Abstract
Tobacco rattle virus (TRV)-based virus-induced gene silencing (VIGS) is widely used in various plant species to downregulate the expression of a target plant gene. TRV is a bipartite, positive-strand RNA virus with the TRV1 and TRV2 genomes. To induce post-transcriptional gene silencing (PTGS), the TRV2 genome is genetically modified to carry a fragment of the target gene and delivered into the plant (along with the TRV1 genome) by agroinoculation. TRV1- and TRV2-carrying Agrobacterium strains are then co-inoculated into 3-week-old plant leaves by one of three methods: a needleless syringe, the agrodrench method or by pricking with a toothpick. Target gene silencing occurs in the newly developed noninoculated leaves within 2–3 weeks of TRV inoculation. The TRV-VIGS protocol described here takes only 4 weeks to implement, and it is faster and easier to perform than other gene silencing techniques that are currently available. Although we use Nicotiana benthamiana as an example, the protocol is adaptable to other plant species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hamilton, J.P. & Robin Buell, C. Advances in plant genome sequencing. Plant J. 70, 177–190 (2012).
Mochida, K. & Shinozaki, K. Advances in omics and bioinformatics tools for systems analyses of plant functions. Plant Cell Physiol. 52, 2017–2038 (2011).
Alonso, J.M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Tsai, H. et al. Discovery of rare mutations in populations: TILLING by sequencing. Plant Physiol. 156, 1257–1268 (2011).
Hilson, P. et al. Versatile gene-specific sequence tags for Arabidopsis functional genomics: transcript profiling and reverse genetics applications. Genome Res. 14, 2176–2189 (2004).
Wesley, S.V. et al. Construct design for efficient, effective and high-throughput gene silencing in plants. Plant J. 27, 581–590 (2001).
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18, 1121–1133 (2006).
de Felippes, F.F., Wang, J.-w. & Weigel, D. MIGS: miRNA-induced gene silencing. Plant J. 70, 541–547 (2012).
Lloyd, A., Plaisier, C.L., Carroll, D. & Drews, G.N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc. Natl. Acad. Sci. USA 102, 2232–2237 (2005).
Zhang, Y. et al. Transcription activator-like effector nucleases enable efficient plant genome engineering. Plant Physiol. 161, 20–27 (2013).
Nekrasov, V., Staskawicz, B., Weigel, D., Jones, J. & Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotech. 31, 691–693 (2013).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotech. 31, 686–688 (2013).
Lu, R., Martin-Hernandez, A.M., Peart, J.R., Malcuit, I. & Baulcombe, D.C. Virus-induced gene silencing in plants. Methods 30, 296–303 (2003).
Waterhouse, P.M., Wang, M.-B. & Lough, T. Gene silencing as an adaptive defence against viruses. Nature 411, 834–842 (2001).
Ruiz, M.T., Voinnet, O. & Baulcombe, D.C. Initiation and maintenance of virus-induced gene silencing. Plant Cell 10, 937–946 (1998).
Baulcombe, D.C. Fast forward genetics based on virus-induced gene silencing. Curr. Opin. Plant Biol. 2, 109–113 (1999).
Burch-Smith, T.M., Anderson, J.C., Martin, G.B. & Dinesh-Kumar, S.P. Applications and advantages of virus-induced gene silencing for gene function studies in plants. Plant J. 39, 734–746 (2004).
Kumagai, M.H. et al. Cytoplasmic inhibition of carotenoid biosynthesis with virus-derived RNA. Proc. Natl. Acad. Sci. USA 92, 1679–1683 (1995).
Senthil-Kumar, M., Ajith, A., Uppalapati, S.R. & Mysore, K.S. Virus-induced gene silencing and its applications. CAB Rev. Perspect. Agri. Vet. Sci. Nutri. Nat. Res. 3, 1–18 (2008).
Lu, R. et al. High throughput virus-induced gene silencing implicates heat shock protein 90 in plant disease resistance. EMBO J. 22, 5690–5699 (2003).
Senthil-Kumar, M. & Mysore, K.S. New dimensions for VIGS in plant functional genomics. Trends Plant Sci. 16, 656–665 (2011).
Senthil-Kumar, M., Govind, G., Kang, L., Mysore, K. & Udayakumar, M. Functional characterization of Nicotiana benthamiana homologs of peanut water deficit-induced genes by virus-induced gene silencing. Planta 225, 523–539 (2007).
Rojas, C.M. et al. Glycolate oxidase plays a major role during nonhost resistance responses by modulating reactive oxygen species mediated signal transduction pathways. Plant Cell 24, 336–352 (2012).
