Abstract
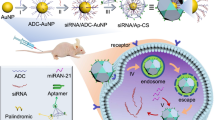
RNA nanotechnology is a term that refers to the design, fabrication and use of nanoparticles that are mainly composed of RNAs via bottom-up self-assembly. The packaging RNA (pRNA) of the bacteriophage phi29 DNA packaging motor has been developed into a nanodelivery platform. This protocol describes the synthesis, assembly and functionalization of pRNA nanoparticles on the basis of three 'toolkits' derived from pRNA structural features: interlocking loops for hand-in-hand interactions, palindrome sequences for foot-to-foot interactions and an RNA three-way junction for branch extension. siRNAs, ribozymes, aptamers, chemical ligands, fluorophores and other functionalities can also be fused to the pRNA before the assembly of the nanoparticles, so as to ensure the production of homogeneous nanoparticles and the retention of appropriate folding and function of the incorporated modules. The resulting self-assembled multivalent pRNA nanoparticles are thermodynamically and chemically stable, and they remain intact at ultralow concentrations. Gene-silencing effects are progressively enhanced with increasing numbers of siRNAs in each pRNA nanoparticle. Systemic injection of the pRNA nanoparticles into xenograft-bearing mice has revealed strong binding to tumors without accumulation in vital organs or tissues. The pRNA-based nanodelivery scaffold paves a new way for nanotechnological application of pRNA-based nanoparticles for disease detection and treatment. The time required for completing one round of this protocol is 3–4 weeks when including in vitro functional assays, or 2–3 months when including in vivo studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Guo, P. The emerging field of RNA nanotechnology. Nat. Nanotechnol. 5, 833–842 (2010).
Shu, D., Huang, L., Hoeprich, S. & Guo, P. Construction of phi29 DNA-packaging RNA (pRNA) monomers, dimers and trimers with variable sizes and shapes as potential parts for nano-devices. J. Nanosci. Nanotechnol. 3, 295–302 (2003).
Shu, D., Moll, W.D., Deng, Z., Mao, C. & Guo, P. Bottom-up assembly of RNA arrays and superstructures as potential parts in nanotechnology. Nano Lett. 4, 1717–1723 (2004).
Hansma, H.G., Oroudjev, E., Baudrey, S. & Jaeger, L. TectoRNA and 'kissing-loop' RNA: atomic force microscopy of self-assembling RNA structures. J. Microsc. 212, 273–279 (2003).
Guo, P., Zhang, C., Chen, C., Trottier, M. & Garver, K. Inter-RNA interaction of phage phi29 pRNA to form a hexameric complex for viral DNA transportation. Mol. Cell 2, 149–155 (1998).
Zhang, F. et al. Function of hexameric RNA in packaging of bacteriophage phi29 DNA in vitro. Mol. Cell 2, 141–147 (1998).
Jaeger, L. & Leontis, N.B. Tecto-RNA: one dimensional self-assembly through tertiary interactions. Angew. Chem. Int. Ed. Engl. 39, 2521–2524 (2000).
Jaeger, L., Westhof, E. & Leontis, N.B. TectoRNA: modular assembly units for the construction of RNA nano-objects. Nucleic Acids Res. 29, 455–463 (2001).
Chworos, A. et al. Building programmable jigsaw puzzles with RNA. Science 306, 2068–2072 (2004).
Guo, P. RNA nanotechnology: engineering, assembly and applications in detection, gene delivery and therapy. J. Nanosci. Nanotechnol. 5 (12): 1964–1982 (2005).
Jaeger, L. & Chworos, A. The architectonics of programmable RNA and DNA nanostructures. Curr. Opin. Struct. Biol. 16, 531–543 (2006).
Abraham, M., Dror, O., Nussinov, R. & Wolfson, H.J. Analysis and classification of RNA tertiary structures. RNA 14, 2274–2289 (2008).
Bindewald, E., Hayes, R., Yingling, Y.G., Kasprzak, W. & Shapiro, B.A. RNAJunction: a database of RNA junctions and kissing loops for three-dimensional structural analysis and nanodesign. Nucleic Acids Res. 36, D392–D397 (2008).
Schroeder, K.T., McPhee, S.A., Ouellet, J. & Lilley, D.M. A structural database for k-turn motifs in RNA. RNA 16, 1463–1468 (2010).
Westhof, E., Masquida, B. & Jaeger, L. RNA tectonics: towards RNA design. Fold. Des. 1, R78–R88 (1996).
