Abstract
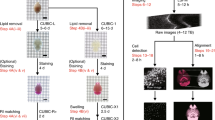
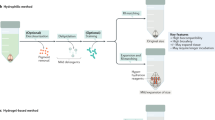
Here we describe a protocol for advanced CUBIC (Clear, Unobstructed Brain/Body Imaging Cocktails and Computational analysis). The CUBIC protocol enables simple and efficient organ clearing, rapid imaging by light-sheet microscopy and quantitative imaging analysis of multiple samples. The organ or body is cleared by immersion for 1–14 d, with the exact time required dependent on the sample type and the experimental purposes. A single imaging set can be completed in 30–60 min. Image processing and analysis can take <1 d, but it is dependent on the number of samples in the data set. The CUBIC clearing protocol can process multiple samples simultaneously. We previously used CUBIC to image whole-brain neural activities at single-cell resolution using Arc-dVenus transgenic (Tg) mice. CUBIC informatics calculated the Venus signal subtraction, comparing different brains at a whole-organ scale. These protocols provide a platform for organism-level systems biology by comprehensively detecting cells in a whole organ or body.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Keller, P.J. & Ahrens, M.B. Visualizing whole-brain activity and development at the single-cell level using light-sheet microscopy. Neuron 85, 462–483 (2015).
Osten, P. & Margrie, T.W. Mapping brain circuitry with a light microscope. Nat. Methods 10, 515–523 (2013).
Spalteholz, W. Über das Durchsichtigmachen von menschlichen und tierischen Präparaten (S. Hirzel, 1914).
Dodt, H.U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Ertürk, A. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 7, 1983–1995 (2012).
Becker, K., Jährling, N., Saghafi, S., Weiler, R. & Dodt, H.U. Chemical clearing and dehydration of GFP expressing mouse brains. PLoS ONE 7, e33916 (2012).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Schwarz, M.K. et al. Fluorescent-protein stabilization and high-resolution imaging of cleared, intact mouse brains. PLoS ONE 10, e0124650 (2015).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Ke, M.T., Fujimoto, S. & Imai, T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 16, 1154–1161 (2013).
Kuwajima, T. et al. ClearT: a detergent- and solvent-free clearing method for neuronal and non-neuronal tissue. Development 140, 1364–1368 (2013).
Aoyagi, Y., Kawakami, R., Osanai, H., Hibi, T. & Nemoto, T. A rapid optical clearing protocol using 2,2′-thiodiethanol for microscopic observation of fixed mouse brain. PLoS ONE 10, e0116280 (2015).
Costantini, I. et al. A versatile clearing agent for multi-modal brain imaging. Sci. Rep. 5, 9808 (2015).
Hou, B. et al. Scalable and DiI-compatible optical clearance of the mammalian brain. Front. Neuroanat. 9, 19 (2015).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Tomer, R., Ye, L., Hsueh, B. & Deisseroth, K. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat. Protoc. 9, 1682–1697 (2014).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Susaki, E.A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Keller, P.J., Schmidt, A.D., Wittbrodt, J. & Stelzer, E.H.K. Reconstruction of zebrafish early embryonic development by scanned light sheet microscopy. Science 322, 1065–1069 (2008).
Keller, P.J. et al. Fast, high-contrast imaging of animal development with scanned light sheet-based structured-illumination microscopy. Nat. Methods 7, 637–642 (2010).
Ahrens, M.B., Orger, M.B., Robson, D.N., Li, J.M. & Keller, P.J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat. Methods 10, 413–420 (2013).
Panier, T. et al. Fast functional imaging of multiple brain regions in intact zebrafish larvae using selective plane illumination microscopy. Front. Neural Circuits 7, 65 (2013).
Murphy, K. et al. Evaluation of registration methods on thoracic CT: the EMPIRE10 challenge. IEEE Trans. Med. Imaging 30, 1901–1920 (2011).
Yushkevich, P.A. et al. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31, 1116–1128 (2006).
Eguchi, M. & Yamaguchi, S. In vivo and in vitro visualization of gene expression dynamics over extensive areas of the brain. Neuroimage 44, 1274–1283 (2009).
Prewitt, J.M.S. Object enhancement and extraction. In Picture Processing and Psychopictorics (eds. Lipkin, B.S. & Rosenfeld, A.) 75–149 (Academic Press, 1970).
Jenkinson, M., Beckmann, C.F., Behrens, T.E., Woolrich, M.W. & Smith, S.M. Fsl. Neuroimage 62, 782–790 (2012).
