Abstract
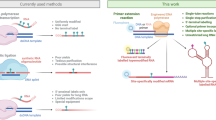
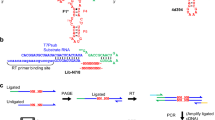
This protocol describes an efficient method for the preparation of riboswitch domains comprising up to ∼200 nt containing site-specific nucleoside modifications. The strategy is based on enzymatic ligation of chemically synthesized RNA fragments. The design of ligation sites strictly follows the criterion that all fragments comprise less than ∼50 nt. This allows the researcher to rely on custom synthesis services and to utilize the large pool of commercially available, functionalized nucleoside phosphoramidites for solid-phase RNA synthesis. Importantly, this design renders utmost flexibility to position a chemical modification (e.g., a fluorescence label) within the RNA. Selection of the appropriate ligation type (using T4 RNA or T4 DNA ligase) is subordinate to the criteria above and is detailed in the protocol. The whole concept is demonstrated for 2-aminopurine containing thiamine pyrophosphate responsive riboswitch domains that are applied in fluorescence spectroscopic folding studies. Labeled samples in 5–35 nmol quantities are obtained within 3–4 d, not including the time for fragment synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Moore, M.J. & Sharp, P.A. Site-specific modification of pre-mRNA: the 2′-hydroxyl groups at the splice sites. Science 256, 992–997 (1992).
Persson, T., Willkomm, D.K. & Hartmann, R.K. T4 RNA ligase. In Handbook of RNA Biochemistry (eds. Hartmann, R.K., Bindereif, A., Schön, A. & Westhof, E.) 53–74 (Wiley-VCH, Weinheim, Germany, 2005).
Frilander, J.M. & Turunen, J.J. RNA ligation using T4 DNA ligase. In Handbook of RNA Biochemistry (eds. Hartmann, R.K., Bindereif, A., Schön, A. & Westhof, E.) 36–52 (Wiley-VCH, Weinheim, Germany, 2005).
Winkler, W., Nahvi, A. & Breaker, R.R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–966 (2002).
Nahvi, A. et al. Genetic control by a metabolite binding mRNA. Chem. Biol. 9, 1043 (2002).
Miranda-Ríos, J., Navarro, M. & Soberón, M. A conserved RNA structure (thi box) is involved in regulation of thiamin biosynthetic gene expression in bacteria. Proc. Natl. Acad. Sci. USA 98, 9736–9741 (2001).
Mironov, A.S. et al. Sensing small molecules by nascent RNA: a mechanism to control transcription in bacteria. Cell 111, 747–756 (2002).
Barrick, J.E. & Breaker, R.R. The distributions, mechanisms, and structures of metabolite-binding riboswitches. Genome Biol. 8, R239 (2007).
Serganov, A. & Patel, D.J. Ribozymes, riboswitches and beyond: regulation of gene expression without proteins. Nat. Rev. Genet. 8, 776–790 (2007).
Edwards, T.E., Klein, D.J. & Ferré-D'Amaré, A.R. Riboswitches: small-molecule recognition by gene regulatory RNAs. Curr. Opin. Struct. Biol. 17, 273–279 (2007).
Bocobza, S. et al. Riboswitch-dependent gene regulation and its evolution in the plant kingdom. Genes Dev. 21, 2874–2879 (2007).
Grundy, F.J. & Henkin, T.M. From ribosome to riboswitch: control of gene expression in bacteria by RNA structural rearrangements. Crit. Rev. Biochem. Mol. Biol. 41, 329–338 (2006).
Cheah, M.T., Wachter, A., Sudarsan, N. & Breaker, R.R. Control of alternative RNA splicing and gene expression by eukaryotic riboswitches. Nature 447, 497–500 (2007).
Wachter, A. et al. Riboswitch control of gene expression in plants by splicing and alternative 3′ end processing of mRNAs. Plant Cell 19, 3437–3450 (2007).
Schwalbe, H., Buck, J., Fürtig, B., Noeske, J. & Wöhnert, J. Structures of RNA switches: insight into molecular recognition and tertiary structure. Angew. Chem. Int. Ed. Engl. 46, 1212–1219 (2007).
Mandal, M. & Breaker, R.R. Gene regulation by riboswitches. Nat. Rev. Mol. Cell Biol. 5, 451–463 (2004).
Nudler, E. & Mironov, A.S. The riboswitch control of bacterial metabolism. Trends Biochem. Sci. 29, 11–17 (2004).
