Abstract
Large-insert BAC (bacterial artificial chromosome) and BIBAC (binary BAC) libraries are essential for modern genomics research for all organisms. We helped pioneer the BAC and BIBAC technologies, and by using them we have constructed hundreds of BAC and BIBAC libraries for different species of plants, animals, marine animals, insects, algae and microbes. These libraries have been used globally for different aspects of genomics research. Here we describe the procedure with the latest improvements that we have made and used for construction of BIBAC libraries. The procedure includes the preparation of BIBAC vectors, the preparation of clonable fragments of the desired size from the source DNA, the construction and transformation of BIBACs and, finally, the characterization and assembly of BIBAC libraries. We also specify the modifications necessary for construction of BAC libraries using the protocol. The entire protocol takes ∼7 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Burke, D.T., Carle, G.F. & Olson, M.V. Cloning of large segments of exogenous DNA into yeast by means of artificial chromosome vectors. Science 236, 806–811 (1987).
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Echerichia coli using an F-factor-based vector. Proc. Natl. Acad. Sci. USA 89, 8794–8797 (1992).
Ioannou, P.A. et al. A new bacteriophage P1-derived vector for the propagation of large human DNA fragments. Nat. Genet. 6, 84–89 (1994).
Hamilton, C.M., Frary, A., Lewis, C. & Tanksley, S.D. Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc. Natl. Acad. Sci. USA 93, 9975–9979 (1996).
Tao, Q.-Z. & Zhang, H.-B. Cloning and stable maintenance of DNA fragments over 300 kb in Escherichia coli with conventional plasmid-based vectors. Nucleic Acids Res. 26, 4901–4909 (1998).
Liu, Y. et al. Complementation of plant mutants with large genomic DNA fragments by a transformation-competent artificial chromosome vector accelerates positional cloning. Proc. Natl. Acad. Sci. USA 96, 6535–6540 (1999).
Zhang, H.-B. Map-based cloning of genes and quantitative trait loci. In Principles and Practices of Plant Genomics Vol. 1 Genome Mapping (eds. Kole, C. & Abbott A.G.) 229–267 (Science Publishers, 2007).
Gregory, S.G., Howell, G.R. & Bentley, D.R. Genome mapping by fluorescent fingerprinting. Genome Res. 7, 1162–1168 (1997).
Marra, M. et al. High throughput fingerprint analysis of large-insert clones. Genome Res. 7, 1072–1084 (1997).
Zhang, H.-B. & Wing, R.A. Physical mapping of the rice genome with BACs. Plant Mol. Biol. 35, 115–127 (1997).
Wu, C. et al. Whole genome physical mapping: an overview on methods for DNA fingerprinting. In Handbook of Plant Genome Mapping: Genetic and Physical Mapping (eds. Meksem, K. & Kahl, G.) 257–284 (Wiley-VCH, 2005).
Chang, Y.-L., Tao, Q.-Z., Scheuring, C., Meksem, K. & Zhang, H.-B. An integrated map of Arabidopsis thaliana for functional analysis of its genome sequence. Genetics 159, 1231–1242 (2001).
Tao, Q.-Z. et al. Bacterial artificial chromosome-based physical map of the rice genome constructed by restriction fingerprint analysis. Genetics 158, 1711–1724 (2001).
Ren, C. et al. A BAC-based physical map of the chicken genome. Genome Res. 13, 2754–2758 (2003).
Wallis, J.W. et al. A physical map of the chicken genome. Nature 432, 761–764 (2004).
Wu, C. et al. A BAC and BIBAC-based physical map of the soybean genome. Genome Res. 14, 319–326 (2004).
Zhang, X. et al. An integrated BAC and genome sequence physical map of Phytophthora sojae. Mol. Plant-Microbe Interact. 19, 1302–1310 (2006).
Zhang, X. et al. A BAC/BIBAC-based physical map of chickpea, Cicer arietinum L. BMC Genomics 11, 501 (2010).
Zhang, Y. et al. A comparative physical map reveals the pattern of chromosomal evolution between the turkey (Meleagris gallopavo) and chicken (Gallus gallus) genomes. BMC Genomics 12, 447 (2011).
Venter, J.C., Smith, H.O. & Hood, L. A new strategy for genome sequencing. Nature 381, 364–366 (1996).
Zhang, H.-B. & Wu, C.C. BACs as tools for genome sequencing. Plant Physiol. Biochem. 39, 195–209 (2001).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature 436, 793–800 (2005).
Tyler, B.M. et al. Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313, 1261–1266 (2006).
Schnable, P.S. et al. The B73 maize genome: complexity, diversity, and dynamics. Science 326, 1112–1115 (2009).
Steuernagel, B. et al. De novo 454 sequencing of barcoded BAC pools for comprehensive gene survey and genome analysis in the complex genome of barley. BMC Genomics 10, 247 (2009).
Dalloul, R.A. et al. Multi-platform next generation sequencing of the domestic turkey (Meleagris gallopavo): Genome assembly and analysis. PLoS Biol. 8, e1000475 (2010).