Becker, A. & Lange, M. VIGS – genomics goes functional. Trends Plant Sci. 15, 1–4 (2010).
Liu, Y.L., Schiff, M., Marathe, R. & Dinesh-Kumar, S.P. Tobacco Rar1, EDS1 and NPR1/NIM1-like genes are required for N-mediated resistance to tobacco mosaic virus. Plant J. 30, 415–429 (2002).
Ratcliff, F., Martin-Hernandez, A.M. & Baulcombe, D.C. Tobacco rattle virus as a vector for analysis of gene function by silencing. Plant J. 25, 237–245 (2001).
Pozo, O., Pedley, K.F. & Martin, G.B. MAPKKKα is a positive regulator of cell death associated with both plant immunity and disease. EMBO J. 23, 3072–3082 (2004).
Vaistij, F.E. & Jones, L. Compromised virus-induced gene silencing in RDR6-deficient plants. Plant Physiol. 149, 1399–1407 (2009).
MacFarlane, S.A. Molecular biology of the tobraviruses. J. Gen. Virol. 80, 2799–2807 (1999).
Ziegler-Graff, V., Guilford, P.J. & Baulcombe, D.C. Tobacco rattle virus RNA-1 29K gene product potentiates viral movement and also affects symptom induction in tobacco. Virology 182, 145–155 (1991).
Visser, P.B. & Bol, J.F. Nonstructural proteins of tobacco rattle virus which have a role in nematode-transmission: expression pattern and interaction with viral coat protein. J. Gen. Virol. 80, 3273–3280 (1999).
Liu, Y., Schiff, M. & Dinesh-Kumar, S.P. Virus-induced gene silencing in tomato. Plant J. 31, 777–786 (2002).
Dong, Y., Burch-Smith, T.M., Liu, Y.L., Mamillapalli, P. & Dinesh-Kumar, S.P. A ligation-independent cloning TRV vector for high-throughput virus-induced gene silencing identifies roles for NbMADS4-1 and -2 in floral development. Plant Physiol. 145, 1161–1170 (2007).
Ryu, C.-M., Anand, A., Kang, L. & Mysore, K.S. Agrodrench: a novel and effective agroinoculation method for virus-induced gene silencing in roots and diverse Solanaceous species. Plant J. 40, 322–331 (2004).
Senthil-Kumar, M. et al. A systematic study to determine the extent of gene silencing in Nicotiana benthamiana and other Solanaceae species when heterologous gene sequences are used for virus-induced gene silencing. New Phytol. 176, 782–791 (2007).
Senthil-Kumar, M., Ramegowda, H.V., Hema, R., Mysore, K.S. & Udayakumar, M. Virus-induced gene silencing and its application in characterizing genes involved in water-deficit-stress tolerance. J. Plant Physiol. 165, 1404–1421 (2008).
Senthil-Kumar, M. & Mysore, K.S. Virus-induced gene silencing can persist for more than 2 years and also be transmitted to progeny seedlings in Nicotiana benthamiana and tomato. Plant Biotechnol. J. 9, 797–806 (2011).
Liu, E. & Page, J. Optimized cDNA libraries for virus-induced gene silencing (VIGS) using tobacco rattle virus. Plant Methods 4, 5 (2008).
Anand, A. et al. Identification and characterization of plant genes involved in Agrobacterium-mediated plant transformation by virus-induced gene silencing. Mol. Plant Microbe Interact. 20, 41–52 (2007).
Wang, K., Senthil-Kumar, M., Ryu, C.-M., Kang, L. & Mysore, K.S. Phytosterols play a key role in plant innate immunity against bacterial pathogens by regulating nutrient efflux into the apoplast. Plant Physiol. 158, 1789–1802 (2012).
Wangdi, T. et al. A virus-induced gene silencing screen identifies a role for thylakoid formation1 in Pseudomonas syringae pv tomato symptom development in tomato and Arabidopsis. Plant Physiol. 152, 281–292 (2010).
Wolpert, T. & Gilbert, B.M. Characterization of the LOV1-mediated, victorin-induced cell death response with virus-induced gene silencing. Mol. Plant Microbe Interact. 26, 903–917 (2013).
Deng, X. et al. Modification of tobacco rattle virus RNA1 to serve as a VIGS vector reveals that the 29K movement protein is an RNA silencing suppressor of the virus. Mol. Plant Microbe Interact. 26, 503–514 (2013).
Ramegowda, V., Senthil-Kumar, M., Udayakumar, M. & Mysore, K. A high-throughput virus-induced gene silencing protocol identifies genes involved in multi-stress tolerance. BMC Plant Biol. 13, 193 (2013).
