Abstract
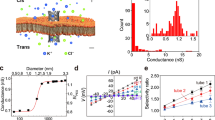
We describe a protocol for the insertion of ultrashort single-walled carbon nanotubes (SWCNTs) to form nanopores in a Montal-Mueller lipid bilayer. The SWCNTs are designed to bind to a specific analyte of interest; binding will result in the reduction of current in single-channel recording experiments. The first stage of the PROCEDURE is to cut and separate the SWCNTs. We cut long, purified SWCNTs with sonication in concentrated sulfuric acid/nitric acid (3/1). Isolation of ultrashort SWCNTs is carried out by size-exclusion HPLC separation. The second stage is to insert these short SWCNTs into the lipid bilayer. This step requires a microinjection probe made from a glass capillary. The setup for protein nanopore research can be adopted for the single-channel recording experiments without any special treatment. The obtained current traces are of very high quality, showing stable baselines and little background noise. Example procedures are shown for investigating ion transport and DNA translocation through these SWCNT nanopores. This nanopore has potential applications in molecular sensing, nanopore DNA sequencing and early disease diagnosis. For example, we have selectively detected modified 5-hydroxymethylcytosine in single-stranded DNA (ssDNA), which may have implications in screening specific genomic DNA sequences. The protocol takes ∼15 d, including SWCNT purification, cutting and separation, as well as the formation of SWCNT nanopores for DNA analyses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Clarke, J. et al. Continuous base identification for single-molecule nanopore DNA sequencing. Nat. Nanotechnol. 4, 265–270 (2009).
Cherf, G.M. et al. Automated forward and reverse ratcheting of DNA in a nanopore at 5-angstrom precision. Nat. Biotechnol. 30, 344–348 (2012).
Manrao, E.A. et al. Reading DNA at single-nucleotide resolution with a mutant MspA nanopore and phi29 DNA polymerase. Nat. Biotechnol. 30, 349–353 (2012).
Ashton, P.M. et al. MinION nanopore sequencing identifies the position and structure of a bacterial antibiotic resistance island. Nat. Biotechnol. 33, 296–300 (2015).
Jain, M. et al. Improved data analysis for the MinION nanopore sequencer. Nat. Methods 12, 351–356 (2015).
Bayley, H. & Cremer, P.S. Stochastic sensors inspired by biology. Nature 413, 226–230 (2001).
Howorka, S. & Siwy, Z. Nanopore analytics: sensing of single molecules. Chem. Soc. Rev. 38, 2360–2384 (2009).
Wanunu, M. et al. Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nat. Nanotechnol. 5, 807–814 (2010).
Wang, Y., Zheng, D.L., Tan, Q.L., Wang, M.X. & Gu, L.Q. Nanopore-based detection of circulating microRNAs in lung cancer patients. Nat. Nanotechnol. 6, 668 (2011).
Kasianowicz, J.J., Brandin, E., Branton, D. & Deamer, D.W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl. Acad. Sci. USA 93, 13770–13773 (1996).
Shin, S.H., Luchian, T., Cheley, S., Braha, O. & Bayley, H. Kinetics of a reversible covalent-bond-forming reaction observed at the single-molecule level. Angew. Chem. Int. Ed. 41, 3707–3709 (2002).
Luchian, T., Shin, S.H. & Bayley, H. Single-molecule covalent chemistry with spatially separated reactants. Angew. Chem. Int. Ed. 42, 3766–3771 (2003).
Braha, O. et al. Simultaneous stochastic sensing of divalent metal ions. Nat. Biotechnol. 18, 1005–1007 (2000).
Hammerstein, A.F., Shin, S.-H. & Bayley, H. Single-molecule kinetics of two-step divalent cation chelation. Angew. Chem. Int. Ed. 49, 5085–5090 (2010).
Wen, S. et al. Highly sensitive and selective DNA-based detection of mercury(II) with α-hemolysin nanopore. J. Am. Chem. Soc. 133, 18312–18317 (2011).