Sato, K., Motoi, Y., Yamaji, N. & Yoshida, H. 454 sequencing of pooled BAC clones on chromosome 3H of barley. BMC Genomics 12, 246 (2011).
Song, R., Segal, G. & Messing, J. Expression of the sorghum 10-member kafirin gene cluster in maize endosperm. Nucleic Acids Res. 32, e189 (2004).
Chang, Y.-L. et al. A plant-transformation-ready large-insert BIBAC library of Arabidopsis and bombardment transformation and expression of its large-insert BIBACs in tobacco. Genome 54, 437–447 (2011).
Heintz, N. BAC to the future: the use of BAC transgenic mice for neuroscience research. Nat. Rev. Neurosci. 2, 861–870 (2001).
Marshall, V.M., Janette Allison, J., Templeton, T. & Foote, S.J. Generation of BAC transgenic mice. Methods Mol Biol. 256, 159–182 (2004).
Valjent, E., Bertran-Gonzalez, J., Hervé, D., Fisone, G. & Girault, J.A. Looking BAC at striatal signaling: cell-specific analysis in new transgenic mice. Trends Neurosci. 32, 538–547 (2009).
Johnson, S.J. & Wade-Martins, R. A BACwards glance at neurodegeneration: molecular insights into disease from LRRK2, SNCA and MAPT BAC-transgenic mice. Biochem. Soc. Trans. 39, 862–867 (2011).
Ren, C. et al. Genomic DNA libraries and physical mapping. In Handbook of Plant Genome Mapping: Genetic and Physical Mapping (eds. Meksem, K. & Kahl, G.) 173–213 (Wiley-VCH, 2005).
Wu, C., Xu, Z. & Zhang, H.-B. DNA Libraries. In Encyclopedia of Molecular Cell Biology and Molecular Medicine, 2nd edn. Vol. 3 (ed. Meyers, R.A.) 385–425 (Wiley-VCH, 2004).
Zhang, H.-B., Woo, S.-S. & Wing, R.A. BAC, YAC and cosmid library construction. In Plant Gene Isolation: Principles and Practice (eds. Foster, G. & Twell, D.) 75–99 (John Wiley & Sons., 1996).
Sambrook, J., Fritsch, E.E. & Maniatis, T. Molecular Cloning: Laboratory Manual 2 edn. Vol. 1, 1.42–1.43, 1.46, 1.90–1.104 (Cold Spring Harbor Laboratory Press, 1989).
Panter, S. et al. Molecular breeding of transgenic white clover (Trifolium repens L.) with field resistance to Alfalfa mosaic virus through the expression of its coat protein gene. Transgenic Res. doi: 10.1007/s11248-011-9557-z63 (2011).
Kamo, K., Jordan, R., Guaragna, M.A., Hsu, H.-T. & Ueng, P. Resistance to cucumber mosaic virus in Gladiolus plants transformed with either a defective replicase or coat protein subgroup II gene from Cucumber mosaic virus. Plant Cell Rep. 29, 695–704 (2010).
Strong, S.J., Ohta, Y., Litman, G.W. & Amemiya, C.T. Marked improvement of PAC and BAC cloning is achieved using electroelution of pulsed-field gel-separated partial digests of genomic DNA. Nucleic Acids Res. 25, 3959–3961 (1997).
Osoegawa, K. et al. An improved approach for construction of bacterial artificial chromosome libraries. Genomics 52, 1–8 (1998).
Zhang, H.-B., Choi, S.-D., Woo, S.-S., Li, Z.-K. & Wing, R.A. Construction and characterization of two rice bacterial artificial chromosome libraries from the parents of a permanent recombinant inbred mapping population. Mol. Breed 2, 11–24 (1996).
Hamilton, C.M., Frary, A., Xu, Y., Tanksley, S.D. & Zhang, H.-B. Construction of tomato genomic DNA libraries in a binary-BAC (BIBAC) vector. Plant J. 18, 223–229 (1999).
Lee, M.-K. et al. Construction and characterization of three complementary BAC libraries for analysis of the chicken genome. Animal Genet. 34, 151–152 (2003).
Zhang, Y. et al. Construction and characterization of two bacterial artificial chromosome libraries of Zhikong Scallop, Chlamys farreri Jones et Preston, and identification of BAC clones containing the genes involved in its innate immune system. Marine Biotechnol. 10, 358–365 (2008).
Zhang, X. et al. Construction and characterization of a bacterial artificial chromosome (BAC) library of pacific white shrimp, Liptopenaeus vannamei. Marine Biotechnol. 12, 141–149 (2010).
Shao, C.-W. et al. Construction of two BAC libraries of half-smooth tongue sole Cynoglossus semilaevis and isolation of clones containing candidate sex-determination genes. Marine Biotechnol. 12, 558–568 (2010).
Hong, Y.S. et al. Construction and characterization of a BAC library and generation of BAC end sequence-tagged connectors for genome sequencing of the malaria mosquito, Anopheles gambiae. Mol. Genet. Genomics 268, 720–728 (2003).
Wu, C.-C. et al. Construction and sequence sampling of deep-coverage, large-insert BAC libraries for three model Lepidopteran species. BMC Genomics 10, 283 (2009).
