle nanostructures for use as contrast agents for ultrasound and MRI. Nat Protoc 12, 2050–2080 (2017). https://doi.org/10.1038/nprot.2017.081
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.081
This article is cited by
-
Phase transition of GvpU regulates gas vesicle clustering in bacteria
Nature Microbiology (2024)
-
Magneto-acoustic protein nanostructures for non-invasive imaging of tissue mechanics in vivo
Nature Materials (2024)
-
Wireless agents for brain recording and stimulation modalities
Bioelectronic Medicine (2023)
-
Cell-cycle dependent nuclear gene delivery enhances the effects of E-cadherin against tumor invasion and metastasis
Signal Transduction and Targeted Therapy (2023)
-
Ultrasensitive ultrasound imaging of gene expression with signal unmixing
Nature Methods (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.