Abstract
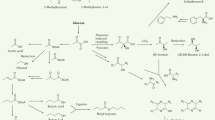
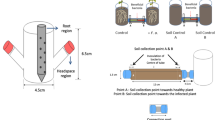
Airborne chemical signals emitted by bacteria influence the behavior of other bacteria and plants. We present an overview of in vitro methods for evaluating bacterial and plant responses to bacterial volatile compounds (BVCs). Three types of equipment have been used to physically separate the bacterial test strains from either other bacterial strains or plants (in our laboratory we use either Arabidopsis or tobacco plant seedlings): a Petri dish containing two compartments (BI Petri dish); two Petri dishes connected with tubing; and a microtiter-based assay. The optimized procedure for the BI Petri dish system is described in this protocol and can be widely used for elucidation of potential function in interactions between diverse microbes and those plant and chemical volatiles emitted by bacteria that are most likely to mediate bacterial or plant responses to BVCs. We also describe a procedure for metabolome-based BVC profiling via dynamic (i.e., continuous airflow) or static headspace sampling using solid-phase microextraction (SPME). Using both these procedures, bacteria–bacteria communications and bacteria–plant interactions mediated by BVCs can be rapidly investigated (within 1–4 weeks).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Farag, M.A., Zhang, H. & Ryu, C.-M. Dynamic chemical communication between plants and bacteria through airborne signals: induced resistance by bacterial volatiles. J. Chem. Ecol. 39, 1007–1018 (2013).
Kloepper, J.W., Leong, J., Teintze, M. & Schroth, M.N. Enhanced plant growth by siderophores produced by plant growth-promoting rhizobacteria. Nature 286, 885–886 (1980).
Ryu, C.M. et al. Bacterial volatiles promote growth in Arabidopsis. Proc. Natl. Acad. Sci. USA 100, 4927–4932 (2003).
Ryu, C.-M., Hu, C.-H., Locy, R.D. & Kloepper, J.W. Study of mechanisms for plant growth promotion elicited by rhizobacteria in Arabidopsis thaliana. Plant Soil 268, 285–292 (2005).
Ryu, C.-M., Kim, J., Choi, O., Kim, S.H. & Park, C.S. Improvement of biological control capacity of Paenibacillus polymyxa E681 by seed pelleting on sesame. Biol. Control 39, 282–289 (2006).
Ryu, C.M., Murphy, J.F., Mysore, K.S. & Kloepper, J.W. Plant growth-promoting rhizobacteria systemically protect Arabidopsis thaliana against Cucumber mosaic virus by a salicylic acid and NPR1-independent and jasmonic acid-dependent signaling pathway. Plant J. 39, 381–392 (2004).
Kloepper, J.W. Plant growth-promoting rhizobacteria as biological control agents. in Soil Microbial Ecology-Applications in Agricultural and Environmental Management (ed., Metting, F.B. Jr.), pp. 255–274 (Marcel Dekker, New York, 1993).
Kloeppe, J. et al. Plant root-bacterial interactions in biological control of soilborne diseases and potential extension to systemic and foliar diseases. Austr. Plant Pathol. 28, 21–26 (1999).
Ryu, C.-M. et al. Bacterial volatiles induce systemic resistance in Arabidopsis. Plant Physiol. 134, 1017–1026 (2004).
Bate, N.J. & Rothstein, S.J. C6-volatiles derived from the lipoxygenase pathway induce a subset of defenserelated genes. Plant J. 16, 561–569 (1998).
Banchio, E., Xie, X., Zhang, H. & Pare, P.W. Soil bacteria elevate essential oil accumulation and emissions in sweet basil. J. Agric. Food Chem. 57, 653–657 (2009).
Blom, D. et al. Production of plant growth modulating volatiles is widespread among rhizosphere bacteria and strongly depends on culture conditions. Environ. Microbiol. 13, 3047–3058 (2011).
