Abstract
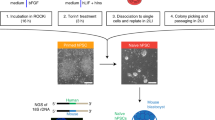
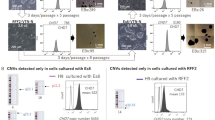
The ability to derive and stably maintain ground-state human pluripotent stem cells (hPSCs) that resemble the cells seen in vivo in the inner cell mass has the potential to be an invaluable tool for researchers developing stem cell–based therapies. To date, derivation of human naive-like pluripotent stem cell lines has been limited to a small number of lineages, and their long-term culturing remains problematic. We describe a protocol for genetic and phenotypic tagging, selecting and maintaining naive-like hPSCs. We tag hPSCs by GFP, expressed by the long terminal repeat (LTR7) of HERVH endogenous retrovirus. This simple and efficient protocol has been reproduced with multiple hPSC lines, including embryonic and induced pluripotent stem cells, and it takes ∼6 weeks. By using the reporter, homogeneous hPSC cultures can be derived, characterized and maintained for the long term by repeated re-sorting and re-plating steps. The HERVH-expressing cells have a similar, but nonidentical, expression pattern to other naive-like cells, suggesting that alternative pluripotent states might exist.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
29 October 2018
In the published version of this paper, the authors omitted a funding source. L.D.H. acknowledges support from the European Research Council (Advanced Grant ERC-2014-ADG 669207). The original article has not been corrected.
References
Stadtfeld, M. & Hochedlinger, K. Induced pluripotency: history, mechanisms, and applications. Genes Dev. 24, 2239–2263 (2010).
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009).
Boroviak, T., Loos, R., Bertone, P., Smith, A. & Nichols, J. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat. Cell Biol. 16, 516–528 (2014).
Brons, I.G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Tesar, P.J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Bao, S. et al. Epigenetic reversion of post-implantation epiblast to pluripotent embryonic stem cells. Nature 461, 1292–1295 (2009).
Evans, M.J. & Kaufman, M.H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Martin, G.R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl. Acad. Sci. USA 78, 7634–7638 (1981).
Ying, Q.L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Thomson, J.A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Hanna, J. et al. Human embryonic stem cells with biological and epigenetic characteristics similar to those of mouse ESCs. Proc. Natl. Acad. Sci. USA 107, 9222–9227 (2010).
Theunissen, T.W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Yan, L. et al. Single-cell RNA-seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Smith, Z.D. et al. DNA methylation dynamics of the human preimplantation embryo. Nature 511, 611–615 (2014).
Chen, Y., Blair, K. & Smith, A. Robust self-renewal of rat embryonic stem cells requires fine-tuning of glycogen synthase kinase-3 inhibition. Stem Cell Rep. 1, 209–217 (2013).
Meek, S. et al. Tuning of β-catenin activity is required to stabilize self-renewal of rat embryonic stem cells. Stem Cells 31, 2104–2115 (2013).
Hoffman, L.M. & Carpenter, M.K. Characterization and culture of human embryonic stem cells. Nat. Biotechnol. 23, 699–708 (2005).
Wang, J. et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516, 405–409 (2014).
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 42, 631–634 (2010).
Belshaw, R. et al. Long-term reinfection of the human genome by endogenous retroviruses. Proc. Natl. Acad. Sci. USA 101, 4894–4899 (2004).
Goke, J. et al. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell 16, 135–141 (2015).
Sundaram, V. et al. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. 24, 1963–1976 (2014).
Lu, X. et al. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 21, 423–425 (2014).
Mates, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Gafni, O. et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 504, 282–286 (2013).
Ware, C.B. et al. Derivation of naive human embryonic stem cells. Proc. Natl. Acad. Sci. USA 111, 4484–4489 (2014).
Takashima, Y. et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 158, 1254–1269 (2014).
Chan, Y.S. et al. Induction of a human pluripotent state with distinct regulatory circuitry that resembles preimplantation epiblast. Cell Stem Cell 13, 663–675 (2013).
Martello, G., Bertone, P. & Smith, A. Identification of the missing pluripotency mediator downstream of leukaemia inhibitory factor. EMBO J. 32, 2561–2574 (2013).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Okamoto, I. et al. Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature 472, 370–374 (2011).
Fang, R. et al. Generation of naive induced pluripotent stem cells from rhesus monkey fibroblasts. Cell Stem Cell 15, 488–496 (2014).
Chen, Y. et al. Generation of cynomolgus monkey chimeric fetuses using embryonic stem cells. Cell Stem Cell 17, 116–124 (2015).
Irie, N. et al. SOX17 is a critical specifier of human primordial germ cell fate. Cell 160, 253–268 (2015).
Huang, K., Maruyama, T. & Fan, G. The naive state of human pluripotent stem cells: a synthesis of stem cell and preimplantation embryo transcriptome analyses. Cell Stem Cell 15, 410–415 (2014).
Schlesinger, S. & Goff, S.P. Retroviral transcriptional regulation and embryonic stem cells: war and peace. Mol. Cell. Biol. 35, 770–777 (2015).
Kelley, D. & Rinn, J. Transposable elements reveal a stem cell-specific class of long noncoding RNAs. Genome Biol. 13, R107 (2012).
Loewer, S. et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat. Genet. 42, 1113–1117 (2010).
Ng, S.Y., Johnson, R. & Stanton, L.W. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J. 31, 522–533 (2012).
