Abstract
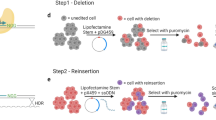
Genome editing of human induced pluripotent stem cells (hiPSCs) offers unprecedented opportunities for in vitro disease modeling and personalized cell replacement therapy. The introduction of Cas9-directed genome editing has expanded adoption of this approach. However, marker-free genome editing using standard protocols remains inefficient, yielding desired targeted alleles at a rate of ∼1–5%. We developed a protocol based on a doxycycline-inducible Cas9 transgene carried on a piggyBac transposon to enable robust and highly efficient Cas9-directed genome editing, so that a parental line can be expeditiously engineered to harbor many separate mutations. Treatment with doxycycline and transfection with guide RNA (gRNA), donor DNA and piggyBac transposase resulted in efficient, targeted genome editing and concurrent scarless transgene excision. Using this approach, in 7 weeks it is possible to efficiently obtain genome-edited clones with minimal off-target mutagenesis and with indel mutation frequencies of 40–50% and homology-directed repair (HDR) frequencies of 10–20%.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Robinton, D.A. & Daley, G.Q. The promise of induced pluripotent stem cells in research and therapy. Nature 481, 295–305 (2012).
Kim, H.S., Bernitz, J.M., Lee, D.F. & Lemischka, I.R. Genomic editing tools to model human diseases with isogenic pluripotent stem cells. Stem Cells Dev. 23, 2673–2686 (2014).
Wang, G. et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat. Med. 20, 616–623 (2014).
Yang, L. et al. Targeted and genome-wide sequencing reveal single nucleotide variations impacting specificity of Cas9 in human stem cells. Nat. Commun. 5, 5507 (2014).
Yang, L. et al. Genome-wide inactivation of porcine endogenous retroviruses (PERVs). Science 350, 1101–1104 (2015).
Gupta, R.M. & Musunuru, K. Expanding the genetic editing tool kit: ZFNs, TALENs, and CRISPR-Cas9. J. Clin. Invest. 124, 4154–4161 (2014).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Yang, L. et al. Optimization of scarless human stem cell genome editing. Nucleic Acids Res. 41, 9049–9061 (2013).
Li, K., Wang, G., Andersen, T., Zhou, P. & Pu, W.T. Optimization of genome engineering approaches with the CRISPR/Cas9 system. PLoS One 9, e105779 (2014).
Ding, Q. et al. Enhanced efficiency of human pluripotent stem cell genome editing through replacing TALENs with CRISPRs. Cell Stem Cell 12, 393–394 (2013).
González, F. et al. An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells. Cell Stem Cell 15, 215–226 (2014).
Bione, S. et al. A novel X-linked gene, G4.5. is responsible for Barth syndrome. Nat. Genet. 12, 385–389 (1996).
Gonzalez, I.L. Barth syndrome: TAZ gene mutations, mRNAs, and evolution. Am. J. Med. Genet. A 134, 409–414 (2005).
Li, X. et al. piggyBac transposase tools for genome engineering. Proc. Natl. Acad. Sci. 110, E2279–E2287 (2013).
Shiba, Y. et al. Human ES-cell-derived cardiomyocytes electrically couple and suppress arrhythmias in injured hearts. Nature 489, 322–325 (2012).
Veres, A. et al. Low incidence of off-target mutations in individual CRISPR-Cas9 and TALEN targeted human stem cell clones detected by whole-genome sequencing. Cell Stem Cell 15, 27–30 (2014).
Suzuki, K. et al. Targeted gene correction minimally impacts whole-genome mutational load in human-disease-specific induced pluripotent stem cell clones. Cell Stem Cell 15, 31–36 (2014).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L.A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Ohnuki, M., Takahashi, K. & Yamanaka, S. Generation and characterization of human induced pluripotent stem cells. Curr. Protoc. Stem Cell Biol. 4A–42 (2009).