Cho, S.M. et al. 2R, 3R-butanediol, a bacterial volatile produced by Pseudomonas chlororaphis O6, is involved in induction of systemic tolerance to drought in Arabidopsis thaliana. Mol. Plant Microbe Interact. 21, 1067–1075 (2008).
Han, S.H. et al. GacS-dependent production of 2R, 3R-butanediol by Pseudomonas chlororaphis O6 is a major determinant for eliciting systemic resistance against Erwinia carotovora but not against Pseudomonas syringae pv. tabaci in tobacco. Mol. Plant Microbe Interact. 19, 924–930 (2006).
Kai, M. et al. Serratia odorifera: analysis of volatile emission and biological impact of volatile compounds on Arabidopsis thaliana. Appl. Microbiol. Biotechnol. 88, 965–976 (2010).
Rudrappa, T. et al. The rhizobacterial elicitor acetoin induces systemic resistance in Arabidopsis thaliana. Commun. Integr. Biol. 3, 130 (2010).
Wenke, K., Kai, M. & Piechulla, B. Belowground volatiles facilitate interactions between plant roots and soil organisms. Planta 231, 499–506 (2010).
Zhang, H. et al. Rhizobacterial volatile emissions regulate auxin homeostasis and cell expansion in Arabidopsis. Planta 226, 839–851 (2007).
Zhang, H. et al. Soil bacteria confer plant salt tolerance by tissue-specific regulation of the sodium transporter HKT1. Mol. Plant Microbe Interact. 21, 737–744 (2008).
Zhang, H. et al. Choline and osmotic-stress tolerance induced in Arabidopsis by the soil microbe Bacillus subtilis (GB03). Mol. Plant Microbe Interact. 23, 1097–1104 (2010).
Zhang, H. et al. A soil bacterium regulates plant acquisition of iron via deficiencyinducible mechanisms. Plant J. 58, 568–577 (2009).
Zhang, H. et al. Soil bacteria augment Arabidopsis photosynthesis by decreasing glucose sensing and abscisic acid levels in planta. Plant J. 56, 264–273 (2008).
Kim, K.-S., Lee, S. & Ryu, C.-M. Interspecific bacterial sensing through airborne signals modulates locomotion and drug resistance. Nat. Commun. 4, 1809 (2013).
Létoffé, S., Audrain, B., Bernier, S.P., Delepierre, M. & Ghigo, J.-M. Aerial exposure to the bacterial volatile compound trimethylamine modifies antibiotic resistance of physically separated bacteria by raising culture medium pH. MBio 5, e00944-13 (2014).
Bernier, S.P., Létoffé, S., Delepierre, M. & Ghigo, J.M. Biogenic ammonia modifies antibiotic resistance at a distance in physically separated bacteria. Mol. Microbiol. 81, 705–716 (2011).
Bitas, V., Kim, H.-S., Bennett, J.W. & Kang, S. Sniffing on microbes: diverse roles of microbial volatile organic compounds in plant health. Mol. Plant Microbe Interact. 26, 835–843 (2013).
Bailly, A. & Weisskopf, L. The modulating effect of bacterial volatiles on plant growth. Plant Signal. Behav. 7, 79–85 (2012).
D'Alessandro, M. et al. Volatiles produced by soil-borne endophytic bacteria increase plant pathogen resistance and affect tritrophic interactions. Plant Cell Environ. 37, 813–826 (2014).
Kai, M. et al. Bacterial volatiles and their action potential. Appl. Microbiol. Biotechnol. 81, 1001–1012 (2009).
Kwon, Y.S. et al. Proteome analysis of Arabidopsis seedlings exposed to bacterial volatiles. Planta 232, 1355–1370 (2010).
Zamioudis, C., Mastranesti, P., Dhonukshe, P., Blilou, I. & Pieterse, C.M. Unraveling root developmental programs initiated by beneficial Pseudomonas spp. bacteria. Plant Physiol. 162, 304–318 (2013).
