Abstract
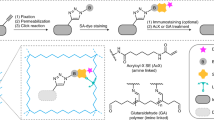
Recent progress in super-resolution imaging (SRI) has created a strong need to improve protein labeling with probes of small size that minimize the target-to-label distance, increase labeling density, and efficiently penetrate thick biological tissues. This protocol describes a method for labeling genetically modified proteins incorporating a small biotin acceptor peptide with a 3-nm fluorescent probe, monomeric streptavidin. We show how to express, purify, and conjugate the probe to organic dyes with different fluorescent properties, and how to label selectively biotinylated membrane proteins for SRI techniques (point accumulation in nanoscale topography (PAINT), stimulated emission depletion (STED), stochastic optical reconstruction microscopy (STORM)). This method is complementary to the previously described anti-GFP-nanobody/SNAP-tag strategies, with the main advantage being that it requires only a short 15-amino-acid tag, and can thus be used with proteins resistant to fusion with large tags and for multicolor imaging. The protocol requires standard molecular biology/biochemistry equipment, making it easily accessible for laboratories with only basic skills in cell biology and biochemistry. The production/purification/conjugation steps take ∼5 d, and labeling takes a few minutes to an hour.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stefan, W.H. et al. The 2015 super-resolution microscopy roadmap. J. Phys. D Appl. Phys. 48, 443001 (2015).
Fernandez-Suarez, M. & Ting, A.Y. Fluorescent probes for super-resolution imaging in living cells. Nat. Rev. Mol. Cell Biol. 9, 929–943 (2008).
Ha, T. & Tinnefeld, P. Photophysics of fluorescent probes for single-molecule biophysics and super-resolution imaging. Annu. Rev. Phys. Chem. 63, 595–617 (2012).
Cranfill, P.J. et al. Quantitative assessment of fluorescent proteins. Nat. Methods 13, 557–562 (2016).
Vreja, I.C. et al. Super-resolution microscopy of clickable amino acids reveals the effects of fluorescent protein tagging on protein assemblies. ACS Nano 9, 11034–11041 (2015).
Stadler, C. et al. Immunofluorescence and fluorescent-protein tagging show high correlation for protein localization in mammalian cells. Nat. Methods 10, 315–323 (2013).
Landgraf, D., Okumus, B., Chien, P., Baker, T.A. & Paulsson, J. Segregation of molecules at cell division reveals native protein localization. Nat. Methods 9, 480–482 (2012).
Deschout, H. et al. Precisely and accurately localizing single emitters in fluorescence microscopy. Nat. Methods 11, 253–266 (2014).
Ries, J., Kaplan, C., Platonova, E., Eghlidi, H. & Ewers, H. A simple, versatile method for GFP-based super-resolution microscopy via nanobodies. Nat. Methods 9, 582–584 (2012).
Chamma, I. et al. Mapping the dynamics and nanoscale organization of synaptic adhesion proteins using monomeric streptavidin. Nat. Commun. 7, 10773 (2016).
Rothbauer, U. et al. Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat. Methods 3, 887–889 (2006).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Los, G.V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Jespersen, L.K., Kuusinen, A., Orellana, A., Keinanen, K. & Engberg, J. Use of proteoliposomes to generate phage antibodies against native AMPA receptor. Eur. J. Biochem. 267, 1382–1389 (2000).
Mikhaylova, M. et al. Resolving bundled microtubules using anti-tubulin nanobodies. Nat. Commun. 6, 7933 (2015).
Nikic, I., Kang, J.H., Girona, G.E., Aramburu, I.V. & Lemke, E.A. Labeling proteins on live mammalian cells using click chemistry. Nat. Protoc. 10, 780–791 (2015).
Lim, K.H., Huang, H., Pralle, A. & Park, S. Stable, high-affinity streptavidin monomer for protein labeling and monovalent biotin detection. Biotechnol. Bioeng. 110, 57–67 (2013).
Demonte, D., Drake, E.J., Lim, K.H., Gulick, A.M. & Park, S. Structure-based engineering of streptavidin monomer with a reduced biotin dissociation rate. Proteins 81, 1621–1633 (2013).
Studier, F.W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Schatz, P.J. Use of peptide libraries to map the substrate specificity of a peptide-modifying enzyme: a 13 residue consensus peptide specifies biotinylation in Escherichia coli. Biotechnology 11, 1138–1143 (1993).
Beckett, D., Kovaleva, E. & Schatz, P.J. A minimal peptide substrate in biotin holoenzyme synthetase-catalyzed biotinylation. Protein Sci. 8, 921–929 (1999).
Parrott, M.B. & Barry, M.A. Metabolic biotinylation of secreted and cell surface proteins from mammalian cells. Biochem. Biophys. Res. Commun. 281, 993–1000 (2001).
Saxton, M.J. & Jacobson, K. Single-particle tracking: applications to membrane dynamics. Annu. Rev. Biophys. Biomol. Struct. 26, 373–399 (1997).