Lescoute, A. & Westhof, E. Topology of three-way junctions in folded RNAs. RNA 12, 83–93 (2006).
Leontis, N.B. & Westhof, E. Analysis of RNA motifs. Curr. Opin. Struct. Biol. 13, 300–308 (2003).
Jossinet, F., Ludwig, T.E. & Westhof, E. RNA structure: bioinformatic analysis. Curr. Opin. Microbiol. 10, 279–285 (2007).
Leontis, N.B., Lescoute, A. & Westhof, E. The building blocks and motifs of RNA architecture. Curr. Opin. Struct. Biol. 16, 279–287 (2006).
Leontis, N.B. & Westhof, E. The annotation of RNA motifs. Comp. Funct. Genomics 3, 518–524 (2002).
Leontis, N.B. & Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001).
Cadd, T.L. & Patterson, J.L. Synthesis of viruslike particles by expression of the putative capsid protein of Leishmania RNA virus in a recombinant baculovirus expression system. J. Virol. 68, 358–365 (1994).
McKinney, S.A., Declais, A.C., Lilley, D.M.J. & Ha, T. Structural dynamics of individual Holliday junctions. Nat. Struct. Biol. 10, 93–97 (2003).
Lilley, D.M. Structure, folding and catalysis of the small nucleolytic ribozymes. Curr. Opin. Struct. Biol. 9, 330–338 (1999).
Delebecque, C.J., Lindner, A.B., Silver, P.A. & Aldaye, F.A. Organization of intracellular reactions with rationally designed RNA assemblies. Science 333, 470–474 (2011).
Afonin, K.A. et al. Design and self-assembly of siRNA-functionalized RNA nanoparticles for use in automated nanomedicine. Nat. Protoc. 6, 2022–2034 (2011).
Huttenhofer, A., Schattner, P. & Polacek, N. Non-coding RNAs: hope or hype? Trends Genet. 21, 289–297 (2005).
Strobel, S.A. & Cochrane, J.C. RNA catalysis: ribozymes, ribosomes, and riboswitches. Curr. Opin. Chem. Biol. 11, 636–643 (2007).
Fabian, M.R., Sonenberg, N. & Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 79, 351–379 (2010).
Taft, R.J., Pang, K.C., Mercer, T.R., Dinger, M. & Mattick, J.S. Non-coding RNAs: regulators of disease. J. Pathol. 220, 126–139 (2010).
Lee, C.Y.F., Rennie, P.S. & Jia, W.W.G. MicroRNA regulation of oncolytic herpes simplex virus-1 for selective killing of prostate cancer cells. Clin. Cancer Res. 15, 5126–5135 (2009).
Zhang, C. Novel functions for small RNA molecules. Curr. Opin. Mol. Ther. 11, 641–651 (2009).
Sudarsan, N. et al. Riboswitches in eubacteria sense the second messenger cyclic di-GMP. Science 321, 411–413 (2008).
Henkin, T.M. Riboswitch RNAs: using RNA to sense cellular metabolism. Genes Dev. 22, 3383–3390 (2008).
Ogawa, A. & Maeda, M. An artificial aptazyme-based riboswitch and its cascading system in E. coli. Chembiochem 9, 206–209 (2008).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Ghildiyal, M. & Zamore, P.D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 10, 94–108 (2009).
Guo, P. et al. Engineering RNA for targeted siRNA delivery and medical application. Adv. Drug Deliv. Rev. 62, 650–666 (2010).
Li, H., Li, W.X. & Ding, S.W. Induction and suppression of RNA silencing by an animal virus. Science 296, 1319–1321 (2002).
Brummelkamp, T.R., Bernards, R. & Agami, R. A system for stable expression of short interfering RNAs in mammalian cells. Science 296, 550–553 (2002).
Jacque, J.M., Triques, K. & Stevenson, M. Modulation of HIV-1 replication by RNA interference. Nature 418, 435–438 (2002).
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002).
Carmichael, G.G. Medicine: silencing viruses with RNA. Nature 418, 379–380 (2002).
Cech, T.R., Zaug, A.J. & Grabowski, P.J. In vitro splicing of the ribosomal RNA precursor of Tetrahymena: involvement of a guanosine nucleotide in the excision of the intervening sequence. Cell 27, 487–496 (1981).
Chowrira, B.M., Berzal-Herranz, A. & Burke, J.M. Novel guanosine requirement for catalysis by the hairpin ribozyme. Nature 354, 320–322 (1991).
