Abstract
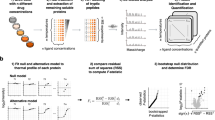
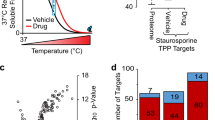
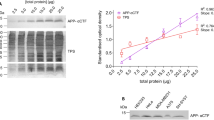
Thermal shift assays are used to study thermal stabilization of proteins upon ligand binding. Such assays have been used extensively on purified proteins in the drug discovery industry and in academia to detect interactions. Recently, we published a proof-of-principle study describing the implementation of thermal shift assays in a cellular format, which we call the cellular thermal shift assay (CETSA). The method allows studies of target engagement of drug candidates in a cellular context, herein exemplified with experimental data on the human kinases p38α and ERK1/2. The assay involves treatment of cells with a compound of interest, heating to denature and precipitate proteins, cell lysis, and the separation of cell debris and aggregates from the soluble protein fraction. Whereas unbound proteins denature and precipitate at elevated temperatures, ligand-bound proteins remain in solution. We describe two procedures for detecting the stabilized protein in the soluble fraction of the samples. One approach involves sample workup and detection using quantitative western blotting, whereas the second is performed directly in solution and relies on the induced proximity of two target-directed antibodies upon binding to soluble protein. The latter protocol has been optimized to allow an increased throughput, as potential applications require large numbers of samples. Both approaches can be completed in a day.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Brandts, J.F. & Lin, L.N. Study of strong to ultratight protein interactions using differential scanning calorimetry. Biochemistry 29, 6927–6940 (1990).
Crothers, D.M. Statistical thermodynamics of nucleic acid melting transitions with coupled binding equilibria. Biopolymers 10, 2147–2160 (1971).
Schellman, J.A. Macromolecular binding. Biopolymers 14, 999–1018 (1975).
Matulis, D., Kranz, J.K., Salemme, F.R. & Todd, M.J. Thermodynamic stability of carbonic anhydrase: measurements of binding affinity and stoichiometry using ThermoFluor. Biochemistry 44, 5258–5266 (2005).
Ericsson, U.B., Hallberg, B.M., Detitta, G.T., Dekker, N. & Nordlund, P. ThermoFluor-based high-throughput stability optimization of proteins for structural studies. Anal. Biochem. 357, 289–298 (2006).
Niesen, F.H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Pantoliano, M.W. et al. High-density miniaturized thermal shift assays as a general strategy for drug discovery. J. Biomol. Screen. 6, 429–440 (2001).
Senisterra, G.A. et al. Screening for ligands using a generic and high-throughput light-scattering-based assay. J. Biomol. Screen. 11, 940–948 (2006).
Fedorov, O. et al. A systematic interaction map of validated kinase inhibitors with Ser/Thr kinases. Proc. Natl. Acad. Sci. USA 104, 20523–20528 (2007).
Vedadi, M. et al. Chemical screening methods to identify ligands that promote protein stability, protein crystallization, and structure determination. Proc. Natl. Acad. Sci. USA 103, 15835–15840 (2006).
Garnett, M.J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Feng, B.Y. & Shoichet, B.K. A detergent-based assay for the detection of promiscuous inhibitors. Nat. Protoc. 1, 550–553 (2006).
Walters, W.P. & Namchuk, M. Designing screens: how to make your hits a hit. Nat. Rev. Drug Disc. 2, 259–266 (2003).
Nordlund, P. Methods for determining ligand binding to a target protein using a thermal shift assay. UK patent no. GB2490404 (2012).
Weibrecht, I. et al. Proximity ligation assays: a recent addition to the proteomics toolbox. Exp. Rev. Proteom. 7, 401–409 (2010).
Knaust, R.K. & Nordlund, P. Screening for soluble expression of recombinant proteins in a 96-well format. Anal. Biochem. 297, 79–85 (2001).
