Abstract
Proteomic characterization of blood plasma is of central importance to clinical proteomics and particularly to biomarker discovery studies. The vast dynamic range and high complexity of the plasma proteome have, however, proven to be serious challenges and have often led to unacceptable tradeoffs between depth of coverage and sample throughput. We present an optimized sample-processing pipeline for analysis of the human plasma proteome that provides greatly increased depth of detection, improved quantitative precision and much higher sample analysis throughput as compared with prior methods. The process includes abundant protein depletion, isobaric labeling at the peptide level for multiplexed relative quantification and ultra-high-performance liquid chromatography coupled to accurate-mass, high-resolution tandem mass spectrometry analysis of peptides fractionated off-line by basic pH reversed-phase (bRP) chromatography. The overall reproducibility of the process, including immunoaffinity depletion, is high, with a process replicate coefficient of variation (CV) of <12%. Using isobaric tags for relative and absolute quantitation (iTRAQ) 4-plex, >4,500 proteins are detected and quantified per patient sample on average, with two or more peptides per protein and starting from as little as 200 μl of plasma. The approach can be multiplexed up to 10-plex using tandem mass tags (TMT) reagents, further increasing throughput, albeit with some decrease in the number of proteins quantified. In addition, we provide a rapid protocol for analysis of nonfractionated depleted plasma samples analyzed in 10-plex. This provides ∼600 quantified proteins for each of the ten samples in ∼5 h of instrument time.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Anderson, N.L. & Anderson, N.G. The human plasma proteome: history, character, and diagnostic prospects. Mol. Cell. Proteomics 1, 845–867 (2002).
Hortin, G.L., Jortani, S.A., Ritchie, J.C. Jr., Valdes, R. Jr. & Chan, D.W. Proteomics: a new diagnostic frontier. Clin. Chem. 52, 1218–1222 (2006).
Pernemalm, M. & Lehtio, J. Mass spectrometry-based plasma proteomics: state of the art and future outlook. Expert Rev. Proteomics 11, 431–448 (2014).
States, D.J. et al. Challenges in deriving high-confidence protein identifications from data gathered by a HUPO plasma proteome collaborative study. Nat. Biotechnol. 24, 333–338 (2006).
Farrah, T. et al. A high-confidence human plasma proteome reference set with estimated concentrations in PeptideAtlas. Mol. Cell. Proteomics 10, M110.006353 (2011).
Farrah, T. et al. State of the human proteome in 2013 as viewed through PeptideAtlas: comparing the kidney, urine, and plasma proteomes for the biology- and disease-driven Human Proteome Project. J. Proteome Res. 13, 60–75 (2014).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol. Cell. Proteomics 11, M111.014050 (2012).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Mertins, P. et al. Ischemia in tumors induces early and sustained phosphorylation changes in stress kinase pathways but does not affect global protein levels. Mol. Cell. Proteomics 13, 1690–1704 (2014).
Hortin, G.L., Sviridov, D. & Anderson, N.L. High-abundance polypeptides of the human plasma proteome comprising the top 4 logs of polypeptide abundance. Clin. Chem. 54, 1608–1616 (2008).
Pieper, R. et al. Multi-component immunoaffinity subtraction chromatography: an innovative step towards a comprehensive survey of the human plasma proteome. Proteomics 3, 422–432 (2003).
Qian, W.J. et al. Enhanced detection of low abundance human plasma proteins using a tandem IgY12-SuperMix immunoaffinity separation strategy. Mol. Cell. Proteomics 7, 1963–1973 (2008).
Shi, T. et al. IgY14 and SuperMix immunoaffinity separations coupled with liquid chromatography-mass spectrometry for human plasma proteomics biomarker discovery. Methods 56, 246–253 (2012).
Cao, Z., Tang, H.Y., Wang, H., Liu, Q. & Speicher, D.W. Systematic comparison of fractionation methods for in-depth analysis of plasma proteomes. J. Proteome Res. 11, 3090–3100 (2012).
Song, C. et al. Reversed-phase-reversed-phase liquid chromatography approach with high orthogonality for multidimensional separation of phosphopeptides. Anal. Chem. 82, 53–56 (2010).
Wang, Y. et al. Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human MCF10A cells. Proteomics 11, 2019–2026 (2011).
Cominetti, O. et al. Proteomic biomarker discovery in 1000 human plasma samples with mass spectrometry. J. Proteome Res. 15, 389–399 (2016).
Dayon, L., Nunez Galindo, A., Corthesy, J., Cominetti, O. & Kussmann, M. Comprehensive and scalable highly automated MS-based proteomic workflow for clinical biomarker discovery in human plasma. J. Proteome Res. http://dx.doi.org/10.1021/pr500635f (2014).
