Abstract
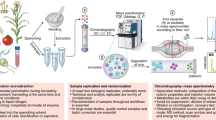
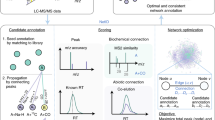
Untargeted metabolomics aims to gather information on as many metabolites as possible in biological systems by taking into account all information present in the data sets. Here we describe a detailed protocol for large-scale untargeted metabolomics of plant tissues, based on reversed phase liquid chromatography coupled to high-resolution mass spectrometry (LC-QTOF MS) of aqueous methanol extracts. Dedicated software, MetAlign, is used for automated baseline correction and alignment of all extracted mass peaks across all samples, producing detailed information on the relative abundance of thousands of mass signals representing hundreds of metabolites. Subsequent statistics and bioinformatics tools can be used to provide a detailed view on the differences and similarities between (groups of) samples or to link metabolomics data to other systems biology information, genetic markers and/or specific quality parameters. The complete procedure from metabolite extraction to assembly of a data matrix with aligned mass signal intensities takes about 6 days for 50 samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bino, R.J. et al. Potential of metabolomics as a functional genomics tool. Trends Plant. Sci. 9, 418–425 (2004).
Hall, R.D. Plant metabolomics: from holistic hope, to hype, to hot topic. New Phytol. 169, 453–468 (2006).
Jenkins, H. et al. A proposed framework for the description of plant metabolomics experiments and their results. Nat. Biotechnol. 22, 1601–1606 (2004).
Sumner, L.W., Mendes, P. & Dixon, R.A. Plant metabolomics: large-scale phytochemistry in the functional genomics era. Phytochemistry 62, 817–836 (2003).
Dixon, R.A. et al. Applications of metabolomics in agriculture. J. Agric. Food Chem. 54, 8984–8994 (2006).
Trethewey, R.N. Metabolite profiling as an aid to metabolic engineering in plants. Curr. Opin. Plant Biol. 7, 196–201 (2004).
Saito, K., Dixon, R. & Willmitzer, L. Plant Metabolomics (Springer Verlag, Heidelberg, Germany, 2006).
Vaidyanathan, S., Harrigan, G.G., Goodacre, R. (eds.) Metabolome Analyses: Strategies for Systems Biology (Springer, New York, 2005).
Van der Greef, J., Stroobant, P. & Van der Heijden, R. The role of analytical sciences in medical systems biology. Curr. Opin. Chem. Biol. 8, 559–565 (2004).
Fernie, A.R. Metabolome characterization in plant system analysis. Funct. Plant Biol. 30, 111–120 (2003).
Fiehn, O. et al. Metabolite profiling for plant functional genomics. Nat. Biotechnol. 18, 1157–1161 (2000).
Lisec, J., Schauer, N., Kopka, J., Willmitzer, L. & Fernie, A.R. Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat. Protoc. 1, 1–10 (2006).
Roessner, U. et al. Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 13, 11–29 (2001).
Roessner, U., Willmitzer, L. & Fernie, A.R. Metabolic profiling and biochemical phenotyping of plant systems. Plant Cell Rep. 21, 189–196 (2002).
Schauer, N. et al. GC-MS libraries for the rapid identification of metabolites in complex biological samples. FEBS Lett. 579, 1332–1337 (2005).
Fiehn, O. et al. Metabolite profiling for plant functional genomics. Nat. Biotechnol. 18, 1157–1161 (2000).
Aharoni, A. et al. Nontargeted metabolome analysis by use of Fourier transform ion cyclotron mass spectrometry. Omics 6, 217–234 (2002).
Hirai, M.Y. et al. Elucidation of gene-to-gene and metabolite-to-gene networks in Arabidopsis by integration of metabolomics and transcriptomics. J. Biol. Chem. 280, 25590–25595 (2005).
Overy, S.A. et al. Application of metabolite profiling to the identification of traits in a population of tomato introgression lines. J. Exp. Bot. 56, 287–296 (2005).
Goodacre, R., York, E.V., Heald, J.K. & Scott, J.M. Chemometric discrimination of unfractionated plant extracts analyzed by electrospray mass spectrometry. Phytochemistry 62, 859–863 (2003).
Jander, G. et al. Application of a high-throughput HPLC-MS/MS assay to Arabidopsis mutant screening; evidence that threonine aldolase plays a role in seed nutritional quality. Plant J. 39, 465–475 (2004).