Yang, C. et al. Highly sensitive simultaneous detection of lead(II) and barium(II) with G-quadruplex DNA in α-hemolysin nanopore. Anal. Chem. 85, 7302–7307 (2013).
Gu, L.Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686–690 (1999).
Wu, H.-C. & Bayley, H. Single-molecule detection of nitrogen mustards by covalent reaction within a protein nanopore. J. Am. Chem. Soc. 130, 6813–6819 (2008).
Movileanu, L., Howorka, S., Braha, O. & Bayley, H. Detecting protein analytes that modulate transmembrane movement of a polymer chain within a single protein pore. Nat. Biotechnol. 18, 1091–1095 (2000).
Howorka, S., Cheley, S. & Bayley, H. Sequence-specific detection of individual DNA strands using engineered nanopores. Nat. Biotechnol. 19, 636–639 (2001).
Yusko, E.C. et al. Controlling protein translocation through nanopores with bio-inspired fluid walls. Nat. Nanotechnol. 6, 253–260 (2011).
Wei, R., Gatterdam, V., Wieneke, R., Tampe, R. & Rant, U. Stochastic sensing of proteins with receptor-modified solid-state nanopores. Nat. Nanotechnol. 7, 257–263 (2012).
Hladky, S.B. & Haydon, D.A. Discreteness of conductance change in bimolecular lipid membranes in presence of certain antibiotics. Nature 225, 451–453 (1970).
Ghadiri, M.R., Granja, J.R., Milligan, R.A., McRee, D.E. & Khazanovich, N. Self-assembling organic nanotubes based on a cyclic peptide architecture. Nature 366, 324–327 (1993).
Ghadiri, M.R., Granja, J.R. & Buehler, L.K. Artificial transmembrane ion channels from self-assembling peptide nanotubes. Nature 369, 301–304 (1994).
Bezrukov, S.M., Vodyanoy, I. & Parsegian, V.A. Counting polymers moving through a single-ion channel. Nature 370, 279–281 (1994).
Li, J. et al. Ion-beam sculpting at nanometre length scales. Nature 412, 166–169 (2001).
Storm, A.J., Chen, J.H., Ling, X.S., Zandbergen, H.W. & Dekker, C. Fabrication of solid-state nanopores with single-nanometre precision. Nat. Mater. 2, 537–540 (2003).
Dekker, C. Solid-state nanopores. Nat. Nanotechnol. 2, 209–215 (2007).
Butler, T.Z., Pavlenok, M., Derrington, I.M., Niederweis, M. & Gundlach, J.H. Single-molecule DNA detection with an engineered MspA protein nanopore. Proc. Natl. Acad. Sci. USA 105, 20647–20652 (2008).
Wendell, D. et al. Translocation of double-stranded DNA through membrane-adapted phi29 motor protein nanopores. Nat. Nanotechnol. 4, 765–772 (2009).
Mohammad, M.M. et al. Engineering a rigid protein tunnel for biomolecular detection. J. Am. Chem. Soc. 134, 9521–9531 (2012).
Soskine, M. et al. An engineered ClyA nanopore detects folded target proteins by selective external association and pore entry. Nano Lett. 12, 4895–4900 (2012).
Liu, H. et al. Translocation of single-stranded DNA through single-walled carbon nanotubes. Science 327, 64–67 (2010).
Liu, L., Yang, C., Zhao, K., Li, J. & Wu, H.C. Ultrashort single-walled carbon nanotubes in a lipid bilayer as a new nanopore sensor. Nat. Commun. 4, 2989 (2013).
Geng, J. et al. Stochastic transport through carbon nanotubes in lipid bilayers and live cell membranes. Nature 514, 612–615 (2014).
Langecker, M. et al. Synthetic lipid membrane channels formed by designed DNA nanostructures. Science 338, 932–936 (2012).
Burns, J.R., Stulz, E. & Howorka, S. Self-assembled DNA nanopores that span lipid bilayers. Nano Lett. 13, 2351–2356 (2013).
