Abstract
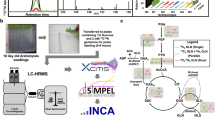
Metabolism has a decisive role in many fundamental biological processes, including organism development and tissue homeostasis. Here we describe a protocol for fast and reliable 13C-isotope-based in vivo metabolic profiling. This protocol covers the loading of isotope precursor; extraction, preparation and quantification of the labeled lipid metabolites (e.g., the prenyl lipid CoQ10) by the means of HPLC-MS; and its analysis in zebrafish embryos. This protocol can be applied to different types of experimental settings, including tissue-specific metabolic analyses or dynamic metabolic changes that occur during vertebrate embryogenesis. The protocol takes 5–7 d to complete, requiring minimal equipment and analytical expertise, and it represents a unique alternative to the existing ex vivo (e.g., cell lines) isotope-based metabolic methods. This procedure represents a valuable approach for researchers interested in studying the effect of gene manipulation on lipid metabolism in zebrafish and in understanding the genetic conditions that result in metabolism dysfunction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Crown, S.B. & Antoniewicz, M.R. Parallel labeling experiments and metabolic flux analysis: past, present and future methodologies. Metab. Eng. 16C, 21–32 (2012).
Klein, S. & Heinzle, E. Isotope labeling experiments in metabolomics and fluxomics. Wiley Interdiscip. Rev. Syst. Biol. Med. 4, 261–272 (2012).
Kalen, A., Appelkvist, E.L., Chojnacki, T. & Dallner, G. Nonaprenyl-4-hydroxybenzoate transferase, an enzyme involved in ubiquinone biosynthesis, in the endoplasmic reticulum-Golgi system of rat liver. J. Biol. Chem. 265, 1158–1164 (1990).
Bequette, B.J., Sunny, N.E., El-Kadi, S.W. & Owens, S.L. Application of stable isotopes and mass isotopomer distribution analysis to the study of intermediary metabolism of nutrients. J. Anim. Sci. 84 (suppl.), E50–E59 (2006).
Mugoni, V. et al. Ubiad1 is an antioxidant enzyme that regulates eNOS activity by CoQ10 synthesis. Cell 152, 504–518 (2013).
Fahy, E. et al. Update of the LIPID MAPS comprehensive classification system for lipids. J. Lipid Res. 50 (suppl.), S9–S14 (2009).
Fahy, E. et al. A comprehensive classification system for lipids. J. Lipid Res. 46, 839–861 (2005).
Raamsdonk, L.M. et al. A functional genomics strategy that uses metabolome data to reveal the phenotype of silent mutations. Nat. Biotechnol. 19, 45–50 (2001).
Zamboni, N. 13C metabolic flux analysis in complex systems. Curr. Opin. Biotechnol. 22, 103–108 (2011).
Trethewey, R.N. Gene discovery via metabolic profiling. Curr. Opin. Biotechnol. 12, 135–138 (2001).
Lv, H. Mass spectrometry-based metabolomics towards understanding of gene functions with a diversity of biological contexts. Mass Spectrom. Rev. 32, 118–128 (2013).
Fiehn, O. Metabolomics--the link between genotypes and phenotypes. Plant Mol. Biol. 48, 155–171 (2002).
Adamski, J. & Suhre, K. Metabolomics platforms for genome wide association studies-linking the genome to the metabolome. Curr. Opin. Biotechnol. 24, 39–47 (2013).
Crane, F.L. Biochemical functions of coenzyme Q10. J. Am. Coll. Nutr. 20, 591–598 (2001).
Bentinger, M., Dallner, G., Chojnacki, T. & Swiezewska, E. Distribution and breakdown of labeled coenzyme Q10 in rat. Free Radic. Biol. Med. 34, 563–575 (2003).
Huang, S.M., Xu, F., Lam, S.H., Gong, Z. & Ong, C.N. Metabolomics of developing zebrafish embryos using gas chromatography- and liquid chromatography-mass spectrometry. Mol. Biosyst. 9, 1372–1380 (2013).
Nath, A.K. et al. Chemical and metabolomic screens identify novel biomarkers and antidotes for cyanide exposure. FASEB J. 27, 1928–1938 (2013).
Hayashi, S. et al. A novel application of metabolomics in vertebrate development. Biochem. Biophys. Res. Commun. 386, 268–272 (2009).
Ong, E.S., Chor, C.F., Zou, L. & Ong, C.N. A multi-analytical approach for metabolomic profiling of zebrafish (Danio rerio) livers. Mol. Biosyst. 5, 288–298 (2009).
Mootha, V.K. & Hirschhorn, J.N. Inborn variation in metabolism. Nat. Genet. 42, 97–98 (2010).
Weiss, R.H. & Kim, K. Metabolomics in the study of kidney diseases. Nat. Rev. Nephrol. 8, 22–33 (2012).
Jin, S.W. et al. A transgene-assisted genetic screen identifies essential regulators of vascular development in vertebrate embryos. Dev. Biol. 307, 29–42 (2007).
Bentinger, M. et al. Polyisoprenoid epoxides stimulate the biosynthesis of coenzyme Q and inhibit cholesterol synthesis. J. Biol. Chem. 283, 14645–14653 (2008).
Kilkenny, C., Browne, W.J., Cuthill, I.C., Emerson, M. & Altman, D.G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
Rosen, J.N., Sweeney, M.F. & Mably, J.D. Microinjection of zebrafish embryos to analyze gene function. J. Vis. Exp. 2009, 1115 (2009).
Singleman, C. & Holtzman, N.G. Heart dissection in larval, juvenile and adult zebrafish, Danio rerio. J. Vis. Exp. 2011, 3165 (2011).
Gupta, T. & Mullins, M.C. Dissection of organs from the adult zebrafish. J. Vis. Exp. 2011, 2839 (2010).
Turunen, M., Olsson, J. & Dallner, G. Metabolism and function of coenzyme Q. Biochim. Biophys. Acta 1660, 171–199 (2004).
Link, V., Shevchenko, A. & Heisenberg, C.P. Proteomics of early zebrafish embryos. BMC Dev. Biol. 6, 1 (2006).
United Kingdom Co-ordinating Committee on Cancer Research (UKCCCR) guidelines for the welfare of animals in experimental neoplasia (second edition). Br. J. Cancer 77, 1–10 (1998).
Acknowledgements
M.M.S.'s laboratory was supported by research grants from the Marie Curie Action IRG 247852, Telethon GGP10195 and AIRC MFAG-8911. We thank E.J. Corcoran for editorial assistance.
Author information
Authors and Affiliations
Contributions
V.M. and M.M.S. designed the study. V.M., C.M. and M.M.S. performed the experiments, analyzed the data and discussed this study. V.M., C.M. and M.M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Mugoni, V., Medana, C. & Santoro, M. 13C-isotope-based protocol for prenyl lipid metabolic analysis in zebrafish embryos. Nat Protoc 8, 2337–2347 (2013). https://doi.org/10.1038/nprot.2013.139
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.139
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.