Bookout, A.L., Cummins, C.L., Mangelsdorf, D.J., Pesola, J.M. & Kramer, M.F. High-throughput real-time quantitative reverse transcription PCR. Curr. Protoc. Mol. Biol. 15–18 (2006).
Hill, J.T. et al. Poly peak parser: method and software for identification of unknown indels using Sanger sequencing of polymerase chain reaction products. Dev. Dyn. 243, 1632–1636 (2014).
Graham, D.B. & Root, D.E. Resources for the design of CRISPR gene editing experiments. Genome Biol. 16, 260 (2015).
Yang, L., Yang, J.L., Byrne, S., Pan, J. & Church, G.M. CRISPR/Cas9-directed genome editing of cultured cells. Curr. Protoc. Mol. Biol. 107, 31.1.1–31.1.17 . (2014).
Acknowledgements
G.W. was supported by grant T32HL007572 and a Progenitor Cell Biology Consortium Jump Start award. W.T.P. was funded by grants U01 HL100401 and R01 HL128694, and the Barth Syndrome Foundation. L.Y., D.G. and X.R. were funded by NIH Centers of Excellence in Genomic Sciences grant P50 HG005550.
Author information
Authors and Affiliations
Contributions
G.W. and L.Y. developed the protocol. L.Y. and D.G. analyzed sequencing data. G.W. and W.T.P. wrote the manuscript with input from the other authors. W.T.P. and G.M.C. supervised the project. L.Y.Y. performed experiments. K.L. contributed to the gRNA design, molecular cloning, and nucleofection. D.Z. contributed to iPSC culture and differentiation. Y.H. contributed genome-editing data. X.R. contributed to development of the Dox-inducible Cas9 expression plasmid.
Corresponding author
Ethics declarations
Competing interests
L.Y. and G.M.C. are inventors on a patent filed by Harvard University on Cas9 genome editing using the technology described in this protocol.
Integrated supplementary information
Supplementary Figure 1 Efficient genome editing in human pluripotent stem cells.
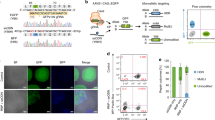
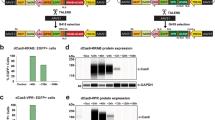
Our genome editing protocol was highly efficient in an additional pluripotent human cell line (CHB10) and at several different loci. a-b. We tested the efficiency of our protocol in a different pluripotent cell line, the human embryonic stem cell line CHB10, and a different locus, an integrated GFP reporter that is inactive due to a stop codon in the open reading frame (red symbol, panel a). Efficiency of HDR-mediated correction of the frameshift mutation was measured by FACS detection of GFP expression (panel b). c. Genome editing at the autosomal DNAJC19 locus of PGP1-hCas9-PB (P) or CHB10-hCas9-PB (C) pluripotent cell lines. Arrowheads indicate bands diagnostic of targeted genome modification. d. Genome editing efficiency at three autosomal sites, one in DNAJC19, and two within JUP (named JUP-M and JUP-C). Graph summarizes the results of Sanger sequencing of individual clones. Both homozygous and heterozygous mutations were efficiently recovered. Genotypes at each of the cell's two alleles is indicated as N, indel mutation from NHEJ; H, point mutation from HDR; +, wild-type. Overall frequency of each genotype class across the 3 experiments is summarized to the right.
Supplementary Figure 2 Efficient genome editing and piggyBac excision in a single step.
a. The co-transfection of the excision-only piggyBac mutant (PB) at the same as gRNA and HDR donor (”one-step” genome editing) did not substantially reduce the yield of genome-edited clones compared to sequential editing followed by excision (”two-step” genome editing). TAZ targeting frequencies were determined by next generation sequencing of genomic DNA from pooled cells. b. The frequency of transgene excision was not substantially different between one-step and two-step protocols. With either protocol, the majority of the recovered clones had successfully undergone excision of the piggyBac transgene, as determined by PCR genotyping of at least 79 independent clones per group over three separate experiments. These results show that including the excision-only piggyBac mutant into the transfection mix with gRNA and donor DNA permits efficient, single step genome editing and transgene excision.
