Abstract
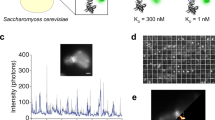
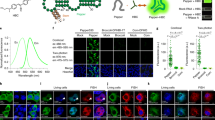
Protein removal has a central role in numerous cellular processes. Obtaining systematic measurements of multiple protein removal rates is necessary to understand the principles that govern these processes, but it is currently a major technical challenge. To address this, we developed 'bleach-chase', a noninvasive method for measuring the half-lives of multiple proteins at high temporal resolution in living cells. The method uses a library of annotated human reporter cell clones, each with a unique fluorescently tagged protein expressed from its native chromosomal location. In this protocol, we detail a simple procedure that bleaches the cells and uses time-lapse fluorescence microscopy and automated image analysis to systematically measure the half-life dynamics of multiple proteins. The duration of the protocol is 4–5 d. The method may be applicable to a wide range of fluorescently tagged proteins and cell lines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ciechanover, A. Proteolysis: from the lysosome to ubiquitin and the proteasome. Nat. Rev. Mol. Cell Biol. 6, 79–87 (2005).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Schwartz, A.L. & Ciechanover, A. The ubiquitin-proteasome pathway and pathogenesis of human diseases. Annu. Rev. Med. 50, 57–74 (1999).
Nakayama, K.I. & Nakayama, K. Ubiquitin ligases: cell-cycle control and cancer. Nat. Rev. Cancer 6, 369–381 (2006).
Pratt, J.M. et al. Dynamics of protein turnover, a missing dimension in proteomics. Mol. Cell. Proteom. 1, 579–591 (2002).
Alon, U. An Introduction to Systems Biology (Chapman & Hall, 2007).
Schimke, R.T. & Doyle, D. Control of enzyme levels in animal tissues. Annu. Rev. Biochem. 39, 929–976 (1970).
Belle, A., Tanay, A., Bitincka, L., Shamir, R. & O'Shea, E.K. Quantification of protein half-lives in the budding yeast proteome. Proc. Natl. Acad. Sci. USA 103, 13004–13009 (2006).
Shwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Yen, H.-C.S., Xu, Q., Chou, D.M., Zhao, Z. & Elledge, S.J. Global protein stability profiling in mammalian cells. Science 322, 918–923 (2008).
Eden, E. et al. Proteome half-life dynamics in living human cells. Science 331, 764–768 (2011).
Cohen, A.A. et al. Dynamic proteomics of individual cancer cells in response to a drug. Science 322, 1511–1516 (2008).
Sigal, A. et al. Generation of a fluorescently labeled endogenous protein library in living human cells. Nat. Protoc. 2, 1515–1527 (2007).
Sigal, A. et al. Dynamic proteomics in individual human cells uncovers widespread cell-cycle dependence of nuclear proteins. Nat. Methods 3, 525–531 (2006).
Hampton, R.Y., Koning, A., Wright, R. & Rine, J. In vivo examination of membrane protein localization and degradation with green fluorescent protein. Proc. Natl. Acad. Sci. USA 93, 828–833 (1996).
Li, X. et al. Characterization of NFκB activation by detection of green fluorescent protein-tagged IκB degradation in living cells. J. Biol. Chem. 274, 21244–21250 (1999).
Zhang, L., Gurskaya, N. & Lukyanov, K. Method for real-time monitoring of protein degradation at the single cell level. Biotechniques 42, 446–450 (2007).
Bellanger, S., Demeret, C., Goyat, S. & Thierry, F. Stability of the human papillomavirus type 18 E2 protein is regulated by a proteasome degradation pathway through its amino-terminal transactivation domain. J. Virol. 75, 7244–7251 (2001).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Jarvik, J.W. et al. CD-tagging: a new approach to gene and protein discovery and analysis. Biotechniques 20, 896–904 (1996).
Morin, X. et al. A protein trap strategy to detect GFP-tagged proteins expressed from their endogenous loci in Drosophila. Proc. Natl. Acad. Sci. USA 98, 15050–15055 (2001).
Ciechanover, A. & Schwartz, A.L. How are substrates recognized by the ubiquitin-mediated proteolytic system? Trends Biochem. Sci. 14, 483–488 (1989).
Varshavsky, A. The N-end rule pathway of protein degradation. Genes Cells 2, 13–28 (1997).
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Liu, H., Krizek, J. & Bretscher, A. Construction of a GAL1-regulated yeast cDNA expression library and its application to the identification of genes whose overexpression causes lethality in yeast. Genetics 132, 665–673 (1992).
Buszczak, M. et al. The carnegie protein trap library: a versatile tool for Drosophila developmental studies. Genetics 175, 1505–1531 (2007).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Megerle, J.A., Fritz, G., Gerland, U., Jung, K. & Rädler, J.O. Timing and dynamics of single cell gene expression in the arabinose utilization system. Biophys. J. 95, 2103–2115 (2008).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Sys. Man Cyber. 9, 62–66 (1979).
Adams, R. & Bischof, L. Seeded region growing. IEEE Trans. Pattern Anal. Mach. Intell. 16, 641–647 (1994).
Eden, E. et al. An automated method for analysis of flow characteristics of circulating particles from in vivo video microscopy. IEEE Trans. Med. Imaging 24, 1011–1024 (2005).
Acknowledgements
We thank Z. Yakhini, Z. Kam, R. Milo, P. Tsvetkov, J. Adler and all members of the Alon laboratory for fruitful discussions and comments. We thank P. Choukroun for assistance with the computer cluster. This work was supported by the European Research Council, the Kahn Family Foundation and by Keren Isra-Pa'amei Tikva.
Author information
Authors and Affiliations
Contributions
E.E., N.G.-Z. and U.A. led the study and designed the experiments. E.E. and A.M. performed the analysis. N.G.-Z., T.D., L.C. and I.I. performed experiments. Y.L. designed the microscope operating software. E.D. and A.C. contributed to the analysis. E.E., N.G.-Z. and U.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Weizmann Institute of Science has filed for a patent on the technology described in this report for measuring protein half-lives.
Rights and permissions
About this article
Cite this article
Geva-Zatorsky, N., Issaeva, I., Mayo, A. et al. Using bleach-chase to measure protein half-lives in living cells. Nat Protoc 7, 801–811 (2012). https://doi.org/10.1038/nprot.2012.028
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.028
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.