Abstract
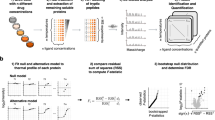
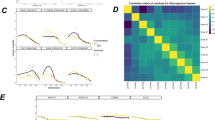
The direct detection of drug-protein interactions in living cells is a major challenge in drug discovery research. Recently, we introduced an approach termed thermal proteome profiling (TPP), which enables the monitoring of changes in protein thermal stability across the proteome using quantitative mass spectrometry. We determined the intracellular thermal profiles for up to 7,000 proteins, and by comparing profiles derived from cultured mammalian cells in the presence or absence of a drug we showed that it was possible to identify direct and indirect targets of drugs in living cells in an unbiased manner. Here we demonstrate the complete workflow using the histone deacetylase inhibitor panobinostat. The key to this approach is the use of isobaric tandem mass tag 10-plex (TMT10) reagents to label digested protein samples corresponding to each temperature point in the melting curve so that the samples can be analyzed by multiplexed quantitative mass spectrometry. Important steps in the bioinformatic analysis include data normalization, melting curve fitting and statistical significance determination of compound concentration-dependent changes in protein stability. All analysis tools are made freely available as R and Python packages. The workflow can be completed in 2 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Schenone, M., Dancik, V., Wagner, B.K. & Clemons, P.A. Target identification and mechanism of action in chemical biology and drug discovery. Nat. Chem. Biol. 9, 232–240 (2013).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Jafari, R. et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 9, 2100–2122 (2014).
Linderstrøm-Lang, K. & Schellman, J.A. Protein structure and enzyme activity. Enzymes 1, 443–510 (1959).
Pantoliano, M.W. et al. High-density miniaturized thermal shift assays as a general strategy for drug discovery. J. Biomol. Screen. 6, 429–440 (2001).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Becher, I. et al. Chemoproteomics reveals time-dependent binding of histone deacetylase inhibitors to endogenous repressor complexes. ACS Chem. Biol. 9, 1736–1746 (2014).
Becher, I. et al. Affinity profiling of the cellular kinome for the nucleotide cofactors ATP, ADP, and GTP. ACS Chem. Biol. 8, 599–607 (2013).
Huang, J. Tracking drugs. N. Engl. J. Med. 369, 1168–1169 (2013).
Werner, T. et al. High-resolution enabled TMT 8-plexing. Anal. Chem. 84, 7188–7194 (2012).
Werner, T. et al. Ion coalescence of neutron encoded TMT 10-plex reporter ions. Anal. Chem. 86, 3594–3601 (2014).
Savitski, M.M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Oda, T. et al. Crkl is the major tyrosine-phosphorylated protein in neutrophils from patients with chronic myelogenous leukemia. J. Biol. Chem. 269, 22925–22928 (1994).
Bantscheff, M., Lemeer, S., Savitski, M.M. & Kuster, B. Quantitative mass spectrometry in proteomics: critical review update from 2007 to the present. Anal. Bioanal. Chem. 404, 939–965 (2012).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Rauniyar, N. & Yates, J.R. 3rd Isobaric labeling-based relative quantification in shotgun proteomics. J. Proteome Res. 13, 5293–5309 (2014).
Atadja, P. Development of the pan-DAC inhibitor panobinostat (LBH589): successes and challenges. Cancer Lett. 280, 233–241 (2009).
Moffat, J.G., Rudolph, J. & Bailey, D. Phenotypic screening in cancer drug discovery—past, present and future. Nat. Rev. Drug Discov. 13, 588–602 (2014).
Paul, S.M. et al. How to improve R&D productivity: the pharmaceutical industry's grand challenge. Nat. Rev. Drug Discov. 9, 203–214 (2010).
Roberts, R.A. et al. Reducing attrition in drug development: smart loading preclinical safety assessment. Drug Discov. Today 19, 341–347 (2014).
Anighoro, A., Bajorath, J. & Rastelli, G. Polypharmacology: challenges and opportunities in drug discovery. J. Med. Chem. 57, 7874–7887 (2014).
Keiser, M.J. et al. Predicting new molecular targets for known drugs. Nature 462, 175–181 (2009).
Jalencasa, X. & Mestres, J. On the origins of drug polypharmacology. Med. Chem. Commun. 4, 80–87 (2013).
Knight, Z.A., Lin, H. & Shokat, K.M. Targeting the cancer kinome through polypharmacology. Nat. Rev. Cancer 10, 130–137 (2010).
Asial, I. et al. Engineering protein thermostability using a generic activity-independent biophysical screen inside the cell. Nat. Commun. 4, 2901 (2013).
Miettinen, T.P. & Bjorklund, M. NQO2 is a reactive oxygen species generating off-target for acetaminophen. Mol. Pharm. 11, 4395–4404 (2014).
Kruse, U. et al. Chemoproteomics-based kinome profiling and target deconvolution of clinical multi-kinase inhibitors in primary chronic lymphocytic leukemia cells. Leukemia 25, 89–100 (2011).
Michalski, A. et al. Mass spectrometry-based proteomics using Q Exactive, a high-performance benchtop quadrupole Orbitrap mass spectrometer. Mol. Cell. Proteomics 10, M111.011015 (2011).
Olsen, J.V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Dayon, L. et al. Relative quantification of proteins in human cerebrospinal fluids by MS/MS using 6-plex isobaric tags. Anal. Chem. 80, 2921–2931 (2008).