Addona, T.A. et al. A pipeline that integrates the discovery and verification of plasma protein biomarkers reveals candidate markers for cardiovascular disease. Nat. Biotechnol. 29, 635–643 (2011).
Huttenhain, R. et al. Reproducible quantification of cancer-associated proteins in body fluids using targeted proteomics. Sci. Transl. Med. 4, 142ra194 (2012).
Whiteaker, J.R. et al. A targeted proteomics-based pipeline for verification of biomarkers in plasma. Nat. Biotechnol. 29, 625–634 (2011).
Bantscheff, M. et al. Robust and sensitive iTRAQ quantification on an LTQ Orbitrap mass spectrometer. Mol. Cell. Proteomics 7, 1702–1713 (2008).
Erickson, B.K. et al. Evaluating multiplexed quantitative phosphopeptide analysis on a hybrid quadrupole mass filter/linear ion trap/orbitrap mass spectrometer. Anal. Chem. 87, 1241–1249 (2015).
Gan, C.S., Chong, P.K., Pham, T.K. & Wright, P.C. Technical, experimental, and biological variations in isobaric tags for relative and absolute quantitation (iTRAQ). J. Proteome Res. 6, 821–827 (2007).
McAlister, G.C. et al. Increasing the multiplexing capacity of TMTs using reporter ion isotopologues with isobaric masses. Anal. Chem. 84, 7469–7478 (2012).
McAlister, G.C. et al. Analysis of the acidic proteome with negative electron-transfer dissociation mass spectrometry. Anal. Chem. 84, 2875–2882 (2012).
Mertins, P. et al. iTRAQ labeling is superior to mTRAQ for quantitative global proteomics and phosphoproteomics. Mol. Cell. Proteomics 11, M111.014423 (2012).
Nilsson, C.L. et al. Quantitative phosphoproteomic analysis of the STAT3/IL-6/HIF1alpha signaling network: an initial study in GSC11 glioblastoma stem cells. J. Proteome Res. 9, 430–443 (2010).
Ow, S.Y. et al. iTRAQ underestimation in simple and complex mixtures: 'The Good, the Bad and the Ugly'. J. Proteome Res. 8, 5347–5355 (2009).
Rauniyar, N. & Yates, J.R. III Isobaric labeling-based relative quantification in shotgun proteomics. J. Proteome Res. 13, 5293–5309 (2014).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Savitski, M.M. et al. Measuring and managing ratio compression for accurate iTRAQ/TMT quantification. J. Proteome Res. 12, 3586–3598 (2013).
Ting, L., Rad, R., Gygi, S.P. & Haas, W. MS3 eliminates ratio distortion in isobaric multiplexed quantitative proteomics. Nat. Methods 8, 937–940 (2011).
Cole, R.N. et al. The plasma proteome identifies expected and novel proteins correlated with micronutrient status in undernourished Nepalese children. J. Nutr. 143, 1540–1548 (2013).
Jones, K.A. et al. Immunodepletion plasma proteomics by tripleTOF 5600 and Orbitrap elite/LTQ-Orbitrap Velos/Q exactive mass spectrometers. J. Proteome Res. 12, 4351–4365 (2013).
Burgess, M.W., Keshishian, H., Mani, D.R., Gillette, M.A. & Carr, S.A. Simplified and efficient quantification of low-abundance proteins at very high multiplex via targeted mass spectrometry. Mol. Cell. Proteomics 13, 1137–1149 (2014).
Keshishian, H., Addona, T., Burgess, M., Kuhn, E. & Carr, S.A. Quantitative, multiplexed assays for low abundance proteins in plasma by targeted mass spectrometry and stable isotope dilution. Mol. Cell. Proteomics 6, 2212–2229 (2007).
Keshishian, H. et al. Multiplexed, quantitative workflow for sensitive biomarker discovery in plasma yields novel candidates for early myocardial injury. Mol. Cell. Proteomics 14, 2375–2393 (2015).
Swearingen, K.E. & Moritz, R.L. High-field asymmetric waveform ion mobility spectrometry for mass spectrometry-based proteomics. Expert Rev. Proteomics 9, 505–517 (2012).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Shadforth, I.P., Dunkley, T.P.J., Lilley, K.S. & Bessant, C. i-Tracker: for quantitative proteomics using iTRAQ (TM). BMC Genomics 6, 145 (2005).
Wenger, C.D. et al. Gas-phase purification enables accurate, multiplexed proteome quantification with isobaric tagging. Nat. Methods 8, 933–935 (2011).
Patel, B.B. et al. Assessment of two immunodepletion methods: off-target effects and variations in immunodepletion efficiency may confound plasma proteomics. J. Proteome Res. 11, 5947–5958 (2012).