Moco, S. et al. A liquid chromatography-mass spectrometry-based metabolome database for tomato. Plant Physiol. 141, 1205–1218 (2006).
Tolstikov, V.V., Lommen, A., Nakanishi, K., Tanaka, N. & Fiehn, O. Monolithic silica-based capillary reversed-phase liquid chromatography/electrospray mass spectrometry for plant metabolomics. Anal. Chem. 75, 6737–6740 (2003).
von Roepenack-Lahaye, E. et al. Profiling of Arabidopsis secondary metabolites by capillary liquid chromatography coupled to electrospray ionization quadrupole time-of-flight mass spectrometry. Plant Physiol. 134, 548–559 (2004).
Vorst, O. et al. A non-directed approach to the differential analysis of multiple LC-MS-derived metabolic profiles. Metabolomics 1, 169–180 (2005).
Rischer, H. et al. Gene-to-metabolite networks for terpenoid indole alkaloid biosynthesis in Catharanthus roseus cells. Proc. Natl. Acad. Sci. USA 103, 5614–5619 (2006).
Sato, S., Soga, T., Nishioka, T. & Tomita, M. Simultaneous determination of the main metabolites in rice leaves using capillary electrophoresis mass spectrometry and capillary electrophoresis diode array detection. Plant J. 40, 151–163 (2004).
Le Gall, G., Colquhoun, I.J., Davis, A.L., Collins, G.J. & Verhoeyen, M.E. Metabolite profiling of tomato (Lycopersicon esculentum) using 1H NMR spectroscopy as a tool to detect potential unintended effects following a genetic modification. J. Agric. Food Chem. 51, 2447–2456 (2003).
Ward, J.L., Harris, C., Lewis, J. & Beale, M.H. Assessment of H-1 NMR spectroscopy and multivariate analysis as a technique for metabolite fingerprinting of Arabidopsis thaliana . Phytochemistry 62, 949–957 (2003).
Huhman, D.V. & Sumner, L.W. Metabolic profiling of saponins in Medicago sativa and Medicago truncatula using HPLC coupled to an electrospray ion-trap mass spectrometer. Phytochemistry 59, 347–360 (2002).
Breitling, R., Pitt, A.R. & Barrett, M.P. Precision mapping of the metabolome. Trends Biotechnol. 24, 543–548 (2006).
Verhoeven, H.A., de Vos, C.H., Bino, R.J. & Hall, R.D. Plant metabolomics strategies based upon quadrupole time of flight mass spectrometry (QTOF-MS). in Plant Metabolomics—Biotechnology and Forestry Vol. 57, pp. 33–48 (eds. Saito, K., Dixon, R.A. & Willmitzer, L.) (Springer-Verlag, Berlin, Heidelberg, 2006).
Beekwilder, J., Jonker, H., Meesters, P., Hall, R.F., van der Meer, I.M. & de Vos, C.H.R. Antioxidants in raspberry: on-line analysis links antioxidant activity to a diversity of individual metabolites. J. Agric. Food Chem. 53, 3313–3320 (2005).
Hall, R.D., de Vos, C.H.R., Verhoeven, H.A. & Bino, R.J. Metabolomics for the assessment of functional diversity and quality traits in plants. in Metabolome Analyses-Strategies for Systems Biology (eds. Vaidyanathan, S., Harrigan, G.G. & Goodacre, R.) (Springer, New York, 2005).
Kopka, J. et al. GMD@CSB.DB: the Golm Metabolome Database. Bioinformatics 21, 1635–1638 (2005).
Tolstikov, V.V. & Fiehn, O. Analysis of highly polar compounds of plant origin: combination of hydrophilic interaction chromatography and electrospray ion mass trap spectrometry. Anal. Biochem. 301, 298–307 (2002).
Peterman, S.M., Duczak, N., Kalgutkar, A.S., Lame, M.E. & Soglia, J.R. Application of a linear ion trap/orbitrap mass spectrometer in metabolite characterization studies: examination of the human liver microsomal metabolism of the non-tricyclic anti-depressant nefazodone using data-dependent accurate mass measurements. J. Am. Soc. Mass Spectrom. 17, 363–375 (2006).