Unwin, R.D., Griffiths, J.R. & Whetton, A.D. Simultaneous analysis of relative protein expression levels across multiple samples using iTRAQ isobaric tags with 2D nano LC-MS/MS. Nat. Protoc. 5, 1574–1582 (2010).
Ting, L., Rad, R., Gygi, S.P. & Haas, W. MS3 eliminates ratio distortion in isobaric multiplexed quantitative proteomics. Nat. Methods 8, 937–940 (2011).
McAlister, G.C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Ow, S.Y. et al. iTRAQ underestimation in simple and complex mixtures: “the good, the bad and the ugly”. J. Proteome Res. 8, 5347–5355 (2009).
Savitski, M.M. et al. Measuring and managing ratio compression for accurate iTRAQ/TMT quantification. J. Proteome Res. 12, 3586–3598 (2013).
Savitski, M.M. et al. Targeted data acquisition for improved reproducibility and robustness of proteomic mass spectrometry assays. J. Am. Soc. Mass Spectrom. 21, 1668–1679 (2010).
Savitski, M.M. et al. Delayed fragmentation and optimized isolation width settings for improvement of protein identification and accuracy of isobaric mass tag quantification on Orbitrap-type mass spectrometers. Anal. Chem. 83, 8959–8967 (2011).
Lemeer, S., Hahne, H., Pachl, F. & Kuster, B. Software tools for MS-based quantitative proteomics: a brief overview. Methods Mol. Biol. 893, 489–499 (2012).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. A practical guide to the MaxQuant computational platform for SILAC-based quantitative proteomics. Nat. Protoc. 4, 698–705 (2009).
Colaert, N. et al. Thermo-msf-parser: an open source Java library to parse and visualize Thermo Proteome Discoverer msf files. J. Proteome Res. 10, 3840–3843 (2011).
Wilhelm, M., Kirchner, M., Steen, J.A. & Steen, H. mz5: space- and time-efficient storage of mass spectrometry data sets. Mol. Cell. Proteomics 11, O111.011379 (2012).
Savitski, M.M., Mathieson, T., Becher, I. & Bantscheff, M. H-score, a mass accuracy driven rescoring approach for improved peptide identification in modification rich samples. J. Proteome Res. 9, 5511–5516 (2010).
Nielsen, M.L., Savitski, M.M. & Zubarev, R.A. Improving protein identification using complementary fragmentation techniques in Fourier transform mass spectrometry. Mol. Cell. Proteomics 4, 835–845 (2005).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Kocher, T., Pichler, P., Swart, R. & Mechtler, K. Analysis of protein mixtures from whole-cell extracts by single-run nanoLC-MS/MS using ultralong gradients. Nat. Protoc. 7, 882–890 (2012).
Chittur, S.V., Sangster-Guity, N. & McCormick, P.J. Histone deacetylase inhibitors: a new mode for inhibition of cholesterol metabolism. BMC Genomics 9, 507 (2008).
Acknowledgements
We thank M. Jundt, K. Kammerer, M. Klös-Hudak, M. Paulmann and T. Rudi for expert technical assistance; F. Weisbrodt for help with the figures; and R. Heinkel for expert advice regarding packaging of the isobarQuant software. We are grateful to G. Neubauer for discussions and support.
Author information
Authors and Affiliations
Contributions
H.F., D.C., T.M., F.B.M.R., W.H. and M.M.S. conceived the project and wrote the manuscript; M.M.S., F.B.M.R., T.W., M.B. and G.D. designed the mass spectrometry experiments; T.W. and I.T. conducted and supervised the experiments; H.F., T.M., D.C., G.M.A.S., T.W., C.D., F.B.M.R. and M.M.S. analyzed proteomics data; T.M. and G.M.A.S. developed the isobarQuant package; D.C., H.F., C.D., S.G. and W.H. developed the TPP package; T.W., I.T., G.M.A.S., M.B. and G.D. contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.F., T.M., G.M.A.S., T.W., I.T., C.D., S.G., M.B., G.D., F.B.M.R. and M.M.S. are employees and/or shareholders of Cellzome and GlaxoSmithKline.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Manual (PDF 3363 kb)
Supplementary Data 1
Results output of the TPP package of the analysis of the TPP-TR panobinostat experiment. (XLSX 9553 kb)
Supplementary Data 2
Results output of the TPP package of the analysis of the TPP-CCR panobinostat experiment. (XLSX 2330 kb)
Rights and permissions
About this article
Cite this article
Franken, H., Mathieson, T., Childs, D. et al. Thermal proteome profiling for unbiased identification of direct and indirect drug targets using multiplexed quantitative mass spectrometry. Nat Protoc 10, 1567–1593 (2015). https://doi.org/10.1038/nprot.2015.101
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.101
This article is cited by
-
The Golgi stacking protein GRASP55 is targeted by the natural compound prodigiosin
Cell Communication and Signaling (2023)
-
USP7/Maged1-mediated H2A monoubiquitination in the paraventricular thalamus: an epigenetic mechanism involved in cocaine use disorder
Nature Communications (2023)
-
Sinomenine ameliorates collagen-induced arthritis in mice by targeting GBP5 and regulating the P2X7 receptor to suppress NLRP3-related signaling pathways
Acta Pharmacologica Sinica (2023)
-
Systematic analysis of drug combinations against Gram-positive bacteria
Nature Microbiology (2023)
-
Experimental strategies to improve drug-target identification in mass spectrometry-based thermal stability assays
Communications Chemistry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.