Pichler, P. et al. Peptide labeling with isobaric tags yields higher identification rates using iTRAQ 4-plex compared to TMT 6-plex and iTRAQ 8-plex on LTQ Orbitrap. Anal. Chem. 82, 6549–6558 (2010).
Acknowledgements
We thank N. Udeshi for reading the manuscript and providing valuable feedback. This work was supported in part by grants from the National Institutes of Health: HHSN268201000033C and R01HL096738 from the National Heart, Lung, and Blood Institute (NHLBI; to S.A.C.) and grants U24CA160034 from the National Cancer Institute (NCI) Clinical Proteomics Tumor Analysis Consortium initiative and U01CA152990 from the NCI Early Detection Research Network program (to M.A.G.).
Author information
Authors and Affiliations
Contributions
H.K., M.W.B., H.S., K.R.C., M.A.G. and S.A.C. developed the protocol. L.W., M.W.B. and H.S. optimized and ran QC checks on many aspects of the protocol. H.K., M.W.B., H.S., K.R.C., M.A.G. and S.A.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Plumbing schematic for IgY14-Supermix tandem depletion columns on an Agilent 1200 LC system.
Diagram illustrates plumbing the second valve for tandem setup of 2 depletion columns. A 2mL needle seat extension has been incorporated to accommodate injections of more than 900μL on the Agilent 1200 system.
Supplementary Figure 2 Chromatograms of plasma after tandem IgY14-Supermix depletion demonstrating column performance over the vendor recommended lifetime.
Injection number 7(blue), 49 (red) and 100 (green) are overlaid showing a reduction in peak area of the IgY14 bound peak with a concomitant increase and change in shape of the IgY14-Supermix depleted peak. In addition, an early peak (1) is also observed by injection 49 increasing over the remainder of the column lifetime. We estimate that area ratio of IgY14-Supemix depleted peak to IgY14 bound peak should be around 0.06-0.07 for effective depletion. As column ages this ratio increases and ratio of more than 0.1 indicates problematic depletion.
Supplementary Figure 3 Summary statistics in four PMI patient samples (Adapted from (38)).
Table gives details about peptide and protein identification and quantitation in each of the patient sample. Venn diagram shows the overlap in between the four patient samples. aProteins identified in at least two patients with two or more peptides. bSubset of identified proteins with two or more distinct peptides observed in at least one patient. cProtein subgroups (groups); that is, 5304 distinct protein subgroups were identified within 4555 protein groups. Proteins that share a detected distinct peptide (length > 8) are combined into a group. A protein group is parsimoniously expanded to one or more subgroups to distinguish proteins that also have one or more distinct peptides that are not shared with the rest of the group, typically isoforms and family members.
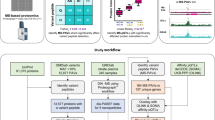
Supplementary Figure 4 Timelines and throughput for (A) plasma sample processing and (B) LC-MS/MS data collection.
Times for each processing step are illustrated for deep plasma profiling employing iTRAQ4-plex and TMT10-plex modes with off-line basic reversed phase peptide fractionation, as well as for single shot (no off-line fractionation) analysis of peptides labeled with TMT10. Times shown are those required for a single person manually processing samples. All sample processing steps can be further parallelized, increasing the throughput, making LC-MS/MS analysis on a single instrument the rate-limiting step of the deep profiling workflow (B). In contrast, in the single shot analysis workflow 4 different plexes (16 – 40 samples, depending on reagent used for labeling) can be analyzed on a single LC-MS/MS instrument in a single day making sample processing the limiting factor for this workflow.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4. (PDF 615 kb)
Rights and permissions
About this article
Cite this article
Keshishian, H., Burgess, M., Specht, H. et al. Quantitative, multiplexed workflow for deep analysis of human blood plasma and biomarker discovery by mass spectrometry. Nat Protoc 12, 1683–1701 (2017). https://doi.org/10.1038/nprot.2017.054
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.054
This article is cited by
-
Deep proteomic analysis of obstetric antiphospholipid syndrome by DIA-MS of extracellular vesicle enriched fractions
Communications Biology (2024)
-
Differential Expression of Fibrinogen Alpha and Its Potential Involvement in Osteoarthritis Pathogenesis
Molecular Biotechnology (2024)
-
The salivary proteome in relation to oral mucositis in autologous hematopoietic stem cell transplantation recipients: a labelled and label-free proteomics approach
BMC Oral Health (2023)
-
Data-independent acquisition-based mass spectrometry(DIA-MS) for quantitative analysis of patients with chronic hepatitis B
Proteome Science (2023)
-
The protein corona from nanomedicine to environmental science
Nature Reviews Materials (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.