Fernandez-Moreno, J.-P., Orzaez, D. & Granell, A. VIGS: a tool to study fruit development in Solanum lycopersicum. in Virus-induced Gene Silencing Vol. 975 (ed. Becker, A), 183–196 (Humana Press, 2013).
Fu, D.-Q., Zhu, B.-Z., Zhu, H.-L., Jiang, W.-B. & Luo, Y.-B. Virus-induced gene silencing in tomato fruit. Plant J. 43, 299–308 (2005).
Yan, H.-x. et al. Sprout vacuum-infiltration: a simple and efficient agroinoculation method for virus-induced gene silencing in diverse solanaceous species. Plant Cell Rep. 31, 1713–1722 (2012).
Senthil-Kumar, M., Lee, H.K. & Mysore, K.S . VIGS-mediated forward genetics screening for identification of genes involved in nonhost resistance. J. Vis. Exp. e51033 (2013).
Vaghchhipawala, Z., Rojas, C., Senthil-Kumar, M. & Mysore, K. Agroinoculation and agroinfiltration: simple tools for complex gene function analyses. in Plant Reverse Genetics Vol. 678. (ed. Pereira, A.) 65–76 (Humana Press, 2011).
Hollings, M. Chenopodium amaranticolor as a test plant for plant viruses. Plant Pathol. 5, 57–60 (1956).
Xu, P., Zhang, Y., Kang, L., Roossinck, M.J. & Mysore, K.S. Computational estimation and experimental verification of off-target silencing during post-transcriptional gene silencing in plants. Plant Physiol. 142, 429–440 (2006).
Hartl, M., Merker, H., Schmidt, D.D. & Baldwin, I.T. Optimized virus-induced gene silencing in Solanum nigrum reveals the defensive function of leucine aminopeptidase against herbivores and the shortcomings of empty vector controls. New Phytol. 179, 356–365 (2008).
Wu, C., Jia, L. & Goggin, F. The reliability of virus-induced gene silencing experiments using tobacco rattle virus in tomato is influenced by the size of the vector control. Mol. Plant Pathol. 12, 299–305 (2011).
Qin, G. et al. Disruption of phytoene desaturase gene results in albino and dwarf phenotypes in Arabidopsis by impairing chlorophyll, carotenoid, and gibberellin biosynthesis. Cell Res. 17, 471–482 (2007).
Hiriart, J.-B., Lehto, K., Tyystjärvi, E., Junttila, T. & Aro, E.-M. Suppression of a key gene involved in chlorophyll biosynthesis by means of virus-inducing gene silencing. Plant Mol. Biol. 50, 213–224 (2002).
Dinesh-Kumar, S.P., Anandalakshmi, R., Marathe, R., Schiff, M. & Liu, Y. Virus-induced gene silencing. in Plant Functional Genomics Vol. 236. (ed. Grotewold, E.) 287–293 (Humana Press, 2003).
Velásquez, A.C., Chakravarthy, S. & Martin, G.B. Virus-induced gene silencing (VIGS) in Nicotiana benthamiana and tomato. J. Vis. Exp. e1292 (2009).
Hayward, A., Padmanabhan, M. & Dinesh-Kumar, S.P. Virus-induced gene silencing in Nicotiana benthamiana and other plant species. in Plant Reverse Genetics Vol. 678. (ed. Pereira, A.) 55–63 (Humana Press, 2011).
Brigneti, G. et al. Viral pathogenicity determinants are suppressors of transgene silencing in Nicotiana benthamiana. EMBO J. 17, 6739–6746 (1998).
Bombarely, A. et al. A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Mol. Plant Microbe Interact. 25, 1523–1530 (2012).
McCormac, A.C., Elliott, M.C. & Chen, D.F. A simple method for the production of highly competent cells of Agrobacterium for transformation via electroporation. Mol. Biotechnol. 9, 155–159 (1998).
Udvardi, M.K., Czechowski, T. & Scheible, W.-R. Eleven golden rules of quantitative RT-PCR. Plant Cell 20, 1736–1737 (2008).
Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Zhang, Y. et al. De novo foliar transcriptome of Chenopodium amaranticolor and analysis of its gene expression during virus-induced hypersensitive response. PLoS ONE 7, e45953 (2012).
Cooper, B. Collateral gene expression changes induced by distinct plant viruses during the hypersensitive resistance reaction in Chenopodium amaranticolor. Plant J. 26, 339–349 (2001).
Acknowledgements
This project was funded by The Samuel Roberts Noble Foundation. We thank J. Gallaway for excellent plant care. TRV constructs were obtained from S.P. Dinesh-Kumar, University of California Davis. We thank B. Stearns and S. McNeill for making and editing the video. We thank K. Brown for artwork in the figures and J. Kelley for editing the manuscript.