Forster, A.C. & Symons, R.H. Self-cleavage of virusoid RNA is performed by the proposed 55-nucleotide active site. Cell 50, 9–16 (1987).
Sarver, N.A. et al. Ribozymes as potential anti-HIV-1 therapeutic agents. Science 24, 1222–1225 (1990).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Coleman, J., Hirashima, A., Inocuchi, Y., Green, P.J. & Inouye, M. A novel immune system against bacteriophage infection using complementary RNA (micRNA). Nature 315, 601–603 (1985).
Knecht, D.A. & Loomis, W.F. Antisense RNA inactivation of myosin heavy chain gene expression in Dictyostelium discoideum. Science 236, 1081–1086 (1987).
Duchaine, T.F. & Slack, F.J. RNA interference and micro-RNA-oriented therapy in cancer: rationales, promises, and challenges. Curr. Oncol. 16, 265–270 (2009).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
He, L. & Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Breaker, R.R. Complex riboswitches. Science 319, 1795–1797 (2008).
Tucker, B.J. & Breaker, R.R. Riboswitches as versatile gene control elements. Curr. Opin. Struct. Biol. 15, 342–348 (2005).
Bouvet, P. Determination of nucleic acid recognition sequences by SELEX. Methods Mol. Biol. 148, 603–610 (2001).
Ciesiolka, J., Gorski, J. & Yarus, M. Selection of an RNA domain that binds Zn2+. RNA 1, 538–550 (1995).
Clark, S. & Remcho, V. Aptamers as analytical reagents. Electrophoresis 23, 1335–1340 (2002).
Kraus, E., James, W. & Barclay, A.N. Cutting edge: novel RNA ligands able to bind CD4 antigen and inhibit CD4+ T lymphocyte function. J. Immunol. 160, 5209–5212 (1998).
Shu, D. & Guo, P. A viral RNA that binds ATP and contains an motif similar to an ATP-binding aptamer from SELEX. J. Biol. Chem. 278, 7119–7125 (2003).
Shapiro, B.A. Computational design strategies for RNA nanostructures. J. Biomol. Str. Dyn. 26, 820 (2009).
Afonin, K.A. et al. In vitro assembly of cubic RNA-based scaffolds designed in silico. Nat. Nanotechnol. 5, 676–682 (2010).
Yingling, Y.G. & Shapiro, B.A. Computational design of an RNA hexagonal nanoring and an RNA nanotube. Nano Lett. 7, 2328–2334 (2007).
Bindewald, E., Grunewald, C., Boyle, B., O'Connor, M. & Shapiro, B.A. Computational strategies for the automated design of RNA nanoscale structures from building blocks using NanoTiler. J. Molec. Graph. Model. 27, 299–308 (2008).
Grabow, W.W. et al. Self-assembling RNA nanorings based on RNAI/II inverse kissing complexes. Nano Lett. 11, 878–887 (2011).
Shapiro, B.A., Bindewald, E., Kasprzak, W. & Yingling, Y. Protocols for the in silico design of RNA nanostructures. Methods Mol. Biol. 474, 93–115 (2008).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Andronescu, M., Fejes, A.P., Hutter, F., Hoos, H.H. & Condon, A. A new algorithm for RNA secondary structure design. J. Mol. Biol. 336, 607–624 (2004).
Ding, Y., Chan, C.Y. & Lawrence, C.E. Sfold web server for statistical folding and rational design of nucleic acids. Nucleic Acids Res. 32, W135–W141 (2004).
Zadeh, J.N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Delebecque, C.J., Silver, P.A. & Lindner, A.B. Designing and using RNA scaffolds to assemble proteins in vivo. Nat. Protoc. 7, 1797–1807 (2012).
Guo, P., Erickson, S. & Anderson, D. A small viral RNA is required for in vitro packaging of bacteriophage phi29 DNA. Science 236, 690–694 (1987).
Lee, T.J. & Guo, P. Interaction of gp16 with pRNA and DNA for genome packaging by the motor of bacterial virus phi29. J. Mol. Biol. 356, 589–599 (2006).
Cairns, J., Overbaugh, J. & Miller, S. The origin of mutants. Nature 335, 142–145 (1988).
Reid, R.J.D., Bodley, J.W. & Anderson, D. Characterization of the prohead-pRNA interaction of bacteriophage phi29. J. Biol. Chem. 269, 5157–5162 (1994).
