Abstract
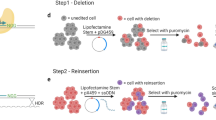
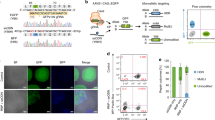
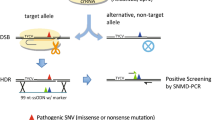
CRISPR/Cas9 is a promising tool for genome-editing DNA in cells with single-base-pair precision, which allows novel in vitro models of human disease to be generated—e.g., in pluripotent stem cells. However, the accuracy of intended sequence changes can be severely diminished by CRISPR/Cas9's propensity to re-edit previously modified loci, causing unwanted mutations (indels) alongside intended changes. Here we describe a genome-editing framework termed consecutive re-guide or re-Cas steps to erase CRISPR/Cas-blocked targets (CORRECT), which, by exploiting the use of highly efficacious CRISPR/Cas-blocking mutations in two rounds of genome editing, enables accurate, efficient and scarless introduction of specific base changes—for example, in human induced pluripotent (iPS) stem cells. This protocol outlines in detail how to implement either the re-Guide or re-Cas variants of CORRECT to generate scarlessly edited isogenic stem cell lines with intended monoallelic and biallelic sequence changes in ∼3 months.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Jinek, M. et al. RNA-programmed genome editing in human cells. elife 2, e00471 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Cho, S.W., Kim, S., Kim, J.M. & Kim, J.-S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Hruscha, A. et al. Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development 140, 4982–4987 (2013).
Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167–170 (2010).
Marraffini, L.A. & Sontheimer, E.J. Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463, 568–571 (2010).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J.A. A Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Dow, L.E. et al. Inducible in vivo genome editing with CRISPR-Cas9. Nat. Biotechnol. 33, 390–394 (2015).
Platt, R.J. et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Canver, M.C. et al. Characterization of genomic deletion efficiency mediated by clustered regularly interspaced palindromic repeats (CRISPR)/Cas9 nuclease system in mammalian cells. J. Biol. Chem. 289, 21312–21324 (2014).
Lieber, M.R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Lin, S., Staahl, B.T., Alla, R.K. & Doudna, J.A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. elife 3, e04766 (2015).
Yu, C. et al. Small molecules enhance CRISPR genome editing in pluripotent stem cells. Cell Stem Cell 16, 142–147 (2015).
Maruyama, T. et al. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat. Biotechnol. 33, 538–542 (2015).
Chu, V.T. et al. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 33, 543–548 (2015).
Richardson, C.D., Ray, G.J., DeWitt, M.A., Curie, G.L. & Corn, J.E. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat. Biotechnol. 34, 339–344 (2016).
Song, J. et al. RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency. Nat. Commun. 7, 1–7 (2016).
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129 (2016).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843 (2013).
Kuscu, C., Arslan, S., Singh, R., Thorpe, J. & Adli, M. Genome-wide analysis reveals characteristics of off-target sites bound by the Cas9 endonuclease. Nat. Biotechnol. 32, 677–683 (2014).
Kleinstiver, B.P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Miyaoka, Y. et al. Isolation of single-base genome-edited human iPS cells without antibiotic selection. Nat. Methods 11, 291–293 (2014).
Yusa, K. Seamless genome editing in human pluripotent stem cells using custom endonuclease-based gene targeting and the piggyBac transposon. Nat. Protoc. 8, 2061–2078 (2013).
Nihongaki, Y., Kawano, F., Nakajima, T. & Sato, M. Photoactivatable CRISPR-Cas9 for optogenetic genome editing. Nat. Biotechnol. 33, 755–760 (2015).
Zetsche, B., Volz, S.E. & Zhang, F. A split-Cas9 architecture for inducible genome editing and transcription modulation. Nat. Biotechnol. 33, 139–142 (2015).
Kleinstiver, B.P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl. Acad. Sci. USA 108, 10098–10103 (2011).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Moreno-Mateos, M.A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Gaj, T. et al. Efficient delivery of nuclease proteins for genome editing in human stem cells and primary cells. Nat. Protoc. 10, 1842–1859 (2015).
Bell, C.C., Magor, G.W., Gillinder, K.R. & Perkins, A.C. A high-throughput screening strategy for detecting CRISPR-Cas9 induced mutations using next-generation sequencing. BMC Genomics 15, 1002 (2014).