Wickiser, J.K., Winkler, W.C., Breaker, R.R. & Crothers, D.M. The speed of RNA transcription and metabolite binding kinetics operate an FMN riboswitch. Mol. Cell 18, 49–60 (2005).
Rieder, R., Lang, K., Graber, D. & Micura, R. Folding of the adenosine deaminase A-riboswitch and implications on translational control. Chembiochem 8, 896–902 (2007).
Lang, K., Rieder, R. & Micura, R. Ligand-induced folding of the thiM TPP riboswitch investigated by a structure-based fluorescence spectroscopic approach. Nucleic Acids Res. 35, 5370–5378 (2007).
Höbartner, C. et al. Syntheses of RNAs with up to 100 nucleotides containing site-specific 2′-methylseleno labels for use in X-ray crystallography. J. Am. Chem. Soc. 127, 12035–12045 (2005).
Lemay, J.F., Penedo, J.C., Tremblay, R., Lilley, D.M. & Lafontaine, D.A. Folding of the adenine riboswitch. Chem. Biol. 13, 857–868 (2006).
Edwards, T.E. & Sigurdsson, S.T. Site-specific incorporation of nitroxide spin-labels into 2′-position of nucleic acids. Nat. Protoc. 2, 1954–1962 (2007).
Tzakos, A.G., Easton, L.E. & Lukavsky, P.J. Preparation of large oligoribonucleotides with complementary isotope-labeled segments for NMR structural studies. Nat. Protoc. 2, 2139–2147 (2007).
Zhang, Q. & Al-Hashimi, H.M. Extending the NMR spatial resolution limit for RNA by motional couplings. Nat. Methods 5, 243–245 (2008).
Bailor, M.H. et al. Characterizing the relative orientation and dynamics of RNA A-form helices using NMR residual dipolar couplings. Nat. Protoc. 2, 1536–1546 (2007).
Schiemann, O., Piton, N., Plackmeyer, J., Bode, B.E., Prisner, T.F. & Engels, J.W. Spin labeling of oligonucleotides with the nitroxide TPA and the use of PELDOR, a pulse EPR method, to measure intramolecular distances. Nat. Protoc. 2, 904–922 (2007).
Srivatsan, S.G. & Tor, Y. Synthesis and enzymatic incorporation of a fluorescent pyrimidine ribonucleotide. Nat. Protoc. 2, 1547–1555 (2007).
Chaulk, S.G. & MacMillan, A.M. Synthesis of oligo-RNAs with photocaged adenosine 2′-hydroxyls. Nat. Protoc. 2, 1052–1058 (2007).
Puffer, B., Moroder, H., Aigner, M. & Micura, R. 2′-Methylseleno modified oligoribonucleotides for X-ray crystallography synthesized by the ACE RNA solid-phase approach. Nucleic Acids Res. 36, 970–983 (2008).
Moroder, H., Kreutz, C., Lang, K., Serganov, A. & Micura, R. Synthesis, oxidation behavior, crystallization and structure of 2′-methylseleno guanosine containing RNAs. J. Am. Chem. Soc. 128, 9909–9918 (2006).
Höbartner, C. & Micura, R. Chemical synthesis of selenium-modified oligoribonucleotides and their enzymatic ligation leading to an U6 snRNA stem-loop segment. J. Am. Chem. Soc. 126, 1141–1149 (2004).
Pallan, P.S. & Egli, M. Selenium modification of nucleic acids: preparation of phosphoroselenoate derivatives for crystallographic phasing of nucleic acid structures. Nat. Protoc. 2, 640–646 (2007).
Pallan, P.S. & Egli, M. Selenium modification of nucleic acids: preparation of oligonucleotides with incorporated 2′-SeMe-uridine for crystallographic phasing of nucleic acid structures. Nat. Protoc. 2, 647–651 (2007).
Arn, E.A. & Abelson, J. RNA ligases: function, mechanism, and sequence conservation. In RNA Structure and Function (eds. Simons, R.W. & Grunberg-Manago, M.) 695–726 (CSHL Press, New York, 1998).
Stark, M.R., Pleiss, J.A., Deras, M., Scaringe, S.A. & Rader, S.D. An RNA ligase-mediated method for the efficient creation of large synthetic RNAs. RNA 12, 2014–2019 (2006).