Dunn, S.J., Martello, G., Yordanov, B., Emmott, S. & Smith, A.G. Defining an essential transcription factor program for naive pluripotency. Science 344, 1156–1160 (2014).
Okamoto, I., Otte, A.P., Allis, C.D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004).
Mak, W. et al. Reactivation of the paternal X chromosome in early mouse embryos. Science 303, 666–669 (2004).
Koyanagi-Aoi, M. et al. Differentiation-defective phenotypes revealed by large-scale analyses of human pluripotent stem cells. Proc. Natl. Acad. Sci. USA 110, 20569–20574 (2013).
Ohnuki, M. et al. Dynamic regulation of human endogenous retroviruses mediates factor-induced reprogramming and differentiation potential. Proc. Natl. Acad. Sci. USA 111, 12426–12431 (2014).
Grow, E.J. et al. Intrinsic retroviral reactivation in human preimplantation embryos and pluripotent cells. Nature 522, 221–225 (2015).
Vigdal, T.J., Kaufman, C.D., Izsvak, Z., Voytas, D.F. & Ivics, Z. Common physical properties of DNA affecting target site selection of sleeping beauty and other Tc1/mariner transposable elements. J. Mol. Biol. 323, 441–452 (2002).
Yant, S.R. et al. High-resolution genome-wide mapping of transposon integration in mammals. Mol. Cell. Biol. 25, 2085–2094 (2005).
Voigt, K. et al. Retargeting Sleeping Beauty transposon insertions by engineered zinc finger DNA-binding domains. Mol. Ther. 20, 1852–1862 (2012).
Grabundzija, I. et al. Comparative analysis of transposable element vector systems in human cells. Mol. Ther. 18, 1200–1209 (2010).
Papapetrou, E.P. et al. Genomic safe harbors permit high β-globin transgene expression in thalassemia induced pluripotent stem cells. Nat. Biotechnol. 29, 73–78 (2011).
Grabundzija, I. et al. Sleeping Beauty transposon-based system for cellular reprogramming and targeted gene insertion in induced pluripotent stem cells. Nucleic Acids Res. 41, 1829–1847 (2013).
Largaespada, D.A. & Collier, L.S. Transposon-mediated mutagenesis in somatic cells: identification of transposon-genomic DNA junctions. Methods Mol. Biol. 435, 95–108 (2008).
Ivics, Z., Izsvak, Z., Medrano, G., Chapman, K.M. & Hamra, F.K. Sleeping Beauty transposon mutagenesis in rat spermatogonial stem cells. Nat. Protoc. 6, 1521–1535 (2011).
Nagy, A. et al. Embryonic stem cells alone are able to support fetal development in the mouse. Development 110, 815–821 (1990).
Kolodziejczyk, A.A. et al. Single cell RNA-sequencing of pluripotent states unlocks modular transcriptional variation. Cell Stem Cell 17, 471–485 (2015).
Tonge, P.D. et al. Divergent reprogramming routes lead to alternative stem-cell states. Nature 516, 192–197 (2014).
Vassena, R. et al. Waves of early transcriptional activation and pluripotency program initiation during human preimplantation development. Development 138, 3699–3709 (2011).
Marti, M. et al. Characterization of pluripotent stem cells. Nat. Protoc. 8, 223–253 (2013).
Padilla-Nash, H.M., Barenboim-Stapleton, L., Difilippantonio, M.J. & Ried, T. Spectral karyotyping analysis of human and mouse chromosomes. Nat. Protoc. 1, 3129–3142 (2006).
Gu, H. et al. Preparation of reduced representation bisulfite sequencing libraries for genome-scale DNA methylation profiling. Nat. Protoc. 6, 468–481 (2011).
Tomoda, K. et al. Derivation conditions impact X-inactivation status in female human induced pluripotent stem cells. Cell Stem Cell 11, 91–99 (2012).
Pasque, V. et al. X chromosome reactivation dynamics reveal stages of reprogramming to pluripotency. Cell 159, 1681–1697 (2014).
Acknowledgements
Z. Izsvák is funded by European Research Council (ERC) grant ERC-2011-AdG 294742. L.D.H. is funded by Medical Research grant MR/L007215/1. We thank E.È. Nagy for her assistance in specifying experimental details.
Author information
Authors and Affiliations
Contributions
Z. Izsvák, L.D.H. and J.W. wrote the manuscript. J.W. designed and performed experiments, and provided technical details to the Protocol. M.S. performed bioinformatic, cross-culture comparative studies. C. Sun assisted in cell culture and gene expression analysis. Z. Ivics provided critical advice. D.B. and A.P. provided materials.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 (PDF 360 kb)
Rights and permissions
About this article
Cite this article
Wang, J., Singh, M., Sun, C. et al. Isolation and cultivation of naive-like human pluripotent stem cells based on HERVH expression. Nat Protoc 11, 327–346 (2016). https://doi.org/10.1038/nprot.2016.016
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.016
This article is cited by
-
ZBTB12 is a molecular barrier to dedifferentiation in human pluripotent stem cells
Nature Communications (2023)
-
Naïve-like pluripotency to pave the way for saving the northern white rhinoceros from extinction
Scientific Reports (2022)
-
Mapping human pluripotent stem cell differentiation pathways using high throughput single-cell RNA-sequencing
Genome Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.