Abstract
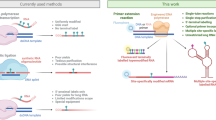
RNA SHAPE chemistry yields quantitative, single-nucleotide resolution structural information based on the reaction of the 2′-hydroxyl group of conformationally flexible nucleotides with electrophilic SHAPE reagents. However, SHAPE technology has been limited by the requirement that sites of RNA modification be detected by primer extension. Primer extension results in loss of information at both the 5′ and 3′ ends of an RNA and requires multiple experimental steps. Here we describe RNase-detected SHAPE that uses a processive, 3′→5′ exoribonuclease, RNase R, to detect covalent adducts in 5′-end–labeled RNA in a one-tube experiment. RNase R degrades RNA but stops quantitatively three and four nucleotides 3′ of a nucleotide containing a covalent adduct at the ribose 2′-hydroxyl or the pairing face of a nucleobase, respectively. We illustrate this technology by characterizing ligand-induced folding for the aptamer domain of the Escherichia coli thiamine pyrophosphate riboswitch RNA. RNase-detected SHAPE is a facile, two-day approach that can be used to analyze diverse covalent adducts in any RNA molecule, including short RNAs not amenable to analysis by primer extension and RNAs with functionally important structures at their 5′ or 3′ ends.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gesteland, R.F., Cech, T.R. & Atkins, J.F. The RNA World (Cold Spring Harbor Laboratory Press, 2004).
Li, P.T., Vieregg, J. & Tinoco, I. Jr. How RNA unfolds and refolds. Annu. Rev. Biochem. 77, 77–100 (2008).
Brunel, C. & Romby, P. Probing RNA structure and RNA-ligand complexes with chemical probes. Methods Enzymol. 318, 3–21 (2000).
Weeks, K.M. Advances in RNA structure analysis by chemical probing. Curr. Opin. Struct. Biol. 20, 295–304 (2010).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. RNA SHAPE chemistry reveals nonhierarchical interactions dominate equilibrium structural transitions in tRNA(Asp) transcripts. J. Am. Chem. Soc. 127, 4659–4667 (2005).
Steen, K.A., Malhotra, A. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by protection from exoribonuclease. J. Am. Chem. Soc. 132, 9940–9943 (2010).
Wilkinson, K.A. et al. High-throughput SHAPE analysis reveals structures in HIV-1 genomic RNA strongly conserved across distinct biological states. PLoS Biol. 6, e96 (2008).
Watts, J.M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Wilkinson, K.A. et al. Influence of nucleotide identity on ribose 2′-hydroxyl reactivity in RNA. RNA 15, 1314–1321 (2009).
Gherghe, C.M., Shajani, Z., Wilkinson, K.A., Varani, G. & Weeks, K.M. Strong correlation between SHAPE chemistry and the generalized NMR order parameter (S2) in RNA. J. Am. Chem. Soc. 130, 12244–12245 (2008).
Mortimer, S.A. & Weeks, K.M. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J. Am. Chem. Soc. 129, 4144–4145 (2007).
Badorrek, C.S. & Weeks, K.M. Architecture of a gamma retroviral genomic RNA dimer. Biochemistry 45, 12664–12672 (2006).
Duncan, C.D. & Weeks, K.M. Nonhierarchical ribonucleoprotein assembly suggests a strain-propagation model for protein-facilitated RNA folding. Biochemistry 49, 5418–5425 (2010).
Wang, B., Wilkinson, K.A. & Weeks, K.M. Complex ligand-induced conformational changes in tRNA(Asp) revealed by single-nucleotide resolution SHAPE chemistry. Biochemistry 47, 3454–3461 (2008).
Gherghe, C. et al. Definition of a high-affinity Gag recognition structure mediating packaging of a retroviral RNA genome. Proc. Natl. Acad. Sci. USA 107, 19248–19253 (2010).
Deigan, K.E., Li, T.W., Mathews, D.H. & Weeks, K.M. Accurate SHAPE-directed RNA structure determination. Proc. Natl. Acad. Sci. USA 106, 97–102 (2009).
Ehresmann, C. et al. Probing the structure of RNAs in solution. Nucleic Acids Res. 15, 9109–9128 (1987).
Stern, S., Moazed, D. & Noller, H.F. Structural analysis of RNA using chemical and enzymatic probing monitored by primer extension. Methods Enzymol. 164, 481–489 (1988).
Mitra, S., Shcherbakova, I.V., Altman, R.B., Brenowitz, M. & Laederach, A. High-throughput single-nucleotide structural mapping by capillary automated footprinting analysis. Nucleic Acids Res. 36, e63 (2008).
McGinnis, J.L., Duncan, C.D. & Weeks, K.M. High-throughput SHAPE and hydroxyl radical analysis of RNA structure and ribonucleoprotein assembly. Methods Enzymol. 468, 67–89 (2009).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat. Protoc. 1, 1610–1616 (2006).
Lalonde, M.S. et al. Exoribonuclease R in Mycoplasma genitalium can carry out both RNA processing and degradative functions and is sensitive to RNA ribose methylation. RNA 13, 1957–1968 (2007).