Author information
Authors and Affiliations
Contributions
M.S.-K. and K.S.M. designed the experiments; M.S.-K. performed the experiments; and M.S.-K. and K.S.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
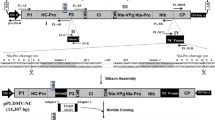
Supplementary Figure 1 Details of TRV1 and TRV2 constructs.
Tobacco rattle virus (TRV) is a bipartite, positive-sense RNA virus. TRV1 (i.e., RNA1) encodes 134 and 194 kDa replicase proteins from the genomic RNA, 29 kDa movement protein, and 16 kDa cysteine-rich protein from subgenomic RNAs. TRV1 can replicate and move systemically without TRV2 (i.e., RNA2). TRV2 encodes coat protein from the genomic and two nonstructural proteins from subgenomic RNAs (A). To develop the TRV-based VIGS vector, a cDNA clone of TRV1 or TRV2 (Ppk20 strain) was placed between a duplicated CaMV 35S promoter (2X35S) and nopaline synthase terminator (NOSt) in an Agrobacterium T-DNA vector. In the TRV2 cDNA construct, the nonessential structural genes were replaced with MCS for cloning the target gene sequences. Further, a self-cleaving ribozyme site was included at the 3' end of TRV2. The TRV1 (AF406990)-based viral vector, pTRV1 (linear plasmid 6.791 kb), is used along with pTRV2 for silencing. The pTRV2 has three variants. The first one is suitable for conventional cloning (AF406991, linear plasmid 9663 nucleotide). The second variant is a gateway-cloning-compatible vector. The third variant is suitable for ligation-independent cloning. Rz, self-cleaving ribozyme; MCS, multiple cloning site; CP, coat protein; MP, movement protein. These vectors were created by Dr. S.P. Dinesh-Kumar, UC Davis, USA. Conventional cloning-compatible (YL156) and ligation-independent-compatible (YY13TRV2) vector plasmid DNA can be obtained from ABRC, Stock No. CD3-1040 vector name YL156. Ligation-independent-compatible TRV2 vector plasmid DNA is ABRC Stock No. CD3-1042, vector name YY13.
Supplementary Figure 2 Applications of VIGS to study gene function in various plant processes.
Some of the possible studies that can be performed on the silenced plants are given in this figure. VIGS has been used to dissect developmental processes like root and leaf development, meristem differentiation, reproductive organ development, the mechanism of symbiotic interaction with microbes, PAMP or effector-induced defense, nonhost disease resistance, nematode resistance, Agrobacterium-mediated T-DNA transfer, and abiotic stress tolerance in plants.
Supplementary information
Supplementary Figure 1
Details of TRV1 and TRV2 constructs. (PDF 390 kb)
Supplementary Figure 2
Applications of VIGS to study gene function in various plant processes. (PDF 434 kb)
Supplementary Table 1
Plant species having publicly available and popularly used mutant resources conducive for cloning the gene of interest. (PDF 108 kb)
Supplementary Table 2
Currently available popular VIGS vectors and their target plant species. (PDF 216 kb)
Supplementary Table 3
Plant species suitable for TRV-VIGS of target genes. (PDF 175 kb)
Supplementary Table 4
Different methods of TRV inoculation for achieving VIGS. (PDF 125 kb)
Supplementary Table 5
Details of some useful primers for performing TRV-VIGS in N. benthamiana. (PDF 178 kb)
Rights and permissions
About this article
Cite this article
Senthil-Kumar, M., Mysore, K. Tobacco rattle virus–based virus-induced gene silencing in Nicotiana benthamiana. Nat Protoc 9, 1549–1562 (2014). https://doi.org/10.1038/nprot.2014.092
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.092
This article is cited by
-
Cucumber mosaic virus-induced gene and microRNA silencing in water dropwort (Oenanthe javanica (Blume) DC)
Plant Methods (2024)
-
Silencing of a Nicotiana benthamiana ascorbate oxidase gene reveals its involvement in resistance against cucumber mosaic virus
Planta (2024)
-
Establishment of virus-induced gene silencing (VIGS) system in Luffa acutangula using Phytoene desaturase (PDS) and tendril synthesis related gene (TEN)
Plant Methods (2023)
-
Divergent sequences of tetraspanins enable plants to specifically recognize microbe-derived extracellular vesicles
Nature Communications (2023)
-
Principles and practice of virus induced gene silencing for functional genomics in plants
Virus Genes (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.