Abstract
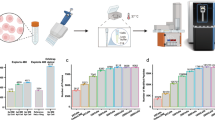
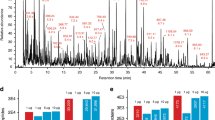
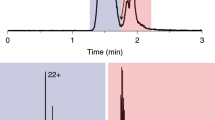
Multidimensional liquid chromatography (LC) combined with mass spectrometry (MS) has become a standard technique in proteomics to reduce sample complexity and to tackle the dynamic range in protein abundance. Fractionation is necessary to obtain a comprehensive analysis of complex biological samples such as tissue and mammalian cell lines. However, extensive fractionation comes at the expense of sample loss, presenting a bottleneck in the analysis of limited amounts of material. In this protocol, we describe a two-dimensional chromatographic strategy based on a combination of hydrophilic interaction liquid chromatography (HILIC; with a zwitterionic packing material, ZIC-cHILIC) and reversed-phase chromatography, which allows proteomic analyses with minimal sample loss. Experimental aspects related to obtaining maximum recovery are discussed, including how to optimally prepare samples for this system. Examples involving protein lysates originating from cultured cell lines and cells sorted by flow cytometry are used to show the power, sensitivity and versatility of the technique. Once the ZIC-cHILIC fractionation system has been optimized and standardized, this protocol requires ∼5–6 d, including sample preparation and fraction analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bensimon, A., Heck, A.J.R. & Aebersold, R. Mass spectrometry–based proteomics and network biology. Annu. Rev. Biochem. 81, 379–405 (2012).
Cox, J. & Mann, M. Quantitative, high-resolution proteomics for data-driven systems biology. Annu. Rev. Biochem. 80, 273–299 (2011).
Anderson, N.L. et al. A human proteome detection and quantitation project. Mol. Cell. Proteomics 8, 883–886 (2009).
Waanders, L.F. et al. Quantitative proteomic analysis of single pancreatic islets. Proc. Natl. Acad. Sci. USA 106, 18902–18907 (2009).
Di Palma, S. et al. Highly sensitive proteome analysis of FACS-sorted adult colon stem cells. J. Proteome Res. 10, 3814–3819 (2011).
Thakur, D. et al. Microproteomic analysis of 10,000 laser captured microdissected breast tumor cells using short-range sodium dodecyl sulfate-polyacrylamide gel electrophoresis and porous layer open tubular liquid chromatography tandem mass spectrometry. J. Chromatogr. A 1218, 8168–8174 (2011).
Masuda, T., Sugiyama, N., Tomita, M. & Ishihama, Y. Microscale phosphoproteome analysis of 10,000 cells from human cancer cell lines. Anal. Chem. 83, 7698–7703 (2011).
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011).
Koroleva, O.A. & Cramer, R. Single-cell proteomic analysis of glucosinolate-rich S-cells in Arabidopsis thaliana. Methods 54, 413–423 (2011).
Makarov, A. & Scigelova, M. Coupling liquid chromatography to Orbitrap mass spectrometry. J. Chromatogr. A 1217, 3938–3945 (2010).
Nagaraj, N. et al. System-wide perturbation analysis with nearly complete coverage of the yeast proteome by single-shot ultra HPLC runs on a bench top orbitrap. Mol. Cell. Proteomics 11, M111.013722 (2012).
Wisniewski, J.R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis 6, 359–362 (2009).
Liu, W. et al. Sample preparation method for isolation of single-cell types from mouse liver for proteomic studies. Proteomics 11, 3556–3564 (2011).
Thakur, S.S. et al. Deep and highly sensitive proteome coverage by LC-MS/MS without prefractionation. Mol. Cell. Proteomics 10, M110.003699 (2011).
Kocher, T., Pichler, P., Swart, R. & Mechtler, K. Analysis of protein mixtures from whole-cell extracts by single-run nanoLC-MS/MS using ultralong gradients. Nat. Protoc. 7, 882–890 (2012).