Kai, M.J.D., Stanforth, S.P. & Dean, J.R. Identification of volatile organic compounds produced by bacteria using HS-SPME-GC–MS. J. Chromatogr. Sci. 52, 363–373 (2014).
Goupry, S., Rochut, N., Robins, R. & Gentil, E. Evaluation of solid-phase microextraction for the isotopic analysis of volatile compounds produced during fermentation by lactic acid bacteria. J. Agric. Food Chem. 48, 2222–2227 (2000).
Risticevic, S., Lord, H., Górecki, T., Arthur, C.L. & Pawliszyn, J. Protocol for solid-phase microextraction method development. Nat. Protoc. 5, 122–139 (2010).
Reaves, M.L. & Rabinowitz, J.D. Metabolomics in systems microbiology. Curr. Opin. Biotechnol. 22, 17–25 (2011).
Halket, J.M. et al. Deconvolution gas chromatography/mass spectrometry of urinary organic acids–potential for pattern recognition and automated identification of metabolic disorders. Rapid Commun. Mass Spectrom. 13, 279–284 (1999).
Broeckling, C.D., Reddy, I.R., Duran, A.L., Zhao, X. & Sumner, L.W. MET-IDEA: data extraction tool for mass spectrometry-based metabolomics. Anal. Chem. 78, 4334–4341 (2006).
Lei, Z., Li, H., Chang, J., Zhao, P.X. & Sumner, L.W. MET-IDEA version 2.06; improved efficiency and additional functions for mass spectrometry-based metabolomics data processing. Metabolomics 8, 105–110 (2012).
Farag, M.A., Ryu, C.-M., Sumner, L.W. & Paré, P.W. GC–MS SPME profiling of rhizobacterial volatiles reveals prospective inducers of growth promotion and induced systemic resistance in plants. Phytochemistry 67, 2262–2268 (2006).
Farag, M.A. & Wessjohann, L.A. Volatiles profiling in medicinal licorice roots using steam distillation and solidphase microextraction (SPME) coupled to chemometrics. J. Food Sci. 77, C1179–C1184 (2012).
Song, G.C., Chio, H.K. & Ryu, C.-M. The folate precursor para-aminobenzoic acid elicits induced resistance against Cucumber mosaic virus and Xanthomonas axonopodis. Ann. Bot. 111, 925–34 (2013).
Song, G.C. & Ryu, C.-M. Two volatile organic compounds trigger plant self-defense against a bacterial pathogen and a sucking insect in cucumber under open field conditions. Int. J. Mol. Sci. 14, 9803–9819 (2013).
Mondello, L., Tranchida, P.Q., Dugo, P. & Dugo, G. Comprehensive two-dimensional gas chromatography-mass spectrometry: a review. Mass Spectrom. Rev. 27, 101–124 (2008).
Lisec, J., Schauer, N., Kopka, J., Willmitzer, L. & Fernie, A.R. Gas chromatography mass spectrometry–based metabolite profiling in plants. Nat. Protoc. 1, 387–396 (2006).
Veriotti, T. & Sacks, R. High-speed characterization and analysis of orange oils with tandem-column stop-flow GC and time-of-flight MS. Anal. Chem. 74, 5635–5640 (2002).
Ramos, H.C. et al. Fermentative metabolism of Bacillus subtilis: physiology and regulation of gene expression. J. Bac. 182, 3072–3080 (2000).
Park, H., Lee, B., Kloepper, J.W. & Ryu, C.M. One shot-two pathogens blocked: exposure of Arabidopsis to hexadecane, a long chain volatile organic compound, confers induced resistance against both Pectobacterium carotovorum and Pseudomonas syringae. Plant Signal. Behav. 8, e24619 (2013).
Bailly, A. & Weisskopf, L. The modulating effect of bacterial volatiles on plant growth: current knowledge and future challenges. Plant Signal. Behav. 7, 79–85 (2012).
