Abstract
Fate allocation in the gastrulating embryo is spatially organized as cells differentiate into specialized cell types depending on their positions with respect to the body axes. There is a need for in vitro protocols that allow the study of spatial organization associated with this developmental transition. Although embryoid bodies and organoids can exhibit some spatial organization of differentiated cells, methods that generate embryoid bodies or organoids do not yield consistent and fully reproducible results. Here, we describe a micropatterning approach in which human embryonic stem cells are confined to disk-shaped, submillimeter colonies. After 42 h of BMP4 stimulation, cells form self-organized differentiation patterns in concentric radial domains, which express specific markers associated with the embryonic germ layers, reminiscent of gastrulating embryos. Our protocol takes 3 d; it uses commercial microfabricated slides (from CYTOO), human laminin-521 (LN-521) as extracellular matrix coating, and either conditioned or chemically defined medium (mTeSR). Differentiation patterns within individual colonies can be determined by immunofluorescence and analyzed with cellular resolution. Both the size of the micropattern and the type of medium affect the patterning outcome. The protocol is appropriate for personnel with basic stem cell culture training. This protocol describes a robust platform for quantitative analysis of the mechanisms associated with pattern formation at the onset of gastrulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Arnold, S.J. & Robertson, E.J. Making a commitment: cell lineage allocation and axis patterning in the early mouse embryo. Nat. Rev. Mol. Cell Biol. 10, 91–103 (2009).
Behringer, R.R., Wakamiya, M., Tsang, T.E. & Tam, P.P. A flattened mouse embryo: leveling the playing field. Genesis 28, 23–30 (2000).
Dobreva, M.P., Pereira, P.N., Deprest, J. & Zwijsen, A. On the origin of amniotic stem cells: of mice and men. Int. J. Dev. Biol. 54, 761–777 (2010).
Rossant, J. Mouse and human blastocyst-derived stem cells: vive les differences. Development 142, 9–12 (2015).
Warmflash, A. et al. Dynamics of TGF-β signaling reveal adaptive and pulsatile behaviors reflected in the nuclear localization of transcription factor Smad4. Proc. Natl. Acad. Sci. USA 109, E1947–E1956 (2012).
Sorre, B., Warmflash, A., Brivanlou, A.H. & Siggia, E.D. Encoding of temporal signals by the TGF-β pathway and implications for embryonic patterning.. Dev. Cell 30, 334–342 (2014).
Thery, M. et al. Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. Proc. Natl. Acad. Sci. USA 103, 19771–19776 (2006).
Thery, M., Jimenez-Dalmaroni, A., Racine, V., Bornens, M. & Julicher, F. Experimental and theoretical study of mitotic spindle orientation. Nature 447, 493–496 (2007).
Peerani, R., Bauwens, C., Kumacheva, E. & Zandstra, P.W. Patterning mouse and human embryonic stem cells using micro-contact printing. Methods Mol. Biol. 482, 21–33 (2009).
Lee, L.H. et al. Micropatterning of human embryonic stem cells dissects the mesoderm and endoderm lineages. Stem Cell Res. 2, 155–162 (2009).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E.D. & Brivanlou, A.H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Kurek, D. et al. Endogenous WNT signals mediate BMP-induced and spontaneous differentiation of epiblast stem cells and human embryonic stem cells. Stem Cell Rep. 4, 114–128 (2015).
Azioune, A., Storch, M., Bornens, M., Thery, M. & Piel, M. Simple and rapid process for single cell micro-patterning. Lab. Chip 9, 1640–1642 (2009).
Ma, Z. et al. Self-organizing human cardiac microchambers mediated by geometric confinement. Nat. Commun. 6, 7413 (2015).
Thery, M. & Piel, M. Adhesive micropatterns for cells: a microcontact printing protocol. Cold Spring Harb. Protoc. 2009 http://dx.doi.org/10.1101/pdb.prot5255 (2009).
van den Brink, S.C. et al. Symmetry breaking, germ layer specification and axial organisation in aggregates of mouse embryonic stem cells. Development 141, 4231–4242 (2014).
