Abstract
Mitophagy is a cellular process that selectively removes damaged, old or dysfunctional mitochondria. Defective mitophagy is thought to contribute to normal aging and to various neurodegenerative and cardiovascular diseases. Previous methods used to detect mitophagy in vivo were cumbersome, insensitive and difficult to quantify. We created a transgenic mouse model that expresses the pH-dependent fluorescent protein mt-Keima in order to more readily assess mitophagy. Keima is a pH-sensitive, dual-excitation ratiometric fluorescent protein that also exhibits resistance to lysosomal proteases. At the physiological pH of the mitochondria (pH 8.0), the shorter-wavelength excitation predominates. Within the acidic lysosome (pH 4.5) after mitophagy, mt-Keima undergoes a gradual shift to longer-wavelength excitation. In this protocol, we describe how to monitor mitophagic flux in living cells over an 18-h time frame, as well as how to quantify mitophagy using the mt-Keima probe. This protocol also describes how to use confocal microscopy to visualize mitophagy in living tissues obtained from mt-Keima transgenic mice. With this protocol, the mt-Keima probe can reliably be imaged within the first 60 min after tissue collection. We also describe how to apply mt-Keima with stimulated emission depletion (STED) microscopy, which can potentially provide substantially higher-resolution images. Typically, the approximate time frame for time-lapse fluorescence imaging of mt-Keima is 20 h for living cells. For confocal analysis of tissue from an mt-Keima mouse, the whole procedure generally takes no longer than 60 min, and the STED imaging usually takes <2 h.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mishra, P. & Chan, D.C. Metabolic regulation of mitochondrial dynamics. J. Cell Biol. 212, 379–387 (2016).
Sun, N., Youle, R.J. & Finkel, T. The mitochondrial basis of aging. Mol. Cell 61, 654–666 (2016).
Youle, R.J. & Narendra, D.P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Bingol, B. et al. The mitochondrial deubiquitinase USP30 opposes Parkin-mediated mitophagy. Nature 510, 370–375 (2014).
Bingol, B. & Sheng, M. Mechanisms of mitophagy: PINK1, Parkin, USP30 and beyond. Free Radic. Biol. Med. 100, 210–222 (2016).
Narendra, D., Tanaka, A., Suen, D.F. & Youle, R.J. Parkin-induced mitophagy in the pathogenesis of Parkinson disease. Autophagy 5, 706–708 (2009).
Sun, N. et al. Measuring in vivo mitophagy. Mol. Cell 60, 685–696 (2015).
Gong, G. et al. Parkin-mediated mitophagy directs perinatal cardiac metabolic maturation in mice. Science 350, aad2459 (2015).
Fang, E.F. et al. NAD+ replenishment improves lifespan and healthspan in ataxia telangiectasia models via mitophagy and DNA repair. Cell Metab. 24, 566–581 (2016).
Cunningham, C.N. et al. USP30 and Parkin homeostatically regulate atypical ubiquitin chains on mitochondria. Nat. Cell Biol. 17, 160–169 (2015).
Liu, L. et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat. Cell Biol. 14, 177–185 (2012).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Klionsky, D.J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016).
Chu, C.T. et al. Cardiolipin externalization to the outer mitochondrial membrane acts as an elimination signal for mitophagy in neuronal cells. Nat. Cell Biol. 15, 1197–1205 (2013).
Kuma, A., Matsui, M. & Mizushima, N. LC3, an autophagosome marker, can be incorporated into protein aggregates independent of autophagy: caution in the interpretation of LC3 localization. Autophagy 3, 323–328 (2007).
Katayama, H., Kogure, T., Mizushima, N., Yoshimori, T. & Miyawaki, A. A sensitive and quantitative technique for detecting autophagic events based on lysosomal delivery. Chem. Biol. 18, 1042–1052 (2011).
Rosado, C.J., Mijaljica, D., Hatzinisiriou, I., Prescott, M. & Devenish, R.J. Rosella: a fluorescent pH-biosensor for reporting vacuolar turnover of cytosol and organelles in yeast. Autophagy 4, 205–213 (2008).
Chan, D.C. Fusion and fission: interlinked processes critical for mitochondrial health. Annu. Rev. Genet. 46, 265–287 (2012).
Westermann, B. Mitochondrial fusion and fission in cell life and death. Nat. Rev. Mol. Cell Biol. 11, 872–884 (2010).
Chen, H. & Chan, D.C. Mitochondrial dynamics—fusion, fission, movement, and mitophagy—in neurodegenerative diseases. Hum. Mol. Genet. 18, R169–176 (2009).
Frey, T.G. & Mannella, C.A. The internal structure of mitochondria. Trends Biochem. Sci. 25, 319–324 (2000).