Exarchou, V., Godejohann, M., van Beek, T.A., Gerothanassis, I.P. & Vervoort, J. LC-UV-solid-phase extraction-NMR-MS combined with a cryogenic flow probe and its application to the identification of compounds present in Greek oregano. Anal. Chem. 75, 6288–6294 (2003).
Wilson, I.D. & Brinkman, U.A.T. Hyphenation and hypernation––the practice and prospects of multiple hyphenation. J. Chromatogr. A 1000, 325–356 (2003).
Wolfender, J.L., Ndjoko, K. & Hostettmann, K. Liquid chromatography with ultraviolet absorbance-mass spectrometric detection and with nuclear magnetic resonance spectroscopy: a powerful combination for the on-line structural investigation of plant metabolites. J. Chromatogr. A 1000, 437–455 (2003).
Nordström, A., O'Maille, G., Qin, C. & Siuzdak, G. Nonlinear data alignment for UPLC-MS and HPLC-MS based metabolomics: quantitative analysis of endogenous and exogenous metabolites in human serum. Anal. Chem. 78, 3289–3295 (2006).
Laaksonen, R. et al. A systems biology strategy reveals biological pathways and plasma biomarker candidates for potentially toxic statin-induced changes in muscle. PLoS ONE e97 (2006).
Keurentjes, J.J.B. et al. The genetics of plant metabolism. Nat. Genet. 38, 842–849 (2006).
Bino, R.J. et al. The light-hyperresponsive high pigment-2dg mutation of tomato: alterations in the fruit metabolome. New Phytol. 166, 427–438 (2005).
Smith, C.A., Want, E.J., O'Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Katajamaa, M. & Oresic, M. Processing software for differential analysis of LC/MS profile data. BMC Bioinformatics 6, 179.1–179.12 (2005).
Idborg, H., Zamani, L., Edlund, P., Schuppe-Koistinen, I. & Jacobsson, S.P. Metabolic fingerprinting of rat urine by LC/MS. Part 2. Data pretreatment methods for handling of complex data. J. Chromatogr. B 828, 14–20 (2005).
Fu, J., Swertz, M.A., Keurentjes, J.J.B. & Jansen, R.C. MetaNetwork: a computational protocol for the genetic study of metabolic networks. Nat. Protoc. (in the press) DOI: 10.1038/nprot.2007.96 (2007).
Tikunov, Y. et al. A novel approach for nontargeted data analysis for metabolomics. Large-scale profiling of tomato fruit volatiles. Plant Physiol. 139, 1125–1137 (2005).
Wolff, J.C., Eckers, C., Sage, A.B., Giles, K. & Bateman, R. Accurate mass liquid chromatography/mass spectrometry on quadrupole orthogonal acceleration time-of-flight mass analyzers using switching between separate sample and reference sprays. 2. Applications using the dual-electrospray ion source. Anal. Chem. 73, 2605–2612 (2001).
Acknowledgements
The preparation of this paper and the work described herein was made possible through funding from the Centre for BioSystems Genomics (which is part of the Netherlands Genomics Initiative and The Netherlands Organisation for Scientific Research), Plant Research International (PRI) and the EU project META-PHOR (Food-CT-2006-03622). We thank Harry Jonker and Bert Schipper (PRI) and Jeroen Jansen (NIOO, Heteren, The Netherlands) for their excellent help in sample preparation and LC-PDA-QTOF MS analyses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
De Vos, R., Moco, S., Lommen, A. et al. Untargeted large-scale plant metabolomics using liquid chromatography coupled to mass spectrometry. Nat Protoc 2, 778–791 (2007). https://doi.org/10.1038/nprot.2007.95
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.95
This article is cited by
-
Metabolomics analysis reveals the metabolite profiles of Rheum tanguticum grown under different altitudinal gradients
BMC Plant Biology (2024)
-
Changes in secondary metabolites of grape skins in response to different postharvest dehydration temperatures as evaluated by UPLC-Q-TOF-MS
Journal of Food Measurement and Characterization (2024)
-
Factors that influence the extraction methods of terpenes from natural sources
Chemical Papers (2024)
-
A combination of conserved and diverged responses underlies Theobroma cacao’s defense response to Phytophthora palmivora
BMC Biology (2024)
-
The multifaceted roles of Arbuscular Mycorrhizal Fungi in peanut responses to salt, drought, and cold stress
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.