Asial, I. et al. Engineering protein thermostability using a generic activity-independent biophysical screen inside the cell. Nat. Commun. 4, 2901 (2013).
Branca, R.M et al. HiRIEF LC-MS enables deep proteome coverage and unbiased proteogenomics. Nat. Methods 11, 59–62 (2014).
Eglen, R.M. et al. The use of AlphaScreen technology in HTS: current status. Curr. Chem. Genomics 1, 2–10 (2008).
Bazin, H., Preaudat, M., Trinquet, E. & Mathis, G. Homogeneous time resolved fluorescence resonance energy transfer using rare earth cryptates as a tool for probing molecular interactions in biology. Spectrochim. Acta. A Mol. Biomol. Spectrosc. 57, 2197–2211 (2001).
Osmond, R.I.W., Das, S. & Crouch, M.F. Development of cell-based assays for cytokine receptor signaling, using an AlphaScreen SureFire assay format. Anal. Biochem. 403, 94–9101 (2010).
Bembenek, M.E. et al. Determination of complementary antibody pairs using protein A capture with the AlphaScreen assay format. Anal. Biochem. 408, 321–327 (2011).
Copeland, R.A., Pompliano, D.L. & Meek, T.D. Drug-target residence time and its implications for lead optimization. Nat. Rev. Drug Disc. 5, 730–739 (2006).
Lepock, J.R. et al. Influence of transition rates and scan rate on kinetic simulations of differential scanning calorimetry profiles of reversible and irreversible protein denaturation. Biochemistry 31, 12706–12712 (1992).
Quistgaard, E.M. et al. Structural and biophysical characterization of the cytoplasmic domains of human BAP29 and BAP31. PLoS ONE 8, e71111 (2013).
Saklatvala, J. et al. Role for p38 mitogen-activated protein kinase in platelet aggregation caused by collagen or a thromboxane analogue. J. Biol. Chem. 271, 6586–6589 (1996).
Dominguez, C., Powers, D.A. & Tamayo, N. p38 MAP kinase inhibitors: many are made, but few are chosen. Curr. Opin. Drug Disc. Dev. 8, 421–430 (2005).
Aronov, A.M. et al. Structure-guided design of potent and selective pyrimidylpyrrole inhibitors of extracellular signal-regulated kinase (ERK) using conformational control. J. Med. Chem. 52, 6362–6368 (2009).
Zhang, J.H., Chung, T.D. & Oldenburg, K.R. A simple statistical parameter for use in evaluation and validation of high-throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Richman, D.D., Cleveland, P.H., Oxman, M.N. & Johnson, K.M. The binding of staphylococcal protein A by the sera of different animal species. J. Immunol. 128, 2300–2305 (1982).
Acknowledgements
D.M.M., R.J., M.I. and P.N. acknowledge Karolinska Institutet (Distinguished Professor Award), the Swedish Research Council (Vetenskapsrådet) and the Swedish Cancer Society (Cancerfonden). T.L., H. Almqvist and H. Axelsson acknowledge Karolinska Institutet and the Swedish Research Council, which funds Chemical Biology Consortium Sweden, a national infrastructure for research in the field of chemical Biology, as well as Biovitrum for a generous donation of instruments and compound libraries when Chemical Biology Consortium Sweden was inaugurated. We are also grateful to C. Zaniol at PerkinElmer for great support in the development of the AlphaScreen assays.
Author information
Authors and Affiliations
Contributions
R.J., H. Axelsson, H. Almqvist, M.I., T.L., P.N. and D.M.M. conceived the study; R.J., H. Axelsson, H. Almqvist, T.L. and D.M.M. designed the experiments and the high-throughput formatting of the CETSA procedure; western blot and the AlphaScreen experiments were conducted by R.J., D.M.M., H. Axelsson, H. Almqvist and T.L.; and the manuscript was prepared by R.J., H. Axelsson, H. Almqvist, M.I., T.L., P.N. and D.M.M.