Poh, Y.C. et al. Generation of organized germ layers from a single mouse embryonic stem cell. Nat. Commun. 5, 4000 (2014).
ten Berge, D. et al. Wnt signaling mediates self-organization and axis formation in embryoid bodies. Cell Stem Cell 3, 508–518 (2008).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Lancaster, M.A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Sato, T. & Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science 340, 1190–1194 (2013).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Engler, A.J., Sen, S., Sweeney, H.L. & Discher, D.E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Migeotte, I., Omelchenko, T., Hall, A. & Anderson, K.V. Rac1-dependent collective cell migration is required for specification of the anterior-posterior body axis of the mouse. PLoS Biol. 8, e1000442 (2010).
Nowotschin, S. et al. The T-box transcription factor eomesodermin is essential for AVE induction in the mouse embryo. Genes Dev. 27, 997–1002 (2013).
Schneider, C.A., Rasband, W.S. & Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Al-Kofahi, Y., Lassoued, W., Lee, W. & Roysam, B. Improved automatic detection and segmentation of cell nuclei in histopathology images. IEEE Trans. Biomed. Eng. 57, 841–852 (2010).
Gniadek, T.J. & Warren, G. WatershedCounting3D: a new method for segmenting and counting punctate structures from confocal image data. Traffic 8, 339–346 (2007).
Mathew, B. et al. Robust and automated three-dimensional segmentation of densely packed cell nuclei in different biological specimens with lines-of-sight decomposition. BMC Bioinformatics 16, 187 (2015).
Acknowledgements
We thank the members of the Brivanlou, Siggia, and Warmflash laboratories for helpful discussions. This work was supported by NIH/NIGMS R01 GM101653 (A.H.B., E.S.), NIH/DHHS R01 HD080699 (A.H.B., E.S.), Cancer Prevention Research Institute of Texas (CPRIT) grant RR140073 (A.W.), NSF grant DGE-1325261 (A.Y.), and NSF grant MCB-1553228 (A.W.).
Author information
Authors and Affiliations
Contributions
All authors contributed to the design of the experiments. A.D., F.E., M.C.G., I.M., J.M., A.R., M.S., A.Y., and A.W. performed and analyzed the experiments. All authors wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
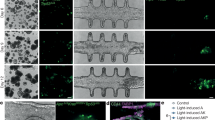
BMP4 differentiation in micropatterned hESC colonies grown in mTeSR.
Immunofluorescence staining of ESI017 hESCs 48 hours after BMP4 (50 ng/ml) treatment. Shown are colonies of 1000, 800, and 500 μm seeded and grown following the alternative protocol described in Box 1. CDX2 (green); BRA (red); SOX2 (blue). Scale bars, 250 μm. ESCRO institutional regulatory board permission was obtained to perform these experiments. (PDF 1525 kb)
Rights and permissions
About this article
Cite this article
Deglincerti, A., Etoc, F., Guerra, M. et al. Self-organization of human embryonic stem cells on micropatterns. Nat Protoc 11, 2223–2232 (2016). https://doi.org/10.1038/nprot.2016.131
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.131
This article is cited by
-
An in vitro culture platform for studying the effect of collective cell migration on spatial self-organization within induced pluripotent stem cell colonies
Journal of Biological Engineering (2023)
-
Nodal is a short-range morphogen with activity that spreads through a relay mechanism in human gastruloids
Nature Communications (2022)
-
Membrane potential drives the exit from pluripotency and cell fate commitment via calcium and mTOR
Nature Communications (2022)
-
The evolution of our understanding of human development over the last 10 years
Nature Communications (2021)
-
lncRNA DIGIT and BRD3 protein form phase-separated condensates to regulate endoderm differentiation
Nature Cell Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.