Xu, Z. et al. Genome-wide physical mapping from large-insert clones by fingerprint analysis with capillary electrophoresis: A robust physical map of Penicillium chrysogenum. Nucleic Acids Res. 33, e50 (2005).
Frary, A. & Hamilton, C.M. Efficiency and stability of high molecular weight DNA transformation: an analysis in tomato. Transgenic Res. 10, 121–132 (2001).
Liu, Y.G. et al. Development of new transformation-competent artificial chromosome vectors and rice genomic libraries for efficient gene cloning. Gene 282, 247–255 (2002).
He, R.F. et al. Construction of a genomic library of wild rice and Agrobacterium-mediated transformation of large insert DNA linked to BPH resistance locus. Gene 321, 113–121 (2003).
Ercolano, M.R. et al. Functional complementation analysis in potato via biolistic transformation with BAC large DNA fragments. Mol. Breed. 13, 15–22 (2004).
Phan, B.H. et al. Transformation of rice with long DNA-segments consisting of random genomic DNA or centromere-specific DNA. Transgenic Res. 16, 341–351 (2007).
Li, Y. et al. A plant-transformation-competent BIBAC/BAC-based map of rice for functional analysis and genetic engineering of its genomic sequence. Genome 50, 278–288 (2007).
Zhang, M.P. et al. Preparation of megabase-sized DNA from a variety of organisms using the nuclei method for advanced genomics research. Nat. Protoc. 7, 467–478 (2012).
Larin, Z., Monaco, A.P. & Lehrach, H. Generation of large insert YAC library. Methods Mol. Biol. 54, 1–11 (1996).
Vinatzer, B.A., Zhang, H.-B. & Sanasavini, S. Construction and characterization of a BAC library of apple (Malus x domestica Borkh.). Theor. Appl. Genet. 97, 1183–1190 (1998).
Jones, J.D.G. et al. Effective vectors for transformation, expression of heterologous genes, and assaying transposon excision in transgenic plants. Transgenic Res. 1, 285–297 (1992).
Hamilton, C.M. A binary-BAC system for plant transformation with high-molecular-weight DNA. Gene 200, 107–116 (1997).
Frijters, A.C.J. et al. Construction of a bacterial artificial chromosome library containing large Eco RI and Hind III genomic fragments of lettuce. Theor. Appl. Genet. 94, 390–399 (1997).
Kim, U.J. et al. Construction and characterization of a human bacterial artificial chromosome library. Genomics 34, 213–218 (1996).
Acknowledgements
We thank the former members of H.-B.Z.'s laboratory who have used the methods described in this article for their research and thus might have directly or indirectly contributed to the procedures. This work is supported in part by Research Grant Award no. IS-4427-11C from BARD, the United States-Israel Binational Agricultural Research and Development Fund (H.-B.Z.) and grants (124329-85360; 124475-85360) from the Texas AgriLife Research Cotton and Monocot Crop Improvement Program (H.-B.Z.).
Author information
Authors and Affiliations
Contributions
H.-B.Z., C.F.S., M.Z., Y.Z., C.-C.W., J.J.D. and Y.L. conducted the experiments on different species with the protocols presented here, and improved and extended the original protocol through these experiments. Furthermore, H.-B.Z. developed the original concept of the protocols and the original protocols, designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
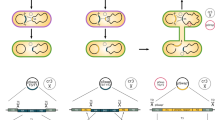
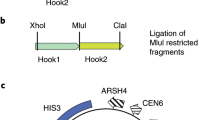
BAC and BIBAC vectors widely used in the construction of BIBAC and BAC libraries. (a) pECBAC1 (ref. 63) and (b) pBeloBAC11 (ref. 64) are both BAC vectors, whereas (c) pCLD04541 (561, d) BIBAC2 (462) and (e) pYLTAC7 (ref. 6) are BIBAC vectors. The cloning sites of restriction enzyme(s), the gene marker for selection of transformants (CMR, chloramphenicol; TetR, tetracycline; KmR, kanamycin) and the gene marker for selection of clones (Lac Z or Sac B) containing inserts or recombinant clones are highlighted or in bold. The figure was from Ref. 35 with permission from its publisher, Wiley-VCH. (TIFF 5426 kb)
Rights and permissions
About this article
Cite this article
Zhang, HB., Scheuring, C., Zhang, M. et al. Construction of BIBAC and BAC libraries from a variety of organisms for advanced genomics research. Nat Protoc 7, 479–499 (2012). https://doi.org/10.1038/nprot.2011.456
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.456
This article is cited by
-
A novel in-situ-process technique constructs whole circular cpDNA library
Plant Methods (2024)
-
A simple plant high-molecular-weight DNA extraction method suitable for single-molecule technologies
Plant Methods (2020)
-
RETRACTED ARTICLE: An assessment of transgenomics as a tool for gene discovery in Populus euphratica Oliv.
Plant Molecular Biology (2018)
-
Oak genome reveals facets of long lifespan
Nature Plants (2018)
-
MISSA 2.0: an updated synthetic biology toolbox for assembly of orthogonal CRISPR/Cas systems
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.