Abstract
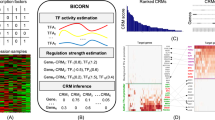
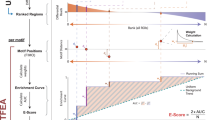
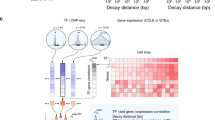
The combination of ChIP-seq and transcriptome analysis is a compelling approach to unravel the regulation of gene expression. Several recently published methods combine transcription factor (TF) binding and gene expression for target prediction, but few of them provide an efficient software package for the community. Binding and expression target analysis (BETA) is a software package that integrates ChIP-seq of TFs or chromatin regulators with differential gene expression data to infer direct target genes. BETA has three functions: (i) to predict whether the factor has activating or repressive function; (ii) to infer the factor's target genes; and (iii) to identify the motif of the factor and its collaborators, which might modulate the factor's activating or repressive function. Here we describe the implementation and features of BETA to demonstrate its application to several data sets. BETA requires ∼1 GB of RAM, and the procedure takes 20 min to complete. BETA is available open source at http://cistrome.org/BETA/.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Jiang, C., Xuan, Z., Zhao, F. & Zhang, M.Q. TRED: a transcriptional regulatory element database, new entries and other development. Nucleic Acids Res. 35, D137–D140 (2007).
Buck, M.J. & Lieb, J.D. A chromatin-mediated mechanism for specification of conditional transcription factor targets. Nat. Genet. 38, 1446–1451 (2006).
Palii, C.G. et al. Differential genomic targeting of the transcription factor TAL1 in alternate haematopoietic lineages. EMBO J. 30, 494–509 (2010).
Tang, Q. et al. A comprehensive view of nuclear receptor cancer cistromes. Cancer Res. 71, 6940–6947 (2011).
Breitling, R., Armengaud, P., Amtmann, A. & Herzyk, P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 573, 83–92 (2004).
Sherman, B.T. et al. DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35, W169–W175 (2007).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X.S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Ma, W. & Wong, W.H. The analysis of ChIP-seq data. Methods Enzymol. 497, 51–73 (2011).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP-seq data. Bioinformatics 25, 1952–1958 (2009).
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Rougemont, J. & Naef, F. Computational analysis of protein-DNA interactions from ChIP-seq data. Methods Mol. Biol. 786, 263–273 (2012).
Cheng, C., Min, R. & Gerstein, M. TIP: a probabilistic method for identifying transcription factor target genes from ChIP-seq binding profiles. Bioinformatics 27, 3221–3227 (2011).
Qian, J., Lin, J., Luscombe, N.M., Yu, H. & Gerstein, M. Prediction of regulatory networks: genome-wide identification of transcription factor targets from gene expression data. Bioinformatics 19, 1917–1926 (2003).
Honkela, A. et al. Model-based method for transcription factor target identification with limited data. Proc. Natl. Acad. Sci. 107, 7793–7798 (2010).
Redestig, H., Weicht, D., Selbig, J. & Hannah, M.A. Transcription factor target prediction using multiple short expression time series from Arabidopsis thaliana. BMC Bioinformatics 8, 454 (2007).
Qin, J., Li, M.J., Wang, P., Zhang, M.Q. & Wang, J. ChIP-Array: combinatory analysis of ChIP-seq/chip and microarray gene expression data to discover direct/indirect targets of a transcription factor. Nucleic Acids Res. 39, W430–W436 (2011).
Maienschein-Cline, M., Zhou, J., White, K.P., Sciammas, R. & Dinner, A.R. Discovering transcription factor regulatory targets using gene expression and binding data. Bioinformatics 28, 206–213 (2012).
Smyth, G.K. Limma: linear models for microarray data. In Bioinformatics and Computational Biology Solutions using R and Bioconductor (eds. Gentleman, R., Carey, V.J., Huber, W., Irizarry, R.A., Dudoit, S.) 397–420 (Springer, 2005).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Johnson, W.E. et al. Model-based analysis of tiling-arrays for ChIP-chip. Proc. Natl. Acad. Sci. USA 103, 12457–12462 (2006).