Waanders, L.F. et al. A novel chromatographic method allows on-line reanalysis of the proteome. Mol. Cell. Proteomics 7, 1452–1459 (2008).
Unwin, R.D., Griffiths, J.R. & Whetton, A.D. Simultaneous analysis of relative protein expression levels across multiple samples using iTRAQ isobaric tags with 2D nano LC-MS/MS. Nat. Protoc. 5, 1574–1582 (2010).
Shen, Y., Smith, R.D., Unger, K.K., Kumar, D. & Lubda, D. Ultrahigh-throughput proteomics using fast RPLC separations with ESI-MS/MS. Anal. Chem. 77, 6692–6701 (2005).
Dai, J. et al. Protein phosphorylation and expression profiling by yin-yang multidimensional liquid chromatography (Yin-Yang MDLC) mass spectrometry. J. Proteome Res. 6, 250–262 (2006).
Motoyama, A., Venable, J.D., Ruse, C.I. & Yates, J.R. Automated ultra-high-pressure multidimensional protein identification technology (UHP-MudPIT) for improved peptide identification of proteomic samples. Anal. Chem. 78, 5109–5118 (2006).
Altelaar, A.F.M. & Heck, A.J.R. Trends in ultrasensitive proteomics. Curr. Opin. Chem. Biol. 16, 206–213 (2012).
Bandura, D.R. et al. Mass cytometry: technique for real time single cell multitarget immunoassay based on inductively coupled plasma time-of-flight mass spectrometry. Anal. Chem. 81, 6813–6822 (2009).
Salehi-Reyhani, A. et al. A first step towards practical single cell proteomics: a microfluidic antibody capture chip with TIRF detection. Lab. Chip. 11, 1256–1261 (2011).
Washburn, M.P., Wolters, D. & Yates, J.R. 3rd. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Wolters, D.A., Washburn, M.P. & Yates, J.R. 3rd. An automated multidimensional protein identification technology for shotgun proteomics. Anal. Chem. 73, 5683–5690 (2001).
Motoyama, A., Xu, T., Ruse, C.I., Wohlschlegel, J.A. & Yates, J.R. 3rd. Anion and cation mixed-bed ion exchange for enhanced multidimensional separations of peptides and phosphopeptides. Anal. Chem. 79, 3623–3634 (2007).
Opiteck, G.J., Jorgenson, J.W. & Anderegg, R.J. Two-dimensional SEC/RPLC coupled to mass spectrometry for the analysis of peptides. Anal. Chem. 69, 2283–2291 (1997).
Dai, J. et al. Fully automatic separation and identification of phosphopeptides by continuous pH-gradient anion exchange online coupled with reversed-phase liquid chromatography mass spectrometry. J. Proteome Res. 8, 133–141 (2009).
Geng, M., Ji, J. & Regnier, F.E. Signature-peptide approach to detecting proteins in complex mixtures. J. Chromatogr. A 870, 295–313 (2000).
Gilar, M., Olivova, P., Daly, A.E. & Gebler, J.C. Two-dimensional separation of peptides using RP-RP-HPLC system with different pH in first and second separation dimensions. J. Sep. Sci. 28, 1694–1703 (2005).
Hao, P. et al. Novel application of electrostatic repulsion-hydrophilic interaction chromatography (ERLIC) in shotgun proteomics: comprehensive profiling of rat kidney proteome. J. Proteome Res. 9, 3520–3526 (2010).
Beausoleil, S.A. et al. Large-scale characterization of HeLa cell nuclear phosphoproteins. Proc. Natl. Acad. Sci. USA 101, 12130–12135 (2004).
Gauci, S. et al. Lys-N and trypsin cover complementary parts of the phosphoproteome in a refined SCX-based approach. Anal. Chem. 81, 4493–4501 (2009).
Ballif, B.A., Villen, J., Beausoleil, S.A., Schwartz, D. & Gygi, S.P. Phosphoproteomic analysis of the developing mouse brain. Mol. Cell. Proteomics 3, 1093–1101 (2004).