Corresponding authors
Ethics declarations
Competing interests
D.M.M. and P.N. are the founders of Pelago Bioscience AB. The remaining authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cellular integrity after heating.
In order to evaluate the integrity of the cell membrane of HL-60 cells during heating a trypan blue dye exclusion experiment was performed. Aliquots of approximately 3.0x104 HL-60 cells in culture medium were heated to 37, 50, 60, 65 and 70°C respectively for 3 min in the presence of 0.2% (w/v) trypan blue dye and allowed to cool to room temperature for 3 minutes. A 10μl aliquot of the heated cell suspension was subsequently analyzed using a TC20™ automated cell counter. Cells with the ability to exclude trypan blue were considered as retaining their cell membrane integrity whereas dye stained cells were identified as cells with disrupted cell membranes. The data demonstrated that the cell membrane of the HL-60 cells remain intact during heating at 50°C (the temperature at which the ITDRFCETSA experiments were conducted) since no statistically significant decrease in dye exclusion was observed compared to the initial dye exclusion capability (control) of the HL-60 cells. In addition, the data demonstrated that the HL-60 cell membranes are not disrupted unless temperatures above 60°C are reached. This is in agreement with our previous study where several other cell lines demonstrated loss of cell membrane integrity at temperatures above 60°C1.Data are given as the average ± standard deviation (n ≥ 4).
Supplementary Figure 2 Chemical structures.
The chemical structures of the p38α inhibitors; AMG-548 (A), SB203580 (B) and the dual Erk1/2 inhibitor ERK 11e (C).
Supplementary Figure 3 Negative control compound on p38α.
Western blot image illustrating remaining amounts of p38α in the presence of a dual Erk1/2 inhibitor, ERK 11e, in HL-60 cells as a function of temperature. All experiments were performed at three independent occasions and the experiments loaded and assayed in chorus using Western blot.
Supplementary Figure 4 ITDRFCETSA confirmation of CBK derived hit compounds.
An ITDRFCETSA experiment involving the human kinase p38α was performed to verify intracellular target binding of CBK200177 and CBK107148, both of which were identified in Box 4 (High throughput screening procedure for identification of stabilizers of intracellular p38α). Data are presented as the fold increase in AlphaScreen signal for each concentration of compound compared to DMSO controls. The data demonstrate dose dependent thermal stabilization of p38α for both compounds, thus confirming the results from the “High throughput screening procedure for identification of stabilizers of intracellular p38α”.
Supplementary information
Supplementary Figure 1
Cellular integrity after heating. (PDF 176 kb)
Supplementary Figure 2
Chemical structures. (PDF 224 kb)
Supplementary Figure 3
Negative control compound on p38α. (PDF 233 kb)
Supplementary Figure 4
ITDRFCETSA confirmation of CBK derived hit compounds. (PDF 260 kb)
Supplementary Data
Positive control compound on ERK1/2. (PDF 445 kb)
Rights and permissions
About this article
Cite this article
Jafari, R., Almqvist, H., Axelsson, H. et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat Protoc 9, 2100–2122 (2014). https://doi.org/10.1038/nprot.2014.138
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.138
This article is cited by
-
Dopamine modification of glycolytic enzymes impairs glycolysis: possible implications for Parkinson’s disease
Cell Communication and Signaling (2024)
-
Structure-guided design of a selective inhibitor of the methyltransferase KMT9 with cellular activity
Nature Communications (2024)
-
Activation of Ca2+ phosphatase Calcineurin regulates Parkin translocation to mitochondria and mitophagy in flies
Cell Death & Differentiation (2024)
-
Deep learning based CETSA feature prediction cross multiple cell lines with latent space representation
Scientific Reports (2024)
-
Deep learning enables the discovery of a novel cuproptosis-inducing molecule for the inhibition of hepatocellular carcinoma
Acta Pharmacologica Sinica (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.