Abstract
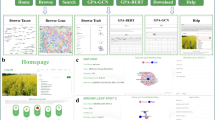
AraNet is a functional gene network for the reference plant Arabidopsis and has been constructed in order to identify new genes associated with plant traits. It is highly predictive for diverse biological pathways and can be used to prioritize genes for functional screens. Moreover, AraNet provides a web-based tool with which plant biologists can efficiently discover novel functions of Arabidopsis genes (http://www.functionalnet.org/aranet/). This protocol explains how to conduct network-based prediction of gene functions using AraNet and how to interpret the prediction results. Functional discovery in plant biology is facilitated by combining candidate prioritization by AraNet with focused experimental tests.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
McGary, K.L., Lee, I. & Marcotte, E.M. Broad network-based predictability of Saccharomyces cerevisiae gene loss-of-function phenotypes. Genome Biol. 8, R258 (2007).
Lehner, B. & Lee, I. Network-guided genetic screening: building, testing and using gene networks to predict gene function. Brief Funct. Genomic Proteomic 7, 217–227 (2008).
Alonso, J.M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Lee, I., Date, S.V., Adai, A.T. & Marcotte, E.M. A probabilistic functional network of yeast genes. Science 306, 1555–1558 (2004).
Marcotte, E.M., Pellegrini, M., Thompson, M.J., Yeates, T.O. & Eisenberg, D. A combined algorithm for genome-wide prediction of protein function. Nature 402, 83–86 (1999).
Fraser, H.B. & Plotkin, J.B. Using protein complexes to predict phenotypic effects of gene mutation. Genome Biol. 8, R252 (2007).
Lee, I. et al. A single gene network accurately predicts phenotypic effects of gene perturbation in Caenorhabditis elegans. Nat. Genet. 40, 181–188 (2008).
Zhong, W. & Sternberg, P.W. Genome-wide prediction of C. elegans genetic interactions. Science 311, 1481–1484 (2006).
Lee, I. et al. Predicting genetic modifier loci using functional gene networks. Genome Res. 20, 1143–1153 (2010).
Franke, L. et al. Reconstruction of a functional human gene network, with an application for prioritizing positional candidate genes. Am. J. Hum. Genet. 78, 1011–1025 (2006).
Huttenhower, C. et al. Exploring the human genome with functional maps. Genome Res. 19, 1093–1106 (2009).
Lage, K. et al. A human phenome-interactome network of protein complexes implicated in genetic disorders. Nat. Biotechnol. 25, 309–316 (2007).
Linghu, B., Snitkin, E.S., Hu, Z., Xia, Y. & Delisi, C. Genome-wide prioritization of disease genes and identification of disease-disease associations from an integrated human functional linkage network. Genome Biol. 10, R91 (2009).
Szklarczyk, D. et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 39, D561–D568 (2011).
Mostafavi, S., Ray, D., Warde-Farley, D., Grouios, C. & Morris, Q. GeneMANIA: a real-time multiple association network integration algorithm for predicting gene function. Genome Biol. 9 (Suppl 1): S4 (2008).
Lee, I., Ambaru, B., Thakkar, P., Marcotte, E.M. & Rhee, S.Y. Rational association of genes with traits using a genome-scale gene network for Arabidopsis thaliana. Nat. Biotechnol. 28, 149–156 (2010).
Cui, J. et al. AtPID: Arabidopsis thaliana protein interactome database—an integrative platform for plant systems biology. Nucleic Acids Res. 36, D999–D1008 (2008).
Geisler-Lee, J. et al. A predicted interactome for Arabidopsis. Plant Physiol. 145, 317–329 (2007).
Gutierrez, R.A. et al. Qualitative network models and genome-wide expression data define carbon/nitrogen-responsive molecular machines in Arabidopsis. Genome Biol. 8, R7 (2007).
Ma, S., Gong, Q. & Bohnert, H.J. An Arabidopsis gene network based on the graphical Gaussian model. Genome Res. 17, 1614–1625 (2007).
Fawcett, T. ROC Graphs: Notes and Practical Considerations for Data Mining Researchers (Hewlett-Packard Company, 2003).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Berardini, T.Z. et al. Functional annotation of the Arabidopsis genome using controlled vocabularies. Plant Physiol. 135, 745–755 (2004).
Avraham, S. et al. The Plant Ontology Database: a community resource for plant structure and developmental stages controlled vocabulary and annotations. Nucleic Acids Res. 36, D449–D454 (2008).
Hunter, S. et al. InterPro: the integrative protein signature database. Nucleic Acids Res. 37, D211–D215 (2009).
Rhee, S.Y. et al. The Arabidopsis Information Resource (TAIR): a model organism database providing a centralized, curated gateway to Arabidopsis biology, research materials and community. Nucleic Acids Res. 31, 224–228 (2003).
Remm, M., Storm, C.E. & Sonnhammer, E.L. Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J. Mol. Biol. 314, 1041–1052 (2001).
Blanc, G., Hokamp, K. & Wolfe, K.H. A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Res. 13, 137–144 (2003).
Lopes, C.T. et al. Cytoscape Web: an interactive web-based network browser. Bioinformatics 26, 2347–2348 (2010).
Cline, M.S. et al. Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2, 2366–2382 (2007).
Bard, J.B. & Rhee, S.Y. Ontologies in biology: design, applications and future challenges. Nat. Rev. Genet. 5, 213–222 (2004).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Danyluk, J., Carpentier, E. & Sarhan, F. Identification and characterization of a low temperature regulated gene encoding an actin-binding protein from wheat. FEBS Lett. 389, 324–327 (1996).
Scholl, R. & Anderson, M. Arabidopsis Biological Resource Center. Plant Mol. Bio. Rep. 12, 242–244 (1994).
Wang, P.I. & Marcotte, E.M. It's the machine that matters: predicting gene function and phenotype from protein networks. J. Proteomics 73, 2277–2289 (2010).
Acknowledgements
This work was supported by the National Research Foundation of Korea grant funded by the Korean government (no. 20100017649); Pohang Iron and Steel Company (POSCO) TJ Park Science Fellowship (to I.L.); grants from the National Science Foundation, National Institutes of Health, Welch (F1515) and Packard Foundations (to E.M.M.); the Carnegie Institution for Science; and a grant from the National Science Foundation (to S.Y.R.).
AUTHOR CONTRIBUTIONS
I.L., E.M.M., S.Y.R. and S.H. conceived the protocol. S.H. constructed the current version of the AraNet web application and wrote the protocol. I.L., E.M.M. and S.Y.R. edited the protocol.
Author information
Authors and Affiliations
Contributions
I.L., E.M.M., S.Y.R. and S.H. conceived the protocol. S.H. constructed the current version of the AraNet web application and wrote the protocol. I.L., E.M.M. and S.Y.R. edited the protocol.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Hwang, S., Rhee, S., Marcotte, E. et al. Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network. Nat Protoc 6, 1429–1442 (2011). https://doi.org/10.1038/nprot.2011.372
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.372
This article is cited by
-
A Network of Splice Isoforms for the Mouse
Scientific Reports (2016)
-
Pan- and core- network analysis of co-expression genes in a model plant
Scientific Reports (2016)
-
Associations of SNPs located at candidate genes to bovine growth traits, prioritized with an interaction networks construction approach
BMC Genetics (2015)
-
Selecting causal genes from genome-wide association studies via functionally coherent subnetworks
Nature Methods (2015)
-
WikiPathways for plants: a community pathway curation portal and a case study in rice and arabidopsis seed development networks
Rice (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.