Abstract
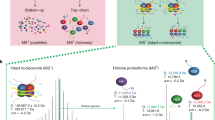
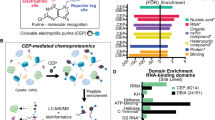
Post-translational modifications (PTMs) of transcription factors alter interactions with co-regulators and epigenetic modifiers. For example, members of the C/EBP transcription factor family are extensively methylated on arginine and lysine residues in short, conserved, modular domains, implying modification-dependent cofactor docking. Here we describe array peptide screening (APS), a systematic and differential approach to detect PTM-dependent interactions in the human proteome using chemically synthesized, biotinylated peptides coupled to fluorophore-labeled streptavidin. Peptides with and without a modified residue are applied in parallel to bacterial expression libraries in an arrayed format. Interactions are detected and quantified by laser scanning to reveal proteins that differentially bind to nonmodified or modified peptides. We have previously used this method to investigate the effect of arginine methylation of C/EBPβ peptides. The method enables determination of PTM-dependent transcription factor interactions, quantification of relative binding affinities and rapid protein classification, all independently of the transactivation potential of peptides or cellular abundance of interactors. The protocol provides a cost-effective alternative to mass spectrometric approaches and takes 3–4 d to complete.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Taverna, S.D. et al. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat. Struct. Mol. Biol. 14, 1025–1040 (2007).
Huang, J. et al. Repression of p53 activity by Smyd2-mediated methylation. Nature 444, 629–632 (2006).
Jansson, M. et al. Arginine methylation regulates the p53 response. Nat. Cell Biol. 10, 1431–1439 (2008).
Yamagata, K. et al. Arginine methylation of FOXO transcription factors inhibits their phosphorylation by Akt. Mol. Cell 32, 221–231 (2008).
Kowenz-Leutz, E. et al. Crosstalk between C/EBPbeta phosphorylation, arginine methylation, and SWI/SNF/Mediator implies an indexing transcription factor code. EMBO J. 29, 1105–1115 (2010).
Edmondson, D.G. & Roth, S.Y. Identification of protein interactions by far western analysis. Curr. Protoc. Protein Sci. 19, 19.7.1–19.7.10 (2001).
Seet, B.T. et al. Reading protein modifications with interaction domains. Nat. Rev. Mol. Cell Biol. 7, 473–483 (2006).
Takayama, S. & Reed, J.C. Protein interaction cloning by far-Western screening of lambda-phage cDNA expression libraries. Methods Mol. Biol. 69, 171–184 (1997).
Pless, O. et al. G9a-mediated lysine methylation alters the function of CCAAT/enhancer-binding protein-beta. J. Biol. Chem. 283, 26357–26363 (2008).
Querfurth, E. et al. Antagonism between C/EBPbeta and FOG in eosinophil lineage commitment of multipotent hematopoietic progenitors. Genes Dev. 14, 2515–2525 (2000).
Mink, S., Haenig, B. & Klempnauer, K.H. Interaction and functional collaboration of p300 and C/EBPbeta. Mol. Cell Biol. 17, 6609–6617 (1997).
Chang, C.J., Chen, Y.L. & Lee, S.C. Coactivator TIF1beta interacts with transcription factor C/EBPbeta and glucocorticoid receptor to induce alpha1-acid glycoprotein gene expression. Mol. Cell Biol. 18, 5880–5887 (1998).
Leutz, A. et al. Crosstalk between phosphorylation and multi-site arginine/lysine methylation in C/EBPs. Transcription 2, 3–8 (2011).
Schulze, W.X. & Mann, M. A novel proteomic screen for peptide-protein interactions. J. Biol. Chem. 279, 10756–10764 (2004).
Ruthenburg, A.J., Allis, C.D. & Wysocka, J. Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol. Cell 25, 15–30 (2007).
Ruthenburg, A.J. et al. Multivalent engagement of chromatin modifications by linked binding modules. Nat. Rev. Mol. Cell Biol. 8, 983–994 (2007).
Finn, R.D. et al. The Pfam protein families database. Nucleic Acids Res. 38, D211–D222 (2010).
Letunic, I., Doerks, T. & Bork, P. SMART 6: recent updates and new developments. Nucleic Acids Res. 37, D229–D232 (2009).
Waterhouse, A.M. et al. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Gould, C.M. et al. ELM: the status of the 2010 eukaryotic linear motif resource. Nucleic Acids Res. 38, D167–D180 (2010).
Bartke, T. et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 143, 470–484 (2010).
Kowenz-Leutz, E. & Leutz, A. A C/EBP beta isoform recruits the SWI/SNF complex to activate myeloid genes. Mol. Cell 4, 735–743 (1999).
Acknowledgements
We thank K.W. Friedrich and U. Englisch from LI-COR Biosciences for technical support and discussions. We thank M. Knoblich for protein purification for subsequent mass spectrometric analysis. O.P. is supported by a grant of the Deutsche Forschungsgemeinschaft (DFG) to A.L. (LE 770/4-1). E.K.-L., G.D. and A.L. are supported by institutional funds from the Helmholtz Association.
Author information
Authors and Affiliations
Contributions
O.P. and A.L. designed experiments, and O.P. and E.K.-L. performed experiments and analyzed data. G.D. performed mass spectrometric analysis and data evaluation. O.P. and A.L. prepared the manuscript. A.L. supervised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Differential interaction partners on UniPEx library, part 1 (DOC 60 kb)
Rights and permissions
About this article
Cite this article
Pless, O., Kowenz-Leutz, E., Dittmar, G. et al. A differential proteome screening system for post-translational modification–dependent transcription factor interactions. Nat Protoc 6, 359–364 (2011). https://doi.org/10.1038/nprot.2011.303
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.303
This article is cited by
-
Peptide array–based interactomics
Analytical and Bioanalytical Chemistry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.