Sherlin, L.D. et al. Chemical and enzymatic synthesis of tRNAs for high-throughput crystallization. RNA 7, 1671–1678 (2001).
Kurschat, W., Müller, J., Wombacher, R. & Helm, M. Optimizing splinted ligation of highly structured small RNAs. RNA 11, 1909–1914 (2005).
Stutz, A.Ph.D. Thesis ETH Nr. 15141. Methoden zur Ligation von Oligoribonukleotiden und ihre Anwendung bei der Totalsynthese aminoacylierter tRNAs. ETH Zurich (2003).
Sudarsan, N., Barrick, J.E. & Breaker, R.R. Metabolite-binding RNA domains are present in the genes of eukaryotes. RNA 9, 644–647 (2003).
Galagan, J.E., Henn, M.R., Ma, L.J., Cuomo, C.A. & Birren, B. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005).
Kubodera, T. et al. Thiamine-regulated gene expression of Aspergillus oryzae thiA requires splicing of the intron containing a riboswitch-like domain in the 5′-UTR. FEBS Lett. 555, 516–520 (2003).
Yamauchi, T. et al. Roles of Mg2+ in TPP-dependent riboswitch. FEBS Lett. 579, 2583–2588 (2005).
Sudarsan, N. et al. Tandem riboswitch architectures exhibit complex gene control functions. Science 314, 300–304 (2006).
Welz, R. & Breaker, R.R. Ligand binding and genetic control characteristics of tandem riboswitches in Bacillus anthracis. RNA 13, 308–317 (2007).
Serganov, A., Polonskaia, A., Phan, A.T., Breaker, R.R. & Patel, D.J. Structural basis for gene regulation by thiamine pyrophosphate-sensing riboswitch. Nature 441, 1167–1171 (2006).
Thore, S., Leibundgut, M. & Ban, N. Structure of the eukaryotic thiamine pyrophosphate riboswitch with its regulatory ligand. Science 312, 1208–1211 (2006).
Edwards, T.E. & Ferré-D'Amaré, A.R. Crystal structure of the thi-box riboswitch bound to thiamine pyrophosphate analogs reveal adaptive RNA-small molecule recognition. Structure 14, 1459–1468 (2006).
Micura, R. Small interfering RNAs and their chemical synthesis. Angew. Chem. Int. Ed. Engl. 41, 2265–2269 (2002).
Pitsch, S., Weiss, P.A., Jenny, L., Stutz, A. & Wu, X.L. Reliable chemical synthesis of oligoribonucleotides (RNA) with 2′-O-[(triisopropylsilyl)oxy]methyl(2′-O-tom)-protected phosphoramidites. Helv. Chim. Acta. 84, 3773–3795 (2001).
Höbartner, C., Ebert, M.O., Jaun, B. & Micura, R. RNA two-state conformation equilibria and the effect of nucleobase methylation. Angew. Chem. Int. Ed. Engl. 41, 605–609 (2002).
Höbartner, C., Mittendorfer, H., Breuker, K. & Micura, R. Triggering of RNA secondary structures by a functionalized nucleobase. Angew. Chem. Int. Ed. Engl. 43, 3922–3925 (2004).
Micura, R. et al. Preparation of 2′-deoxy-2′-methylseleno-modified RNA. Curr. Protoc. Nucleic Acid Chem. 1, C1.15.1–C1.15.34 (2007).
Stephenson, F. Calculations for Molecular Biology and Biotechnology: A Guide to Mathematics in the Laboratory (Academic Press, Amsterdam, 2003).
Acknowledgements
The work is supported by the Austrian Science Fund FWF (P17864) and the bm:wf (Gen-AU programme; project 'Non-coding RNAs' no. P7260-012-011) for funding.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lang, K., Micura, R. The preparation of site-specifically modified riboswitch domains as an example for enzymatic ligation of chemically synthesized RNA fragments. Nat Protoc 3, 1457–1466 (2008). https://doi.org/10.1038/nprot.2008.135
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.135
This article is cited by
-
Incorporation of isotopic, fluorescent, and heavy-atom-modified nucleotides into RNAs by position-selective labeling of RNA
Nature Protocols (2018)
-
EPR-aided approach for solution structure determination of large RNAs or protein–RNA complexes
Nature Communications (2014)
-
Enzymatic incorporation of an azide-modified UTP analog into oligoribonucleotides for post-transcriptional chemical functionalization
Nature Protocols (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.