Smith, C. et al. Whole-genome sequencing analysis reveals high specificity of CRISPR/Cas9 and TALEN-based genome editing in human iPSCs. Cell Stem Cell 15, 12–13 (2014).
Crosetto, N. et al. Nucleotide-resolution DNA double-strand break mapping by next-generation sequencing. Nat. Methods 10, 361–365 (2013).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Kim, D. et al. Digenome-seq: genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat. Methods 12, 237–243 (2015).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Braam, S.R. et al. Feeder-free culture of human embryonic stem cells in conditioned medium for efficient genetic modification. Nat. Protoc. 3, 1435–1443 (2008).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Ren, X. et al. Enhanced specificity and efficiency of the CRISPR/Cas9 system with optimized sgRNA parameters in Drosophila. Cell Rep. 9, 1151–1162 (2014).
Fu, B.X.H., St Onge, R.P., Fire, A.Z. & Smith, J.D. Distinct patterns of Cas9 mismatch tolerance in vitro and in vivo. Nucleic Acids Res. 44, 5365–5377 (2016).
Haass, C. et al. The Swedish mutation causes early-onset Alzheimer's disease by beta-secretase cleavage within the secretory pathway. Nat. Med. 1, 1291–1296 (1995).
Jonsson, T. et al. A mutation in APP protects against Alzheimer's disease and age-related cognitive decline. Nature 488, 96–99 (2012).
Inui, M. et al. Rapid generation of mouse models with defined point mutations by the CRISPR/Cas9 system. Sci. Rep. 4, 5396 (2014).
Yang, L. et al. Optimization of scarless human stem cell genome editing. Nucleic Acids Res. 41, 9049–9061 (2013).
Bialk, P., Rivera-Torres, N., Strouse, B. & Kmiec, E.B. Regulation of gene editing activity directed by single-stranded oligonucleotides and CRISPR/Cas9 Systems. PLoS One 10, e0129308 (2015).
Acknowledgements
This research was supported by The Rockefeller University, The New York Stem Cell Foundation, The Ellison Foundation, the Cure Alzheimer's Fund and the Empire State Stem Cell fund through New York State Department of Health. D.K. is a Howard Hughes Medical Institute International Student Research Fellow and received a fellowship from the National Sciences and Engineering Research Council of Canada. D.P. is a New York Stem Cell Foundation Druckenmiller Fellow and received a fellowship from the German Academy of Sciences Leopoldina. S.T. is supported by the Agency for Science, Technology and Research of Singapore. We thank A. Sproul, S. Jacob and S. Noggle for sharing protocols and know-how for stem cell culture and for sharing stem cell lines. We thank members of the Tessier-Lavigne laboratory for discussions. Our thanks also go to S. Mazel and the team at the Rockefeller University Flow Cytometry Resource Center, C. Zhao and the team at the Rockefeller University Genomics Resource Center, and A. Chen and M. Duffield for technical help. Opinions expressed here are solely those of the authors and do not necessarily reflect those of the Empire State Stem Cell Fund, the New York State Department of Health or the State of New York.
Author information
Authors and Affiliations
Contributions
D.K., D.P. and M.T.-L. conceived and designed the protocol. D.K. and D.P. performed and analyzed the experiments. S.T. helped analyze NGS data. D.K., D.P. and M.T.-L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
sgRNA, primer and ssODN repair template sequences used in our experiments. (XLSX 39 kb)
Rights and permissions
About this article
Cite this article
Kwart, D., Paquet, D., Teo, S. et al. Precise and efficient scarless genome editing in stem cells using CORRECT. Nat Protoc 12, 329–354 (2017). https://doi.org/10.1038/nprot.2016.171
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.171
This article is cited by
-
Proteomics of mouse brain endothelium uncovers dysregulation of vesicular transport pathways during aging
Nature Aging (2024)
-
Homology-based repair induced by CRISPR-Cas nucleases in mammalian embryo genome editing
Protein & Cell (2022)
-
Off-target effects of base editors: what we know and how we can reduce it
Current Genetics (2022)
-
Simple and reliable detection of CRISPR-induced on-target effects by qgPCR and SNP genotyping
Nature Protocols (2021)
-
A Cas9-mediated adenosine transient reporter enables enrichment of ABE-targeted cells
BMC Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.