Vincent, H.A. & Deutscher, M.P. The roles of individual domains of RNase R in substrate binding and exoribonuclease activity. The nuclease domain is sufficient for digestion of structured RNA. J. Biol. Chem. 284, 486–494 (2009).
Matos, R.G., Barbas, A. & Arraiano, C.M. RNase R mutants elucidate the catalysis of structured RNA: RNA-binding domains select the RNAs targeted for degradation. Biochem. J. 423, 291–301 (2009).
Frazao, C. et al. Unravelling the dynamics of RNA degradation by ribonuclease II and its RNA-bound complex. Nature 443, 110–114 (2006).
Lorentzen, E., Basquin, J., Tomecki, R., Dziembowski, A. & Conti, E. Structure of the active subunit of the yeast exosome core, Rrp44: diverse modes of substrate recruitment in the RNase II nuclease family. Mol. Cell 29, 717–728 (2008).
Barbas, A. et al. Determination of key residues for catalysis and RNA cleavage specificity: one mutation turns RNase II into a 'SUPER-ENZYME'. J. Biol. Chem. 284, 20486–20498 (2009).
Mortimer, S.A. & Weeks, K.M. Time-resolved RNA SHAPE chemistry. J. Am. Chem. Soc. 130, 16178–16180 (2008).
Qin, P.Z. & Pyle, A.M. Site-specific labeling of RNA with fluorophores and other structural probes. Methods 18, 60–70 (1999).
Bakin, A.V., Borisova, O.F., Shatsky, I.N. & Bogdanov, A.A. Spatial organization of template polynucleotides on the ribosome determined by fluorescence methods. J. Mol. Biol. 221, 441–453 (1991).
Gherghe, C.M., Mortimer, S.A., Krahn, J.M., Thompson, N.L. & Weeks, K.M. Slow conformational dynamics at C2′-endo nucleotides in RNA. J. Am. Chem. Soc. 130, 8884–8885 (2008).
Weeks, K.M. & Mauger, D.M. Exploring RNA structural codes with SHAPE chemistry. Acc. Chem. Res. 44 (26 May 2011).
Das, R., Laederach, A., Pearlman, S.M., Herschlag, D. & Altman, R.B. SAFA: semi-automated footprinting analysis software for high-throughput quantification of nucleic acid footprinting experiments. RNA 11, 344–354 (2005).
Steen, K.A., Siegfried, N.A., & Weeks, K.M. Synthesis of 1-methyl-7-nitroisatoic anhydride (1M7). Protoc. Exch. Published online, doi:10.1038/protex.2011.255.
Vincent, H.A. & Deutscher, M.P. Insights into how RNase R degrades structured RNA: analysis of the nuclease domain. J. Mol. Biol. 387, 570–583 (2009).
Serganov, A., Polonskaia, A., Phan, A.T., Breaker, R.R. & Patel, D.J. Structural basis for gene regulation by a thiamine pyrophosphate-sensing riboswitch. Nature 441, 1167–1171 (2006).
Lang, K., Rieder, R. & Micura, R. Ligand-induced folding of the thiM TPP riboswitch investigated by a structure-based fluorescence spectroscopic approach. Nucleic Acids Res. 35, 5370–5378 (2007).
Kulshina, N., Edwards, T.E. & Ferre-D'Amare, A.R. Thermodynamic analysis of ligand binding and ligand binding-induced tertiary structure formation by the thiamine pyrophosphate riboswitch. RNA 16, 186–196 (2010).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Schagger, H. Tricine-SDS-PAGE. Nat. Protoc. 1, 16–22 (2006).
Acknowledgements
We are indebted to A. Malhotra for many productive discussions regarding the mechanism of RNase R enzymes and to Z. Li for the gift of a plasmid encoding M. genitalium RNase R optimized for expression in E. coli. We thank P. Homan for helpful discussions regarding synthesis of 1M7. This work was supported by a grant from the National Science Foundation (MCB-0919666 to K.M.W.); N.A.S. is a Fellow of the UNC Lineberger Comprehensive Cancer Center.
Author information
Authors and Affiliations
Contributions
K.-A.S. and K.M.W. collaborated on all aspects of the conception and writing of this paper. N.A.S. and K.M.W. wrote the Supporting Protocol on RNase R purification.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
A representative gel showing purification of the M. genitalium RNase R (75,000 Da) (PDF 215 kb)
Supplementary Data 1
Dataset corresponding to the experiment shown in Figure 3. (XLS 40 kb)
Rights and permissions
About this article
Cite this article
Steen, KA., Siegfried, N. & Weeks, K. Selective 2′-hydroxyl acylation analyzed by protection from exoribonuclease (RNase-detected SHAPE) for direct analysis of covalent adducts and of nucleotide flexibility in RNA. Nat Protoc 6, 1683–1694 (2011). https://doi.org/10.1038/nprot.2011.373
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.373
This article is cited by
-
Reversible 2′-OH acylation enhances RNA stability
Nature Chemistry (2023)
-
The chemistry and applications of RNA 2′-OH acylation
Nature Reviews Chemistry (2019)
-
New windows into retroviral RNA structures
Retrovirology (2018)
-
Transcriptional pausing at the translation start site operates as a critical checkpoint for riboswitch regulation
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.