Abstract
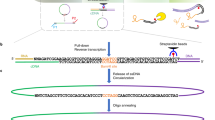
The structure of RNA molecules has a critical role in regulating gene expression, largely through influencing their interactions with RNA-binding proteins (RBPs). RNA hybrid and individual-nucleotide resolution UV cross-linking and immunoprecipitation (hiCLIP) is a transcriptome-wide method of monitoring these interactions by identifying RNA duplexes bound by a specific RBP. The hiCLIP protocol consists of the following steps: in vivo cross-linking of RBPs to their bound RNAs; partial RNA digestion and purification of RNA duplexes interacting with the specific RBP using immunoprecipitation; ligation of the two arms of RNA duplexes via a linker; reverse transcription; cDNA library amplification; and finally high-throughput DNA sequencing. Mapping of the sequenced arms to a reference transcriptome identifies the exact locations of duplexes. hiCLIP data can directly identify all types of RNA duplexes bound by RBPs, including those that are challenging to predict computationally, such as intermolecular and long-range intramolecular duplexes. Moreover, the use of an adaptor that links the two arms of the RNA duplex permits hiCLIP to unambiguously identify the duplexes. Here we describe in detail the procedure for a hiCLIP experiment and the subsequent streamlined data analysis with an R package, 'hiclipr' (https://github.com/luslab/hiclipr/). Preparation of the library for high-throughput DNA sequencing takes ∼7 d and the basic bioinformatic pipeline takes 1 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Wan, Y., Kertesz, M., Spitale, R.C., Segal, E. & Chang, H.Y. Understanding the transcriptome through RNA structure. Nat. Rev. Genet. 12, 641–655 (2011).
Lu, Z. & Chang, H.Y. Decoding the RNA structurome. Curr. Opin. Struct. Biol. 36, 142–148 (2016).
Tian, B., Bevilacqua, P.C., Diegelman-Parente, A. & Mathews, M.B. The double-stranded-RNA-binding motif: interference and much more. Nat. Rev. Mol. Cell Biol. 5, 1013–1023 (2004).
He, L. & Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Gong, C. & Maquat, L.E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–288 (2011).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Gong, C., Tang, Y. & Maquat, L.E. mRNA-mRNA duplexes that autoelicit Staufen1-mediated mRNA decay. Nat. Struct. Mol. Biol. 20, 1214–1220 (2013).
Sugimoto, Y. et al. hiCLIP reveals the in vivo atlas of mRNA secondary structures recognized by Staufen 1. Nature 519, 491–494 (2015).
Aw, J.G. et al. In vivo mapping of eukaryotic RNA interactomes reveals principles of higher-order organization and regulation. Mol. Cell 62, 603–617 (2016).
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016).
Sharma, E., Sterne-Weiler, T., O'Hanlon, D. & Blencowe, B.J. Global mapping of human RNA-RNA interactions. Mol. Cell 62, 618–626 (2016).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J.S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Spitale, R.C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Flynn, R.A. et al. Transcriptome-wide interrogation of RNA secondary structure in living cells with icSHAPE. Nat. Protoc. 11, 273–290 (2016).
Kwok, C.K., Ding, Y., Tang, Y., Assmann, S.M. & Bevilacqua, P.C. Determination of in vivo RNA structure in low-abundance transcripts. Nat. Commun. 4, 2971 (2013).
Weeks, K.M. & Mauger, D.M. Exploring RNA structural codes with SHAPE chemistry. Acc. Chem. Res. 44, 1280–1291 (2011).
Zuker, M. & Stiegler, P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 9, 133–148 (1981).
Kudla, G., Granneman, S., Hahn, D., Beggs, J.D. & Tollervey, D. Cross-linking, ligation, and sequencing of hybrids reveals RNA-RNA interactions in yeast. Proc. Natl. Acad. Sci. USA 108, 10010–10015 (2011).
Helwak, A., Kudla, G., Dudnakova, T. & Tollervey, D. Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 153, 654–665 (2013).
Helwak, A. & Tollervey, D. Mapping the miRNA interactome by cross-linking ligation and sequencing of hybrids (CLASH). Nat. Protoc. 9, 711–728 (2014).
Calvet, J.P. & Pederson, T. Heterogeneous nuclear RNA double-stranded regions probed in living HeLa cells by crosslinking with the psoralen derivative aminomethyltrioxsalen. Proc. Natl. Acad. Sci. USA 76, 755–759 (1979).