Garver, K. & Guo, P. Boundary of pRNA functional domains and minimum pRNA sequence requirement for specific connector binding and DNA packaging of phage phi29. RNA 3, 1068–1079 (1997).
Zhang, C.L., Lee, C.-S. & Guo, P. The proximate 5′ and 3′ ends of the 120-base viral RNA (pRNA) are crucial for the packaging of bacteriophage f29 DNA. Virology 201, 77–85 (1994).
Reid, R.J.D., Bodley, J.W. & Anderson, D. Identification of bacteriophage phi29 prohead RNA (pRNA) domains necessary for in vitro DNA-gp3 packaging. J. Biol. Chem. 269, 9084–9089 (1994).
Reid, R.J.D., Zhang, F., Benson, S. & Anderson, D. Probing the structure of bacteriophage phi29 prohead RNA with specific mutations. J. Biol. Chem. 269, 18656–18661 (1994).
Chen, C., Sheng, S., Shao, Z. & Guo, P. A dimer as a building block in assembling RNA: A hexamer that gears bacterial virus phi29 DNA-translocating machinery. J. Biol. Chem. 275, 17510–17516 (2000).
Chen, C., Zhang, C. & Guo, P. Sequence requirement for hand-in-hand interaction in formation of pRNA dimers and hexamers to gear phi29 DNA translocation motor. RNA 5, 805–818 (1999).
Guo, S., Tschammer, N., Mohammed, S. & Guo, P. Specific delivery of therapeutic RNAs to cancer cells via the dimerization mechanism of phi29 motor pRNA. Hum. Gene Ther. 16, 1097–1109 (2005).
Khaled, A., Guo, S., Li, F. & Guo, P. Controllable self-assembly of nanoparticles for specific delivery of multiple therapeutic molecules to cancer cells using RNA nanotechnology. Nano Lett. 5, 1797–1808 (2005).
Hoeprich, S. et al. Bacterial virus phi29 pRNA as a hammerhead ribozyme escort to destroy hepatitis B virus. Gene Ther. 10, 1258–1267 (2003).
Guo, S., Huang, F. & Guo, P. Construction of folate-conjugated pRNA of bacteriophage phi29 DNA packaging motor for delivery of chimeric siRNA to nasopharyngeal carcinoma cells. Gene Ther. 13, 814–820 (2006).
Liu, H. et al. Phi29 pRNA vector for efficient escort of hammerhead ribozyme targeting survivin in multiple cancer cells. Cancer Biol. Ther. 6, 697–704 (2007).
Li, L., Liu, J., Diao, Z., Guo, P. & Shen, G. Evaluation of specific delivery of chimeric phi29 pRNA/siRNA nanoparticles to multiple tumor cells. Molec. BioSyst. 5, 1361–1368 (2009).
Zhang, H.M. et al. Targeted delivery of anti-coxsackievirus siRNAs using ligand-conjugated packaging RNAs. Antiviral Res. 83, 307–316 (2009).
Tarapore, P., Shu, Y., Guo, P. & Ho, S.M. Application of Phi29 motor pRNA for targeted therapeutic delivery of siRNA silencing metallothionein-IIA and survivin in ovarian cancers. Molec. Ther. 19, 386–394 (2010).
Shu, D., Shu, Y., Haque, F., Abdelmawla, S. & Guo, P. Thermodynamically stable RNA three-way junctions for constructing multifuntional nanoparticles for delivery of therapeutics. Nat. Nanotechnol. 6, 658–667 (2011).
Haque, F. et al. Ultrastable synergistic tetravalent RNA nanoparticles for targeting to cancers. Nano Today 7, 245–257 (2012).
Shu, Y. et al. Fabrication of 14 different RNA nanoparticles for specific tumor targeting without accumulation in normal organs. RNA 19, 766–777 (2013).
Douglas, S.J., Davis, S.S. & Illum, L. Nanoparticles in drug delivery. Crit. Rev. Ther. Drug Carrier Syst. 3, 233–261 (1987).
Moon, M.H. & Giddings, J.C. Size distribution of liposomes by flow field-flow fractionation. J. Pharm. Biomed. Anal. 11, 911–920 (1993).
Singh, A. & Marchywka, S. Synthesis and characterization of head group modified 1,3 diacetylenic phospholipids. Abstr. Pap. Am. Chem. Soc. 198, 147 (1989).
Agarwal, K., Bali, A. & Gupta, C.M. Influence of the phospholipid structure on the stability of liposomes in serum. Biochim. Biophys. Acta 856, 36–40 (1986).