Jakobs, S. & Wurm, C.A. Super-resolution microscopy of mitochondria. Curr. Opin. Chem. Biol. 20, 9–15 (2014).
McWilliams, T.G. et al. mito-QC illuminates mitophagy and mitochondrial architecture in vivo. J. Cell Biol. 214, 333–345 (2016).
Koval, M. & Pagano, R.E. Sorting of an internalized plasma membrane lipid between recycling and degradative pathways in normal and Niemann-Pick, type A fibroblasts. J. Cell Biol. 111, 429–442 (1990).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Acknowledgements
We are grateful to Z.-X. Yu (NHLBI Pathology Core) and P. McCoy (NHLBI FACS Core) for experimental assistance. We thank R. Youle for the Parkin-overexpressing HeLa cell line, and A. Miyawaki for the original mt-Keima construct. This work was supported by Intramural NIH funds and a Leducq Transatlantic Network grant to T.F.
Author information
Authors and Affiliations
Contributions
N.S., D.M. and J.L. performed the experiments; N.S., D.M., J.L., I.I.R. and C.A.C. analyzed the data; and N.S., D.M. and T.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Pixel intensity map of the mt-Keima signals in HeLa cells shown in time-lapse analysis of cellular mitophagic flux (figure 2a-c).
(a) Corresponding pixel intensity map for time-lapse analysis of a HeLa cell line expressing mt-Keima without treatment for 0, 2 and 18 hours as in figure 2a; (b) Corresponding pixel intensity map for time-lapse analysis of a HeLa cell line expressing mt-Keima subjected to hypoxia for 0, 2 and 18 hours as in figure 2b; (c) Corresponding pixel intensity map for time-lapse analysis of a HeLa cells expressing mt-Keima and Parkin treated with FCCP (5 μM) and Oligomycin (5 μM) for 0, 2 and 18 hours in figure 2c. Crosshairs are set as described in Step 19. Mitophagy is quantified as the number of pixels in quadrant 2 (red pixels) divided by the number of total pixels [quadrant 1+2+3+4-B].
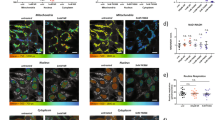
Supplementary Figure 2 mt-Keima signal in the hippocampal region of mice.
(a) High power confocal image of the dentate gyrus in the mt-Keima mouse (Scale bar = 10 μm; color coding is as in Figure 2); (b) Region selected from the SGZ for analysis, and (c) corresponding pixel intensity map.
Supplementary Figure 3 mt-Keima signal in the cerebellum region of mice.
(a) High power confocal mage from the cerebellum of the mt-Keima mouse (Scale bar = 10 μm; color coding is as in Figure 2); (b) Purkinje cells selected for analysis, and (c) corresponding pixel intensity map.
Supplementary information
Combo PDF
Supplementary Figures 1–3. (PDF 898 kb)
Confocal time-lapse of a HeLa cell line with stable expression of mt-Keima without treatment.
Images were obtained at 15-min intervals over an 18-h period. There is only a modest level of mitophagy observed, indicating little evidence of laser-induced photo-damage. (MP4 2924 kb)
Confocal time-lapse of a HeLa cell line with stable expression of mt-Keima and Parkin treated with FCCP (5 μM) and oligomycin (5 μM).
Images were obtained at 15-min intervals over an 18-h period. An increase in mitophagy begins early, and this treatment eventually results in the near total loss of green mt-Keima fluorescence and a marked increase in overall red mt-Keima fluorescence. (MP4 2066 kb)
STED time-lapse of mt-Keima mouse liver imaged ex vivo at 6,000 lines per second for 7 min with a 480-nm excitation laser.
Mapped pseudocolor was used to assess hepatocyte mitochondria morphology and dynamics. Little evidence of STED-induced photo-bleaching is observed. (MP4 3584 kb)
Rights and permissions
About this article
Cite this article
Sun, N., Malide, D., Liu, J. et al. A fluorescence-based imaging method to measure in vitro and in vivo mitophagy using mt-Keima. Nat Protoc 12, 1576–1587 (2017). https://doi.org/10.1038/nprot.2017.060
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.060
This article is cited by
-
NME3 is a gatekeeper for DRP1-dependent mitophagy in hypoxia
Nature Communications (2024)
-
The mitophagy pathway and its implications in human diseases
Signal Transduction and Targeted Therapy (2023)
-
PPTC7 maintains mitochondrial protein content by suppressing receptor-mediated mitophagy
Nature Communications (2023)
-
TREM2hi resident macrophages protect the septic heart by maintaining cardiomyocyte homeostasis
Nature Metabolism (2023)
-
Colorimetric and fluorescence dual-mode pH sensor based on nitrogen-doped carbon dots and its diverse applications
Microchimica Acta (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.