Sharonov, A. & Hochstrasser, R.M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl. Acad. Sci. USA 103, 18911–18916 (2006).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Klar, T.A., Jakobs, S., Dyba, M., Egner, A. & Hell, S.W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl. Acad. Sci. USA 97, 8206–8210 (2000).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Endesfelder, U. et al. Chemically induced photoswitching of fluorescent probes--a general concept for super-resolution microscopy. Molecules 16, 3106–3118 (2011).
Lukinavicius, G. et al. Fluorogenic probes for multicolor imaging in living cells. J. Am. Chem. Soc. 138, 9365–9368 (2016).
Lavis, L.D. et al. Bright photoactivatable fluorophores for single-molecule imaging. bioRxiv 13, 985–988 (2016).
Howarth, M. et al. A monovalent streptavidin with a single femtomolar biotin binding site. Nat. Methods 3, 267–273 (2006).
Howarth, M. & Ting, A.Y. Imaging proteins in live mammalian cells with biotin ligase and monovalent streptavidin. Nat. Protoc. 3, 534–545 (2008).
Grunwald, C. et al. Quantum-yield-optimized fluorophores for site-specific labeling and super-resolution imaging. J. Am. Chem. Soc. 133, 8090–8093 (2011).
Wieneke, R., Raulf, A., Kollmannsperger, A., Heilemann, M. & Tampe, R. SLAP: small labeling pair for single-molecule super-resolution imaging. Angew. Chem. Int. Ed. Engl. 54, 10216–10219 (2015).
Yin, J., Lin, A.J., Golan, D.E. & Walsh, C.T. Site-specific protein labeling by Sfp phosphopantetheinyl transferase. Nat. Protoc. 1, 280–285 (2006).
Yagi, T., Terada, N., Baba, T. & Ohno, S. Localization of endogenous biotin-containing proteins in mouse Bergmann glial cells. Histochem. J. 34, 567–572 (2002).
Viens, A. et al. Use of protein biotinylation in vivo for immunoelectron microscopic localization of a specific protein isoform. J. Histochem. Cytochem. 56, 911–919 (2008).
Paetzel, M., Karla, A., Strynadka, N.C. & Dalbey, R.E. Signal peptidases. Chem. Rev. 102, 4549–4580 (2002).
Lim, K.H., Huang, H., Pralle, A. & Park, S. Engineered streptavidin monomer and dimer with improved stability and function. Biochemistry 50, 8682–8691 (2011).
Demonte, D., Dundas, C.M. & Park, S. Expression and purification of soluble monomeric streptavidin in Escherichia coli. Appl. Microbiol. Biotechnol. 98, 6285–6295 (2014).
Chamma, I., Levet, F., Sibarita, J.B., Sainlos, M. & Thoumine, O. Nanoscale organization of synaptic adhesion proteins revealed by single-molecule localization microscopy. Neurophotonics 3, 041810 (2016).
Hughes, L.D., Rawle, R.J. & Boxer, S.G. Choose your label wisely: water-soluble fluorophores often interact with lipid bilayers. PLoS One 9, e87649 (2014).
Dempsey, G.T., Vaughan, J.C., Chen, K.H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
van de Linde, S. et al. Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat. Protoc. 6, 991–1009 (2011).
Acknowledgements
We thank D. Perrais (Interdisciplinary Institute for Neuroscience, University of Bordeaux (IINS)), A. Ting (Stanford University), and S. Park (Buffalo University) for the generous gift of DNA plasmids; M. Letellier (IINS) for the slice cultures; Z. Karatas, B. Tessier, S. Antoine, and I. Gauthereau for molecular biology and biochemistry; and C. Poujol and the Bordeaux Imaging Center for providing access and support to confocal and STED setups. This work received funding from the Centre National de la Recherche Scientifique, Agence Nationale pour la Recherche (grants Synapse-2Dt and Nanodom (O.T.), Integractome and FastNano (O.R., G.G.), and SynAdh (O.T., M.S.), Conseil Régional Aquitaine, Fondation pour la Recherche Médicale, the National Infrastructure France BioImaging (grant ANR-10INBS-04-01), and the Labex BRAIN.
Author information
Authors and Affiliations
Contributions
I.C., M.S., and O.T. designed research and coordinated the research project. O.R. and G.G. provided their expertise and tools for the integrin model and contributed to related experiments. I.C. and M.S. performed experiments and analysis, and wrote the article. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Chamma, I., Rossier, O., Giannone, G. et al. Optimized labeling of membrane proteins for applications to super-resolution imaging in confined cellular environments using monomeric streptavidin. Nat Protoc 12, 748–763 (2017). https://doi.org/10.1038/nprot.2017.010
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.010
This article is cited by
-
Molecular motion and tridimensional nanoscale localization of kindlin control integrin activation in focal adhesions
Nature Communications (2021)
-
Functional expression of monomeric streptavidin and fusion proteins in Escherichia coli: applications in flow cytometry and ELISA
Applied Microbiology and Biotechnology (2018)
-
Visualizing endocytic recycling and trafficking in live neurons by subdiffractional tracking of internalized molecules
Nature Protocols (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.