Gupta, C.M. & Bali, A. Carbamyl analogs of phosphatidylcholines. Synthesis, interaction with phospholipases and permeability behavior of their liposomes. Biochim. Biophys. Acta 663, 506–515 (1981).
Koziara, J.M., Lockman, P.R., Allen, D.D. & Mumper, R.J. The blood-brain barrier and brain drug delivery. J. Nanosci. Nanotechnol. 6, 2712–2735 (2006).
Lockman, P.R., Koziara, J.M., Mumper, R.J. & Allen, D.D. Nanoparticle surface charges alter blood-brain barrier integrity and permeability. J. Drug Target 12, 635–641 (2004).
Vasey, P.A. et al. Phase I clinical and pharmacokinetic study of PK1 [N-(2-hydroxypropyl)methacrylamide copolymer doxorubicin]: first member of a new class of chemotherapeutic agents-drug-polymer conjugates. Cancer Research Campaign phase I/II committee. Clin. Cancer Res. 5, 83–94 (1999).
Liu, C. & Zhang, N. Nanoparticles in gene therapy principles, prospects, and challenges. Prog. Mol. Biol. Transl. Sci. 104, 509–562 (2011).
Maham, A., Tang, Z., Wu, H., Wang, J. & Lin, Y. Protein-based nanomedicine platforms for drug delivery. Small 5, 1706–1721 (2009).
Ulery, B.D., Nair, L.S. & Laurencin, C.T. Biomedical applications of biodegradable polymers. J. Polym. Sci. B Polym. Phys. 49, 832–864 (2011).
Curcio, M. et al. Grafted thermo-responsive gelatin microspheres as delivery systems in triggered drug release. Eur. J. Pharm. Biopharm. 76, 48–55 (2010).
Curcio, M. et al. Negative thermo-responsive microspheres based on hydrolyzed gelatin as drug delivery device. AAPS PharmSciTech. 11, 652–662 (2010).
Rizi, K., Green, R.J., Khutoryanskaya, O., Donaldson, M. & Williams, A.C. Mechanisms of burst release from pH-responsive polymeric microparticles. J. Pharm. Pharmacol. 63, 1141–1155 (2011).
Manchester, M. & Singh, P. Virus-based nanoparticles (VNPs): platform technologies for diagnostic imaging. Adv. Drug Deliv. Rev. 58, 1505–1522 (2006).
Rae, C.S. et al. Systemic trafficking of plant virus nanoparticles in mice via the oral route. Virology 343, 224–235 (2005).
Huang, H.C., Barua, S., Sharma, G., Dey, S.K. & Rege, K. Inorganic nanoparticles for cancer imaging and therapy. J. Control Release 155, 344–357 (2011).
Shukla, R. et al. Biocompatibility of gold nanoparticles and their endocytotic fate inside the cellular compartment: a microscopic overview. Langmuir 21, 10644–10654 (2005).
Kallenbach, N., Ma, R. & Seeman, N. An immobile nucleic acid junction constructed from oligonucleotides. Nature 305, 829–831 (1983).
Rothemund, P.W.K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Chen, J.H. & Seeman, N.C. Synthesis from DNA of a molecule with the connectivity of a cube. Nature 350, 631–633 (1991).
Ke, Y., Liu, Y., Zhang, J. & Yan, H. A study of DNA tube formation mechanisms using 4-, 8-, and 12-helix DNA nanostructures. J. Am. Chem. Soc. 128, 4414–4421 (2006).
Kuzuya, A., Wang, R., Sha, R. & Seeman, N.C. Six-helix and eight-helix DNA nanotubes assembled from half-tubes. Nano. Lett. 7, 1757–1763 (2007).
Winfree, E., Liu, F., Wenzler, L.A. & Seeman, N.C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Gu, H., Chao, J., Xiao, S.J. & Seeman, N.C. Dynamic patterning programmed by DNA tiles captured on a DNA origami substrate. Nat. Nanotechnol. 4, 245–248 (2009).
Ke, Y. et al. Multilayer DNA origami packed on a square lattice. J. Am. Chem. Soc. 131, 15903–15908 (2009).
Mao, C., Sun, W., Shen, Z. & Seeman, N.C. A nanomechanical device based on the B-Z transition of DNA. Nature 397, 144–146 (1999).
Yan, H., Zhang, X., Shen, Z. & Seeman, N.C. A robust DNA mechanical device controlled by hybridization topology. Nature 415, 62–65 (2002).