Wang, Q. et al. A hierarchical network of transcription factors governs androgen receptor-dependent prostate cancer growth. Mol. Cell 27, 380–392 (2007).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Chan, C.S. & Song, J.S. CCCTC-binding factor confines the distal action of estrogen receptor. Cancer Res. 68, 9041–9049 (2008).
Sun, Z., Pan, J. & Balk, S.P. Androgen receptor-associated protein complex binds upstream of the androgen-responsive elements in the promoters of human prostate-specific antigen and kallikrein 2 genes. Nucleic Acids Res. 25, 3318–3325 (1997).
Steketee, K., Ziel-van der Made, A.C., van der Korput, H.A., Houtsmuller, A.B. & Trapman, J. A bioinformatics-based functional analysis shows that the specifically androgen-regulated gene SARG contains an active direct repeat androgen response element in the first intron. J. Mol. Endocrinol. 33, 477–491 (2004).
Cai, C., Wang, H., Xu, Y., Chen, S. & Balk, S.P. Reactivation of androgen receptor-regulated TMPRSS2:ERG gene expression in castration-resistant prostate cancer. Cancer Res. 69, 6027–6032 (2009).
Thorvaldsdottir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Korhonen, J., Martinmäki, P., Pizzi, C., Rastas, P. & Ukkonen, E. MOODS: fast search for position weight matrix matches in DNA sequences. Bioinformatics 25, 3181–3182 (2009).
Liu, T. et al. Cistrome: an integrative platform for transcriptional regulation studies. Genome Biol. 12, R83 (2011).
Habib, N., Kaplan, T., Margalit, H. & Friedman, N. A novel Bayesian DNA motif comparison method for clustering and retrieval. PLoS Comput. Biol. 4, e1000010 (2008).
Fulton, D.L. et al. TFCat: the curated catalog of mouse and human transcription factors. Genome Biol. 10, R29 (2009).
Da Wei Huang, B.T.S., Stephens, R., Baseler, M.W., Lane, H.C. & Lempicki, R.A. DAVID gene ID conversion tool. Bioinformation 2, 428 (2008).
Fujita, P.A. et al. The UCSC genome browser database: update 2011. Nucleic Acids Res. 39, D876–D882 (2011).
Dai, M. et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res. 33, e175–e175 (2005).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Hu, M., Yu, J., Taylor, J.M., Chinnaiyan, A.M. & Qin, Z.S. On the detection and refinement of transcription factor binding sites using ChIP-seq data. Nucleic Acids Research 38, 2154–2167 (2010).
Carroll, J.S. et al. Genome-wide analysis of estrogen receptor binding sites. Nat. Genet. 38, 1289–1297 (2006).
Acknowledgements
This project was supported by the National Basic Research (973) Program of China (2010CB944904), the National Natural Science Foundation of China (31329003) and the US National Institutes of Health (HG4069 and U41 HG007000).
Author information
Authors and Affiliations
Contributions
S.W., C.A.M., Q.T. and X.S.L. designed the method; S.W., H.S., J.M., C.W., C.Z., J.W. and X.S.L. implemented the algorithm; S.W. performed the data analysis; and S.W. and X.S.L. wrote the initial manuscript. All authors contributed to the discussion and writing of the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Wang, S., Sun, H., Ma, J. et al. Target analysis by integration of transcriptome and ChIP-seq data with BETA. Nat Protoc 8, 2502–2515 (2013). https://doi.org/10.1038/nprot.2013.150
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.150
This article is cited by
-
CBX3 antagonizes IFNγ/STAT1/PD-L1 axis to modulate colon inflammation and CRC chemosensitivity
EMBO Molecular Medicine (2024)
-
IL-33 priming and antigenic stimulation synergistically promote the transcription of proinflammatory cytokine and chemokine genes in human skin mast cells
BMC Genomics (2023)
-
Chromatin accessibility uncovers KRAS-driven FOSL2 promoting pancreatic ductal adenocarcinoma progression through up-regulation of CCL28
British Journal of Cancer (2023)
-
The EGR3 regulome of infant KMT2A-r acute lymphoblastic leukemia identifies differential expression of B-lineage genes predictive for outcome
Leukemia (2023)
-
Prolyl endopeptidase remodels macrophage function as a novel transcriptional coregulator and inhibits fibrosis
Experimental & Molecular Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.