Abstract
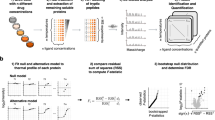
The detection and quantification of protein-ligand binding interactions is crucial in a number of different areas of biochemical research from fundamental studies of biological processes to drug discovery efforts. Described here is a protocol that can be used to identify the protein targets of biologically relevant ligands (e.g., drugs such as tamoxifen or cyclosporin A) in complex protein mixtures such as cell lysates. The protocol utilizes quantitative, bottom-up, shotgun proteomics technologies (isobaric mass tags for relative and absolute quantification, or iTRAQ) with a covalent labeling technique, termed stability of proteins from rates of oxidation (SPROX). In SPROX, the thermodynamic properties of proteins and protein-ligand complexes are assessed using the hydrogen peroxide–mediated oxidation of methionine residues as a function of the chemical denaturant (e.g., guanidine hydrochloride or urea) concentration. The proteome-wide SPROX experiments described here enable the ligand-binding properties of hundreds of proteins to be simultaneously assayed in the context of complex biological samples. The proteomic capabilities of the protocol render it amenable to the detection of both the on- and off-target effects of ligand binding. The protocol can be completed in 5 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
West, G.M., Tang, L. & Fitzgerald, M.C. Thermodynamic analysis of protein stability and ligand binding using a chemical modification- and mass spectrometry-based strategy. Anal. Chem. 80, 4175–4185 (2008).
Ghaemmaghami, S., Fitzgerald, M.C. & Oas, T.G. A quantitative, high-throughput screen for protein stability. Proc. Natl. Acad. Sci. USA 97, 8296–8301 (2000).
Powell, K.D. & Fitzgerald, M.C. Accuracy and precision of a new H/D exchange- and mass spectrometry-based technique for measuring the thermodynamic properties of protein-peptide complexes. Biochemistry 42, 4962–4670 (2003).
West, G.M. et al. Quantitative proteomics approach for identifying protein-drug interactions in complex mixtures using protein stability measurements. Proc. Natl. Acad. Sci. USA 107, 9078–9082 (2010).
Dearmond, P.D., Xu, Y., Strickland, E.C., Daniels, K.G. & Fitzgerald, M.C. Thermodynamic analysis of protein-ligand interactions in complex biological mixtures using a shotgun proteomics approach. J. Proteome Res. 10, 4948–4958 (2011).
Fields, S. & Song, O. A novel genetic system to detect protein-protein interactions. Nature 340, 245–246 (1989).
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 17, 1030–1032 (1999).
Lomenick, B. et al. Target identification using drug affinity responsive target stability (DARTS). Proc. Natl. Acad. Sci. USA 106, 21984–21989 (2009).
Liu, P.F., Kihara, D. & Park, C. Energetics-based discovery of protein-ligand interactions on a proteomic scale. J. Mol. Biol. 408, 147–162 (2011).
DeArmond, P.D., West, G.M., Huang, H.T. & Fitzgerald, M.C. Stable isotope labeling strategy for protein-ligand binding analysis in multi-component protein mixtures. J. Am. Soc. Mass Spectrom. 22, 418–430 (2011).
West, G.M. et al. Mass spectrometry-based thermal shift assay for protein-ligand binding analysis. Anal. Chem. 82, 5573–5581 (2010).
Nozaki, Y. The preparation of guanidine hydrochloride. Methods Enzymol. 26, 43–50 (1972).
Pace, C.N. Determination and analysis of urea and guanidine hydrochloride denaturation curves. Methods Enzymol. 131, 266–280 (1986).
Noble, R.W. & Gibson, Q.H. The reaction of ferrous horseradish peroxidase with hydrogen peroxide. J. Biol. Chem. 245, 2409–2413 (1970).
Shen, M. et al. Isolation and isotope labeling of cysteine- and methionine-containing tryptic peptides: application to the study of cell surface proteolysis. Mol. Cell Proteomics 2, 315–324 (2003).
Acknowledgements
This work was supported in part by US National Institutes of Health grant no. GM084174 (to M.C.F.) and in part by National Science Foundation (NSF) grant no. CHE-0848462 (to M.C.F.). The NSF grant, which was made possible with funds from the American Recovery and Reinvestment Act (ARRA), is jointly funded by the Analytical and Surface Chemistry Program in the Chemistry Division at NSF and by the Biomolecular Systems Cluster in the Division of Molecular and Cellular Biosciences at NSF. The mass spectrometer system used in this work was purchased with funds from National Institutes of Health grant no. S10RR027746 (to M.C.F.).
Author information
Authors and Affiliations
Contributions
G.M.W., P.D.D., Y.X., E.C.S., M.A.G., D.T.T. and J.A. contributed to the development and optimization of the described SPROX protocol. E.C.S. collected and analyzed the SPROX data highlighted in this manuscript. E.C.S., M.A.G., D.T.T., J.A. and M.C.F. drafted the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data 1
A sample data set on which Steps 36–43 were performed ("Sample_Data_Set.xlsx"). The data set is from the NAD binding study described in reference 5. (XLSX 6251 kb)
Supplementary Data 2
Runcompare input file used as a sample data set as described in Steps 44–46 (TXT 222 kb)
Supplementary Data 3
Runcompare input file used as a sample data set as described in Steps 44–46 (TXT 764 kb)
Supplementary Data 4
Output file generated from the sample data set as described in Steps 44–46 (TXT 303 kb)
Supplementary Data 5
The sample data set on which Steps 47–52 were performed ("Matched_NAD_Binding.xlsx") (XLSX 367 kb)
Rights and permissions
About this article
Cite this article
Strickland, E., Geer, M., Tran, D. et al. Thermodynamic analysis of protein-ligand binding interactions in complex biological mixtures using the stability of proteins from rates of oxidation. Nat Protoc 8, 148–161 (2013). https://doi.org/10.1038/nprot.2012.146
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.146
This article is cited by
-
Target identification of small molecules: an overview of the current applications in drug discovery
BMC Biotechnology (2023)
-
Sinomenine ameliorates collagen-induced arthritis in mice by targeting GBP5 and regulating the P2X7 receptor to suppress NLRP3-related signaling pathways
Acta Pharmacologica Sinica (2023)
-
Challenges and Perspectives in Target Identification and Mechanism Illustration for Chinese Medicine
Chinese Journal of Integrative Medicine (2023)
-
Target identification of natural medicine with chemical proteomics approach: probe synthesis, target fishing and protein identification
Signal Transduction and Targeted Therapy (2020)
-
Thermal proteome profiling: unbiased assessment of protein state through heat-induced stability changes
Proteome Science (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.