Supplementary Figure 3 Efficient transposon removal by piggyBac transposase transient transfection yielded high quality iPSCs.
a. iPSC lines before after Cas9 genome editing had normal 46-XY karyotype. PGP1-hCas9-PB after transposon removal was designated PGP1e, and PGP1-hCas9-PB-TAZc.517delG was designated PGP1-TAZc.517delG. bar = 20 μm. b-c. Expression of pluripotency markers by control and mutant lines, as determined by qRTPCR (b) or immunostaining (c). d-g. Hematoxylin and eosin staining of teratomas indicated formation of structures from all three germ layers. n, neural. g, glandular. c, cartilagenous. m, musclar. white bar = 100 μm; pink bar = 200 μm. h. Cardiac differentiation of genome-edited, piggyBac excised iPSCs. bar = 20 μm.
Supplementary Figure 4 iPSCs with induced mutation at the TAZ locus recapitulate features of Barth Syndrome patients.
TAZ mutation causes mitochondrial dysfunction and cardiomyopathy by blocking maturation of cardiolipin, the major phospholipid of the inner mitochondrial membrane. (Wang, G., McCain, M. L., Yang, L., He, A., et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat Med 20, 616-623 (2014); Houtkooper, R. H., Turkenburg, M., Poll-The, B. T., Karall, D., et al. The enigmatic role of tafazzin in cardiolipin metabolism. Biochim Biophys Acta 1788, 2003-2014 (2009)). a. Cardiolipin phospholipid mass spectroscopy analysis control (PGP1e) and TAZ mutant (PGP1e-TAZc.517delG) iPSC-derived cardiomyocytes. As expected, PGP1e-TAZc.517delG iPSC-derived cardiomyocytes (iPSC-CMs) showed abnormal cardiolipin maturation, a signature of Barth syndrome b. Normal CL content is required for optimal function of the mitochondrial electron transport chain (Pfeiffer, K., Gohil, V., Stuart, R. A., Hunte, C., et al. Cardiolipin stabilizes respiratory chain supercomplexes. J Biol Chem 278, 52873-52880 (2003)). Respiratory capacity, the rate of oxygen consumption in the presence of the mitochondrial uncoupler trifluorocarbonylcyanide phenylhydrazone (FCCP), is a measure of maximal electron transport chain activity. Oxygen consumption rate of control and mutant iPSC-CMs. Cells were treated with FCCP and analyzed using a Seahorses Biosciences Extracellular Flux Analyzer. Respiratory capacity, the rate of oxygen consumption in the presence of the mitochondrial uncoupler trifluorocarbonylcyanide phenylhydrazone (FCCP), is a measure of maximal electron transport chain activity. We confirmed that respiratory capacity was markedly impaired in iPSC-CMs containing introduced TAZ mutations.
Supplementary information
Supplementary Figures and Supplementary Data
Supplementary Figures 1–4 and Supplementary Data 1 and 2 (PDF 1001 kb)
Rights and permissions
About this article
Cite this article
Wang, G., Yang, L., Grishin, D. et al. Efficient, footprint-free human iPSC genome editing by consolidation of Cas9/CRISPR and piggyBac technologies. Nat Protoc 12, 88–103 (2017). https://doi.org/10.1038/nprot.2016.152
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.152
This article is cited by
-
Enrichment strategies to enhance genome editing
Journal of Biomedical Science (2023)
-
Novel immortalization approach defers senescence of cultured canine adipose-derived mesenchymal stromal cells
GeroScience (2022)
-
A piggyBac-based platform for genome editing and clonal rhesus macaque iPSC line derivation
Scientific Reports (2021)
-
Modular deep learning enables automated identification of monoclonal cell lines
Nature Machine Intelligence (2021)
-
A simple, quick, and efficient CRISPR/Cas9 genome editing method for human induced pluripotent stem cells
Acta Pharmacologica Sinica (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.