Dolch, M. et al. Volatile compound profiling for the identification of Gram-negative bacteria by ion-molecule reaction–mass spectrometry. J. Appl. Microbiol. 113, 1097–1105 (2012).
Kunze, N. et al. Detection and validation of volatile metabolic patterns over different strains of two human pathogenic bacteria during their growth in a complex medium using multi-capillary column-ion mobility spectrometry (MCC-IMS). Appl. Microbiol. Biotechnol. 97, 3665–3676 (2013).
Broekaert, K., Noseda, B., Heyndrickx, M., Vlaemynck, G. & Devlieghere, F. Volatile compounds associated with Psychrobacter spp. and Pseudoalteromonas spp., the dominant microbiota of brown shrimp (Crangon crangon) during aerobic storage. Int. J. Food Microbiol. 166, 487–493 (2013).
Roessner, U. et al. Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 13, 11–29 (2001).
Doleschall, F., Recseg, K., Kemény, Z. & Kővári, K. Comparison of differently coated SPME fibres applied for monitoring volatile substances in vegetable oils. Eur. J. Lipid Sci. Technol. 105, 333–338 (2003).
Setkova, L., Risticevic, S. & Pawliszyn, J. Rapid headspace solid-phase microextraction-gas chromatographic–time-of-flight mass spectrometric method for qualitative profiling of ice wine volatile fraction. I. Method development and optimization. J. Chromatogr. A 1147, 213–223 (2007).
Lemfack, M.C., Nickel, J., Dunkel, M., Preissner, R. & Piechulla, B. mVOC: a database of microbial volatiles. Nucleic Acids Res. 42, D744–D748 (2014).
Yoon, S.H. et al. Comparative multi-omics systems analysis of Escherichia coli strains B and K-12. Genome Biol. 13, R37 (2012).
Acknowledgements
This research was supported by grants from the BioNano Health-Guard Research Center, funded by the Ministry of Science, ICT and Future Planning of Korea as a Global Frontier Project (Grant H-GUARD_2013M3A6B2078953); the Woo Jang-Choon Project (PJ01093904) of the Rural Development Administration (RDA); and KRIBB Initiative Program South Korea to C.-M.R. B.A. and J.-M.G. are supported by grant no.: ANR-10-LABX-62-IBEID (Investissement d'Avenir Program) and by Fondation pour la Recherche Médicale grant Equipe FRM DEQ20140329508. M.A.F. acknowledges funding received from the Alexander von Humboldt Foundation, Germany, and Cairo University Research Grant no. 31.
Author information
Authors and Affiliations
Contributions
C.-M.R. and M.A.F. designed the experiments; M.A.F., G.C.S., S.L., B.A. and J.-M.G. performed the experiments; and M.A.F., C.-M.R., Y.-S.P., J.-M.G., B.A. and J.W.K. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Chromatographic profiles of volatiles released from B. subtilis PGPR strain GBO3 (A), and uninoculated media (B)
Compounds positively identified include acetoin (1), 2,3-butanediol (2), decanal (3), decane (4), tetramethyl pyrazine (5), and undecane (6). Highlighted in blue is the earlier elution region where most identified bioactive volatiles are eluted including 1, acetoin and 2, 2,3-butanediol. Peaks commonly released from media include some short chain hydrocarbons and tetramethylpyrazine (s).
Supplementary information
Supplementary Figure 1
Chromatographic profiles of volatiles released from (a) B. subtilis PGPR strain GBO3 and (b) uninoculated medium. (PDF 130 kb)
Rights and permissions
About this article
Cite this article
Farag, M., Song, G., Park, YS. et al. Biological and chemical strategies for exploring inter- and intra-kingdom communication mediated via bacterial volatile signals. Nat Protoc 12, 1359–1377 (2017). https://doi.org/10.1038/nprot.2017.023
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.023
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.