Matile, S., Jentzsch, A.V., Montenegro, J. & Fin, A. Recent synthetic transport systems. Chem. Soc. Rev. 40, 2453–2474 (2011).
Takeuchi, T. & Matile, S. Sensing applications of synthetic transport systems. Chem. Commun. 49, 19–29 (2013).
Montenegro, J., Ghadiri, M.R. & Granja, J.R. Ion channel models based on self-assembling cyclic peptide nanotubes. Acc. Chem. Res. 46, 2955–2965 (2013).
Garaj, S. et al. Graphene as a subnanometre trans-electrode membrane. Nature 467, 190–194 (2010).
Schneider, G.F. et al. DNA translocation through graphene nanopores. Nano Lett. 10, 3163–3167 (2010).
Merchant, C.A. et al. DNA translocation through graphene nanopores. Nano Lett. 10, 2915–2921 (2010).
Liu, S. et al. Boron nitride nanopores: highly sensitive DNA single-molecule detectors. Adv. Mater. 25, 4549–4554 (2013).
Liu, K., Feng, J., Kis, A. & Radenovic, A. Atomically thin molybdenum disulfide nanopores with high sensitivity for DNA translocation. ACS Nano 8, 2504–2511 (2014).
Holden, M.A. & Bayley, H. Direct introduction of single protein channels and pores into lipid bilayers. J. Am. Chem. Soc. 127, 6502–6503 (2005).
Holden, M.A., Jayasinghe, L., Daltrop, O., Mason, A. & Bayley, H. Direct transfer of membrane proteins from bacteria to planar bilayers for rapid screening by single-channel recording. Nat. Chem. Biol. 2, 314–318 (2006).
Lee, C.Y., Choi, W., Han, J.-H. & Strano, M.S. Coherence resonance in a single-walled carbon nanotube ion channel. Science 329, 1320–1324 (2010).
Wu, J., Gerstandt, K., Zhang, H., Liu, J. & Hinds, B.J. Electrophoretically induced aqueous flow through single-walled carbon nanotube membranes. Nat. Nanotechnol. 7, 133–139 (2012).
Yang, F. et al. Chirality-specific growth of single-walled carbon nanotubes on solid alloy catalysts. Nature 510, 522–524 (2014).
Tanaka, M. & Sackmann, E. Polymer-supported membranes as models of the cell surface. Nature 437, 656–663 (2005).
Holt, J.K. et al. Fast mass transport through sub-2-nanometer carbon nanotubes. Science 312, 1034–1037 (2006).
Maglia, G., Heron, A.J., Stoddart, D., Japrung, D. & Bayley, H. Analysis of single nucleic acid molecules with protein nanopores. Methods Enzymol. 475, 591–623 (2010).
Acknowledgements
This work was funded by the National Basic Research Program of China (973 program, grant no. 2013CB932800), the National Natural Science Foundation of China (grant nos. 21175135, 21375130, 21205119 and 21475132) and the Chinese Academy of Sciences Hundred Talents Program.
Author information
Authors and Affiliations
Contributions
L.L. and H.-C.W. conceived the project; L.L., J.X. and T.L. performed the experiments; L.L., T.L. and H.-C.W. performed data analysis; and L.L., J.X. and H.-C.W. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Micro-injection probe interaction with the aperture
This movie illustrates how the micro-injection probe is controlled to interact with the PTFE aperture in the absence of buffer solutions. The round aperture is about 100 μm in diameter over which the lipid bilayer forms. The distance between the probe tip and the aperture can be precisely controlled. Starting from 40 s, the micro-injector is shown to inject solutions from the probe syringe. (MOV 3338 kb)
Rights and permissions
About this article
Cite this article
Liu, L., Xie, J., Li, T. et al. Fabrication of nanopores with ultrashort single-walled carbon nanotubes inserted in a lipid bilayer. Nat Protoc 10, 1670–1678 (2015). https://doi.org/10.1038/nprot.2015.112
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.112
This article is cited by
-
Construction of an aerolysin nanopore in a lipid bilayer for single-oligonucleotide analysis
Nature Protocols (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.