Lein, E.S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Alnuami, A.A., Zeedi, B., Qadri, S.M. & Ashraf, S.S. Oxyradical-induced GFP damage and loss of fluorescence. Int. J. Biol. Macromol. 43, 182–186 (2008).
Steinke, H. & Wolff, W. A modified Spalteholz technique with preservation of the histology. Ann. Anat. 183, 91–95 (2001).
Faisal, A.A., White, J.A. & Laughlin, S.B. Ion-channel noise places limits on the miniaturization of the brain's wiring. Curr. Biol. 15, 1143–1149 (2005).
Richards, K.L. et al. Hippocampal volume and cell density changes in a mouse model of human genetic epilepsy. Neurology 80, 1240–1246 (2013).
Feng, G.P. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Okabe, M., Ikawa, M., Kominami, K., Nakanishi, T. & Nishimune, Y. 'Green mice' as a source of ubiquitous green cells. FEBS Lett. 407, 313–319 (1997).
Ertürk, A. et al. Three-dimensional imaging of the unsectioned adult spinal cord to assess axon regeneration and glial responses after injury. Nat. Med. 18, 166–171 (2012).
Soderblom, C. et al. 3D imaging of axons in transparent spinal cords from rodents and nonhuman primates. eNeuro 2 doi:10.1523/ENEURO.0001-15.2015 (2015).
Weber, T.G. et al. Apoptosis imaging for monitoring DR5 antibody accumulation and pharmacodynamics in brain tumors noninvasively. Cancer Res. 74, 1913–1923 (2014).
Acknowledgements
We thank the lab members at RIKEN QBiC and The University of Tokyo, in particular S.I. Kubota for his kind help in preparing the materials; A. Millius for his critical reading and editing of the manuscript; and T. Mano for his kind contributions and suggestions to discuss image resolution. This work was supported by the Program for Innovative Cell Biology by Innovative Technology and the Brain Mapping by Integrated Neurotechnologies for Disease Studies (Brain/MINDS) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan; a Grant-in-Aid for Scientific Research (S) (grant No. 25221004), for Scientific Research on Innovative Areas (grant no. 23115006) and for Young Scientists (A) (grant no. 15H05650) from MEXT/Japan Society for the Promotion of Science (JSPS); by the strategic programs for R&D (President's discretionary fund) of RIKEN; by an intramural Grant-in-Aid from the RIKEN QBiC; by a grant from AMED-CREST; by the RIKEN Special Postdoctoral Research Program; by the RIKEN Foreign Postdoctoral Researcher Program; by a Grant-in-Aid from the Japan Foundation for Applied Enzymology; by the Brain Sciences Project of the Center for Novel Science Initiatives of the National Institutes of Natural Sciences (grant nos. BS261004 and BS271005); by the Tokyo Society of Medical Science; and by the Shimabara Science Promotion Foundation.
Author information
Authors and Affiliations
Contributions
H.R.U., E.A.S., K.T. and D.P. designed the study. E.A.S., H.Y. and A.K. performed most of the immersion protocol. K.T. performed most of the CB-perfusion protocol. D.P. performed most of the computational image analysis. A.K. developed the improved immersion protocol. All authors discussed the results and commented on the manuscript text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data
Scripts and guide for software described. (ZIP 49 kb)
Spot-counting analysis of hippocampal neurons in Thy1-YFP-H Tg mouse brain shown in Fig. 4b-f.
Almost all cells are roughly detected as single spots in such a relatively dense region. The analysis was performed with Imaris software. Some of parameters are manually adjusted. (MP4 26325 kb)
Rights and permissions
About this article
Cite this article
Susaki, E., Tainaka, K., Perrin, D. et al. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat Protoc 10, 1709–1727 (2015). https://doi.org/10.1038/nprot.2015.085
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.085
This article is cited by
-
Dual electrical stimulation at spinal-muscular interface reconstructs spinal sensorimotor circuits after spinal cord injury
Nature Communications (2024)
-
ISL1 controls pancreatic alpha cell fate and beta cell maturation
Cell & Bioscience (2023)
-
Dysregulation of hypoxia-inducible factor 1α in the sympathetic nervous system accelerates diabetic cardiomyopathy
Cardiovascular Diabetology (2023)
-
Whole-mouse clearing and imaging at the cellular level with vDISCO
Nature Protocols (2023)
-
Brain regulation of gastric dysfunction induced by stress
Nature Metabolism (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.