Yurke, B., Turberfield, A.J., Mills, A.P. Jr., Simmel, F.C. & Neumann, J.L. DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Sherman, W.B. & Seeman, N.C. A precisely controlled DNA biped walking device. Nano Lett. 4, 1203–1207 (2004).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Li, J. & Bai, L. DNA nanotechnology. A metamaterial with memory. Nat. Nanotechnol. 7, 773–774 (2012).
Seeman, N.C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Lin, C., Liu, Y. & Yan, H. Designer DNA nanoarchitectures. Biochemistry 48, 1663–1674 (2009).
Simmel, F.C. Three-dimensional nanoconstruction with DNA. Angew. Chem. Int. Ed. 47, 5884–5887 (2008).
Feldkamp, U. & Niemeyer, C.M. Rational design of DNA nanoarchitectures. Angew. Chem. Int. Ed. 45, 1856–1876 (2006).
Deng, Z., Lee, S.H. & Mao, C. DNA as nanoscale building blocks. J. Nanosci. Nanotechnol. 5, 1954–1963 (2005).
Seeman, N.C. DNA in a material world. Nature 421, 427–431 (2003).
Eckardt, L.H. et al. DNA nanotechnology: chemical copying of connectivity. Nature 420, 286 (2002).
Seeman, N.C. DNA engineering and its application to nanotechnology. Trends Biotechnol. 17, 437–443 (1999).
Seeman, N.C. DNA nanotechnology: novel DNA constructions. Annu. Rev. Biophys. Biomol. Struct. 27, 225–248 (1998).
Mbindyo, J.K.N. et al. DNA-directed assembly of gold nanowires on complementary surfaces. Adv. Mater. 13, 249–254 (2001).
Perez, J.M., O'Loughin, T., Simeone, F.J., Weissleder, R. & Josephson, L. DNA-based magnetic nanoparticle assembly acts as a magnetic relaxation nanoswitch allowing screening of DNA-cleaving agents. J. Am. Chem. Soc. 124, 2856–2857 (2002).
Xiao, S. et al. Self-assembly of metallic nanoparticle arrays by DNA scaffolding. J. Nanopart. Res. 4, 313–317 (2002).
Deng, Z. & Mao, C. DNA-templated fabrication of 1D parallel and 2D crossed metallic nanowire arrays. Nano Lett. 3, 1545–1548 (2003).
Monson, C.F. & Woolley, A.T. DNA-templated construction of copper nanowires. Nano Lett. 3, 359–363 (2003).
Yan, H., Park, S.H., Finkelstein, G., Reif, J.H. & LaBean, T.H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
Keren, K., Berman, R.S., Buchstab, E., Sivan, U. & Braun, E. DNA-templated carbon nanotube field-effect transistor. Science 302, 1380–1382 (2003).
Mitchell, G.P., Mirkin, C.A. & Letsinger, R.L. Programmed assembly of DNA functionalized quantum dots. J. Am. Chem. Soc. 121, 8122–8123 (1999).
Xiong, X. et al. DNA aptamer-mediated cell targeting. Angew. Chem. Int. Ed. Engl. 52, 1472–1476 (2013).
Zhang, C. et al. Conformational flexibility facilitates self-assembly of complex DNA nanostructures. Proc. Natl. Acad. Sci. USA 105, 10665–10669 (2008).
Guo, P., Haque, F., Hallahan, B., Reif, R. & Li, H. Uniqueness, advantages, challenges, solutions, and perspectives in therapeutics applying RNA nanotechnology. Nucleic Acid Ther. 22, 226–245 (2012).
de Fougerolles, A., Vornlocher, H.P., Maraganore, J. & Lieberman, J. Interfering with disease: a progress report on siRNA-based therapeutics. Nat. Rev. Drug Discov. 6, 443–453 (2007).
Kim, D.H. & Rossi, J.J. Strategies for silencing human disease using RNA interference. Nat. Rev. Genet. 8, 173–184 (2007).
Rozema, D.B. et al. Dynamic PolyConjugates for targeted in vivo delivery of siRNA to hepatocytes. Proc. Natl. Acad. Sci. 104, 12982–12987 (2007).
Seth, S., Johns, R. & Templin, M.V. Delivery and biodistribution of siRNA for cancer therapy: challenges and future prospects. Ther. Deliv. 3, 245–261 (2012).