Lewandrowski, U., Zahedi, R.P., Moebius, J., Walter, U. & Sickmann, A. Enhanced N-glycosylation site analysis of sialoglycopeptides by strong cation exchange prefractionation applied to platelet plasma membranes. Mol. Cell. Proteomics 6, 1933–1941 (2007).
Boersema, P.J., Divecha, N., Heck, A.J. & Mohammed, S. Evaluation and optimization of ZIC-HILIC-RP as an alternative MudPIT strategy. J. Proteome Res. 6, 937–946 (2007).
Di Palma, S., Hennrich, M.L., Heck, A.J.R. & Mohammed, S. Recent advances in peptide separation by multidimensional liquid chromatography for proteome analysis. J. Proteomics 75, 3791–3813 (2012).
Alpert, A.J. Hydrophilic-interaction chromatography for the separation of peptides, nucleic acids and other polar compounds. J. Chromatogr. 499, 177–196 (1990).
Want, E.J. et al. Global metabolic profiling procedures for urine using UPLC-MS. Nat. Protoc. 5, 1005–1018 (2010).
Palmisano, G. et al. Selective enrichment of sialic acid-containing glycopeptides using titanium dioxide chromatography with analysis by HILIC and mass spectrometry. Nat. Protoc. 5, 1974–1982 (2010).
Jandera, P. Stationary phases for hydrophilic interaction chromatography, their characterization and implementation into multidimensional chromatography concepts. J. Sep. Sci. 31, 1421–1437 (2008).
McNulty, D.E. & Annan, R.S. Hydrophilic interaction chromatography reduces the complexity of the phosphoproteome and improves global phosphopeptide isolation and detection. Mol. Cell. Proteomics 7, 971–980 (2008).
Kolarich, D., Jensen, P.H., Altmann, F. & Packer, N.H. Determination of site-specific glycan heterogeneity on glycoproteins. Nat. Protoc. 7, 1285–1298 (2012).
Lindner, H., Sarg, B., Meraner, C. & Helliger, W. Separation of acetylated core histones by hydrophilic-interaction liquid chromatography. J. Chromatogr. A 743, 137–144 (1996).
Alpert, A.J. Electrostatic repulsion hydrophilic interaction chromatography for isocratic separation of charged solutes and selective isolation of phosphopeptides. Anal. Chem. 80, 62–76 (2008).
Intoh, A., Kurisaki, A., Fukuda, H. & Asashima, M. Separation with zwitterionic hydrophilic interaction liquid chromatography improves protein identification by matrix-assisted laser desorption/ionization-based proteomic analysis. Biomed. Chromatogr. 23, 607–614 (2009).
Jiang, W. & Irgum, K. Tentacle-type Zwitterionic stationary phase, prepared by surface-initiated graft polymerization of 3-N,N-dimethyl-N-(methacryloyloxyethyl)-ammonium propanesulfonate through peroxide groups tethered on porous silica. Anal. Chem. 74, 4682–4687 (2002).
Di Palma, S., Boersema, P.J., Heck, A.J.R. & Mohammed, S. Zwitterionic hydrophilic interaction liquid chromatography (ZIC-HILIC and ZIC-cHILIC) provide high resolution separation and increase sensitivity in proteome analysis. Anal. Chem. 83, 3440–3447 (2011).
Fountain, K.J., Xu, J., Diehl, D.M. & Morrison, D. Influence of stationary phase chemistry and mobile-phase composition on retention, selectivity, and MS response in hydrophilic interaction chromatography. J. Sep. Sci. 33, 740–751 (2010).
Boersema, P.J., Mohammed, S. & Heck, A.J. Hydrophilic interaction liquid chromatography (HILIC) in proteomics. Anal. Bioanal. Chem. 391, 151–159 (2008).
Grumbach, E.S., Diehl, D.M. & Neue, U.D. The application of novel 1.7 μm ethylene bridged hybrid particles for hydrophilic interaction chromatography. J. Sep. Sci. 31, 1511–1518 (2008).