Cimino, G.D., Gamper, H.B., Isaacs, S.T. & Hearst, J.E. Psoralens as photoactive probes of nucleic acid structure and function: organic chemistry, photochemistry, and biochemistry. Annu. Rev. Biochem. 54, 1151–1193 (1985).
Kaufmann, G., Klein, T. & Littauer, U.Z. T4 RNA ligase: substrate chain length requirements. FEBS Lett. 46, 271–275 (1974).
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Konig, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Moore, M.J. et al. Mapping Argonaute and conventional RNA-binding protein interactions with RNA at single-nucleotide resolution using HITS-CLIP and CIMS analysis. Nat. Protoc. 9, 263–293 (2014).
Sander, J.D. & Joung, J.K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Darnell, R.B. HITS-CLIP: panoramic views of protein-RNA regulation in living cells. Wiley Interdiscip. Rev. RNA 1, 266–286 (2010).
Van Nostrand, E.L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Zarnegar, B.J. et al. irCLIP platform for efficient characterization of protein-RNA interactions. Nat. Methods 13, 489–492 (2016).
Grosswendt, S. et al. Unambiguous identification of miRNA:target site interactions by different types of ligation reactions. Mol. Cell 54, 1042–1054 (2014).
Moore, M.J. et al. miRNA-target chimeras reveal miRNA 3′-end pairing as a major determinant of Argonaute target specificity. Nat. Commun. 6, 8864 (2015).
Huppertz, I. et al. iCLIP: protein-RNA interactions at nucleotide resolution. Methods 65, 274–287 (2014).
Tollervey, J.R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Harlow, E. & Lane, D. Using Antibodies : A Laboratory Manual (Cold Spring Harbor Laboratory Press, 1999).
Liu, J., Haorah, J. & Xiong, H. Western blotting technique in biomedical research. in : Current Laboratory Methods in Neuroscience Research (eds. Xiong, H. & Gendelman, H.E.) 187–200 (Springer, 2014).
Hymer, W.C. & Kuff, E.L. Isolation of nuclei from mammalian tissues through the use of Triton X-100. J. Histochem. Cytochem. 12, 359–363 (1964).
Stockley, P.G. Filter-binding assays. Methods Mol. Biol. 543, 1–14 (2009).
Farrell, R.E. RNA Methodologies: A Laboratory Guide for Isolation and Characterization (Academic Press, 1993).
Konig, J. et al. iCLIP--transcriptome-wide mapping of protein-RNA interactions with individual nucleotide resolution. J. Vis. Exp. 50, e2638 (2011).
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 10, 1507–1517 (2004).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, 1–10 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Tarasov, A., Vilella, A.J., Cuppen, E., Nijman, I.J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Acknowledgements
We thank F. Agostini (Luscombe laboratory) for helpful advice on the 'hiclipr' package; C. Militti (Ule laboratory) for valuable comments on the manuscript; F. Lee and I. Ruiz de los Mozos (Ule laboratory) for testing the 'hiclipr' package; and all the members of the Ule and Luscombe laboratories for providing fruitful discussions throughout the study. The hiCLIP project was supported by funding from the European Research Council (617837-Translate) to J.U., a Wellcome Trust Joint Investigator Award to J.U. and N.M.L. (103760/Z/14/Z), the Nakajima Foundation Fellowship and an MRC Centenary Early Career Award to Y.S., a Wellcome Trust PhD Training Fellowship for Clinicians to A.M.C., and the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001002), the UK Medical Research Council (FC001002), and the Wellcome Trust (FC001002).
Author information
Authors and Affiliations
Contributions
Y.S. and J.U. conceived the hiCLIP protocol; Y.S. and A.M.C. wrote and described the software for data analysis; A.M.C. developed the 'hiclipr' package; Y.S., A.M.C., N.M.L., and J.U. wrote the manuscript; and J.U. and N.M.L. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Sugimoto, Y., Chakrabarti, A., Luscombe, N. et al. Using hiCLIP to identify RNA duplexes that interact with a specific RNA-binding protein. Nat Protoc 12, 611–637 (2017). https://doi.org/10.1038/nprot.2016.188
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.188
This article is cited by
-
Recent advances in RNA structurome
Science China Life Sciences (2022)
-
Epitranscriptomic technologies and analyses
Science China Life Sciences (2020)
-
Advances and challenges towards the study of RNA-RNA interactions in a transcriptome-wide scale
Quantitative Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.