Liu, J. et al. Fabrication of stable and RNase-resistant RNA nanoparticles active in gearing the nanomotors for viral DNA packaging. ACS Nano 5, 237–246 (2010).
Abdelmawla, S. et al. Pharmacological characterization of chemically synthesized monomeric pRNA nanoparticles for systemic delivery. Molec. Ther. 19, 1312–1322 (2011).
Jain, K.K. The role of nanobiotechnology in drug discovery. Drug Discov. Today 10, 1435–1442 (2005).
Li, W. & Szoka, F. Lipid-based nanoparticles for nucleic acid delivery. Pharm. Res. 24, 438–449 (2007).
Gao, H., Shi, W. & Freund, L.B. Mechanics of receptor-mediated endocytosis. Proc. Natl. Acad. Sci. USA 102, 9469–9474 (2005).
Afonin, K.A., Danilov, E.O., Novikova, I.V. & Leontis, N.B. TokenRNA: A new type of sequence-specific, label-free fluorescent biosensor for folded RNA molecules. Chembiochem 9, 1902–1905 (2008).
Ferapontova, E.E., Olsen, E.M. & Gothelf, K.V. An RNA aptamer-based electrochemical biosensor for detection of theophylline in serum. J. Am. Chem. Soc. 130, 4256–4258 (2008).
Watts, J.K., Deleavey, G.F. & Damha, M.J. Chemically modified siRNA: tools and applications. Drug Discov. Today 13, 842–855 (2008).
Behlke, M.A. Chemical modification of siRNAs for in vivo use. Oligonucleotides 18, 305–319 (2008).
Corey, D.R. Chemical modification: the key to clinical application of RNA interference? J. Clin. Invest. 117, 3615–3622 (2007).
Choung, S., Kim, Y.J., Kim, S., Park, H.O. & Choi, Y.C. Chemical modification of siRNAs to improve serum stability without loss of efficacy. Biochem. Biophys. Res. Commun. 342, 919–927 (2006).
Layzer, J.M. et al. In vivo activity of nuclease-resistant siRNAs. RNA 10, 766–771 (2004).
Braasch, D.A. et al. RNA interference in mammalian cells by chemically-modified RNA. Biochemistry 42, 7967–7975 (2003).
Salomone, F. et al. A novel chimeric cell-penetrating peptide with membrane-disruptive properties for efficient endosomal escape. J. Control Release 163, 293–303 (2012).
Liou, J.S. et al. Protein transduction in human cells is enhanced by cell-penetrating peptides fused with an endosomolytic HA2 sequence. Peptides 37, 273–284 (2012).
Mellman, I., Fuchs, R. & Helenius, A. Acidification of the endocytic and exocytic pathways. Annu. Rev. Biochem. 55, 663–700 (1986).
Varkouhi, A.K., Scholte, M., Storm, G. & Haisma, H.J. Endosomal escape pathways for delivery of biologicals. J. Control Release 151, 220–228 (2011).
Li, N., Yu, C. & Huang, F. Novel cyanine-AMP conjugates for efficient 5′ RNA fluorescent labeling by one-step transcription and replacement of [γ-32P]ATP in RNA structural investigation. Nucleic Acids Res. 33, e37 (2005).
Sousa, R. Use of T7 RNA polymerase and its mutants for incorporation of nucleoside analogs into RNA. Methods Enzymol. 317, 65–74 (2000).
Sousa, R. & Padilla, R. A mutant T7 RNA polymerase as a DNA polymerase. EMBO J. 14, 4609–4621 (1995).
Shu, Y., Cinier, M., Fox, S.R., Ben-Johnathan, N. & Guo, P. Assembly of therapeutic pRNA-siRNA nanoparticles using bipartite approach. Molec. Ther. 19, 1304–1311 (2011).
Guha, M. et al. Endogenous tumor suppression mediated by PTEN involves survivin gene silencing. Cancer Res. 69, 4954–4958 (2009).
Fulda, S. & Debatin, K.M. Sensitization for tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis by the chemopreventive agent resveratrol. Cancer Res. 64, 337–346 (2004).
Bookout, A.L. & Mangelsdorf, D.J. Quantitative real-time PCR protocol for analysis of nuclear receptor signaling pathways. Nucl. Recept. Signal. 1, e012 (2003).
Gold, L. The SELEX process: a surprising source of therapeutic and diagnostic compounds. Harvey Lect. 91, 47–57 (1995).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA ploymerase. Science 249, 505–510 (1990).