Gilar, M., Olivova, P., Daly, A.E. & Gebler, J.C. Orthogonality of separation in two-dimensional liquid chromatography. Anal. Chem. 77, 6426–6434 (2005).
Jiang, W., Fischer, G., Girmay, Y. & Irgum, K. Zwitterionic stationary phase with covalently bonded phosphorylcholine type polymer grafts and its applicability to separation of peptides in the hydrophilic interaction liquid chromatography mode. J. Chromatogr. A 1127, 82–91 (2006).
Di Palma, S., Raijmakers, R., Heck, A.J.R. & Mohammed, S. Evaluation of the deuterium isotope effect in zwitterionic hydrophilic interaction liquid chromatography separations for implementation in a quantitative proteomic approach. Anal. Chem. 83, 8352–8356 (2011).
Hemstrom, P. & Irgum, K. Hydrophilic interaction chromatography. J. Sep. Sci. 29, 1784–1821 (2006).
Boersema, P.J., Aye, T.T., van Veen, T.A., Heck, A.J. & Mohammed, S. Triplex protein quantification based on stable isotope labeling by peptide dimethylation applied to cell and tissue lysates. Proteomics 8, 4624–4632 (2008).
Boersema, P.J., Raijmakers, R., Lemeer, S., Mohammed, S. & Heck, A.J. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat. Protoc. 4, 484–494 (2009).
Raijmakers, R. et al. Automated online sequential isotope labeling for protein quantitation applied to proteasome tissue-specific diversity. Mol. Cell. Proteomics 7, 1755–1762 (2008).
Meiring, H.D., van der Heeft, E., ten Hove, G.J. & de Jong, A.P.J.M Nanoscale LC–MS(n): technical design and applications to peptide and protein analysis. J. Sep. Sci. 25, 557–568 (2002).
Frese, C.K. et al. Improved peptide identification by targeted fragmentation using CID, HCD and ETD on an LTQ-Orbitrap Velos. J. Proteome Res. 10, 2377–2388 (2011).
Cortes, H.J., Pfeiffer, C.D., Richter, B.E. & Stevens, T.S. Porous ceramic bed supports for fused silica packed capillary columns used in liquid chromatography. J. High Resol. Chromatogr. 10, 446–448 (1987).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Acknowledgements
We thank P.J. Boersema and R. Raijmakers for suggestions and critical reading of the manuscript. We thank all members of the Biomolecular Mass Spectrometry and Proteomics Group, particularly H. Post, A. Barendregt and S. Goerdayal, for their help. This work was in part supported by the PRIME-XS project, grant agreement number 262067, funded by the European Union Seventh Framework Program. The Netherlands Proteomics Centre, embedded in The Netherlands Genomics Initiative and The Netherlands Organization for Scientific Research (NWO) with the VIDI grant 700.10.429 (for S.M.), is acknowledged for funding.
Author information
Authors and Affiliations
Contributions
S.D.P. designed and performed the experiments, analyzed data and wrote the paper. S.M. and A.J.R.H. designed the experiments, supervised the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Methods
Alternative sample preparation procedure for larger amounts of starting material (PDF 395 kb)
Rights and permissions
About this article
Cite this article
Di Palma, S., Mohammed, S. & Heck, A. ZIC-cHILIC as a fractionation method for sensitive and powerful shotgun proteomics. Nat Protoc 7, 2041–2055 (2012). https://doi.org/10.1038/nprot.2012.124
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.124
This article is cited by
-
Improving deep proteome and PTMome coverage using tandem HILIC-HPRP peptide fractionation strategy
Analytical and Bioanalytical Chemistry (2019)
-
Genome-wide transcription start site mapping of Bradyrhizobium japonicum grown free-living or in symbiosis – a rich resource to identify new transcripts, proteins and to study gene regulation
BMC Genomics (2016)
-
Mass spectrometry-based N-glycoproteomics for cancer biomarker discovery
Clinical Proteomics (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.