Baugh, C., Grate, D. & Wilson, C. 2.8 A crystal structure of the malachite green aptamer. J. Mol. Biol. 301, 117–128 (2000).
Srisawat, C. & Engelke, D.R. Streptavidin aptamers: affinity tags for the study of RNAs and ribonucleoproteins. RNA 7, 632–641 (2001).
Guo, P., Shu, Y., Binzel, D. & Cinier, M. Synthesis, conjugation, and labeling of multifunctional pRNA nanoparticles for specific delivery of siRNA, drugs and other therapeutics to target cells. Methods Molec. Biol. 928, 197–219 (2012).
Huang, F. Efficient incorporation of CoA, NAD and FAD into RNA by in vitro transcription. Nucleic Acids Res. 31, e8 (2003).
Shu, D., Zhang, H., Jin, J. & Guo, P. Counting of six pRNAs of phi29 DNA-packaging motor with customized single molecule dual-view system. EMBO J. 26, 527–537 (2007).
Garver, K. & Guo, P. Mapping the inter-RNA interaction of phage phi29 by site-specific photoaffinity crosslinking. J. Biol. Chem. 275, 2817–2824 (2000).
Sampson, J.R. & Uhlenbeck, O.C. Biochemical and physical characterization of an unmodified yeast phenylalanine transfer RNA transcribed in vitro. Proc. Natl. Acad. Sci. USA 85, 1033–1037 (1988).
Shlyakhtenko, L.S. et al. Silatrane-based surface chemistry for immobilization of DNA, protein-DNA complexes and other biological materials. Ultramicroscopy 97, 279–287 (2003).
Lyubchenko, Y.L. & Shlyakhtenko, L.S. AFM for analysis of structure and dynamics of DNA and protein-DNA complexes. Methods 47, 206–213 (2009).
Searle, M.S. & Williams, D.H. On the stability of nucleic acid structures in solution: enthalpy-entropy compensations, internal rotations and reversibility. Nucleic Acids Res. 21, 2051–2056 (1993).
Kitamura, A., Jardine, P.J., Anderson, D.L., Grimes, S. & Matsuo, H. Analysis of intermolecular base pair formation of prohead RNA of the phage phi29 DNA packaging motor using NMR spectroscopy. Nucleic Acids Res. 36, 839–848 (2008).
Acknowledgements
The research was supported by US National Institutes of Health (NIH) grants EB003730 and CA151648, and by the Arnold and Mabel Beckman Initiative for Macular Research External Grant 1108 to P. Guo. AFM images were obtained by L. Shlyakhtenko and Y. Lyubchenko at the Nanoimaging Core Facility, University of Nebraska Medical Center, supported by the NIH Shared Instrumentations Grant Program and the University of Nebraska Medical Center Program Excellence (POE) and the Nebraska Research Initiative (NRI). We thank D. Rodger and M. Chow at University of Kentucky for the help with dynamic light scattering; F. Huang at The University of Southern Mississippi for providing Cy3- and Cy5-labeled AMP and GMP; and J. Haak, E. Khisamutdinov, E. Beabout, and F. Pi for help with the manuscript preparation.
Author information
Authors and Affiliations
Contributions
P.G. conceived and led the project. Y.S., D.S. and F.H. designed and conducted the experiments and co-wrote the manuscript with P.G.
Corresponding author
Ethics declarations
Competing interests
P.G. is a co-founder of Kylin Therapeutics and Biomotor and Nucleic Acid Nanotechnology Development Corp.
Rights and permissions
About this article
Cite this article
Shu, Y., Shu, D., Haque, F. et al. Fabrication of pRNA nanoparticles to deliver therapeutic RNAs and bioactive compounds into tumor cells. Nat Protoc 8, 1635–1659 (2013). https://doi.org/10.1038/nprot.2013.097
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.097
This article is cited by
-
Development of targeted therapy therapeutics to sensitize triple-negative breast cancer chemosensitivity utilizing bacteriophage phi29 derived packaging RNA
Journal of Nanobiotechnology (2021)
-
Short Interfering RNA (siRNA)-Based Therapeutics for Cartilage Diseases
Regenerative Engineering and Translational Medicine (2021)
-
Nanotechnology based approaches for detection and delivery of microRNA in healthcare and crop protection
Journal of Nanobiotechnology (2018)
-
Arrowtail RNA for Ligand Display on Ginger Exosome-like Nanovesicles to Systemic Deliver siRNA for Cancer Suppression
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.