Abstract
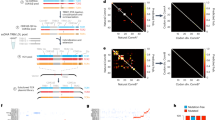
The confident discrimination of nucleic acids that share a high degree of sequence identity is the major obstacle for the widespread applicability of multiplex DNA-based techniques. This diagnostic uncertainty originates in the insufficient specificity of hybridization, allowing cross-hybridization between unwanted probe–target combinations. Starting from a random mixture of oligonucleotides, we describe a protocol to selectively amplify the probes that bind to the target but not to the similar, unintended targets. The procedure involves five forward hybridizations to generate pools of probes with significant affinity, but not necessarily specificity, for the target. Specificity is then achieved during subtractive hybridization steps, where only probes having differential diagnostic performance are retained. Iterative hybridizations, cloning, sequencing and testing of the performance of selected probes can all be fully automated. Eight weeks are required for the full completion of a project composed of 40 probe–target pairs, even when targets share as much as 87% of sequence identity. While alternative, computer-assisted, rational oligonucleotide design may produce an uncertain outcome, the present protocol generates robust and specific probes suitable for a variety of multiplex, nucleic acid-based detection/typing platforms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dean, F.B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl. Acad. Sci. USA 99, 5261–5266 (2002).
Makrigiorgos, G.M., Chakrabarti, S., Zhang, Y., Kaur, M. & Price, B.D. A PCR-based amplification method retaining the quantitative difference between two complex genomes. Nat. Biotechnol. 20, 936–939 (2002).
Bertilsson, S., Cavanaugh, C.M. & Polz, M.F. Sequencing-independent method to generate oligonucleotide probes targeting a variable region in bacterial 16S rRNA by PCR with detachable primers. Appl. Environ. Microbiol. 67, 6077–6086 (2002).
Regnault, B., Grimont, F. & Grimont, P.A. Universal ribotyping method using a chemically labelled oligonucleotide probe mixture. Res. Microbiol. 148, 649–659 (1997).
Perez-Luz, S., Adela Yanez, M. & Catalan, V. Identification of waterborne bacteria by the analysis of 16S–23S rRNA intergenic spacer region. J. Appl. Microbiol. 97, 191–204 (2004).
Mohammadi, T., Pietersz, R.N., Vandenbroucke-Grauls, C.M., Savelkoul, P.H. & Reesink, H.W. Detection of bacteria in platelet concentrates: comparison of broad-range real-time 16S rDNA polymerase chain reaction and automated culturing. Transfusion 45, 731–736 (2005).
Nadkarni, M.A., Martin, F.E., Jacques, N.A. & Hunter, N. Determination of bacterial load by real-time PCR using a broad-range (universal) probe and primers set. Microbiology 148 (Pt. 1): 257–266 (2002).
Santos, S. & Ochman, H. Identification and phylogenetic sorting of bacterial lineages with universally conserved genes and proteins. Environ. Microbiol. 6, 754–759 (2004).
Watanabe, K., Kodama, Y. & Harayama, S. Design and evaluation of PCR primers to amplify bacterial 16S ribosomal DNA fragments used for community fingerprinting. J. Microbiol. Methods. 44, 253–262 (2001).
Okada, M., Ogawa, T., Kubonoya, H., Yoshizumi, H. & Shinozaki, K. Detection and sequence-based typing of human adenoviruses using sensitive universal primer sets for the hexon gene. Arch. Virol. 152, 1–9 (2007).
Schaefer, S., Glebe, D., Wend, U.C., Oyunbileg, J. & Gerlich, W.H. Universal primers for real-time amplification of DNA from all known Orthohepadnavirus species. J. Clin. Virol. 27, 30–37 (2003).
Hoffmann, E., Stech, J., Guan, Y., Webster, R.G. & Perez, D.R. Universal primer set for the full-length amplification of all influenza A viruses. Arch. Virol. 146, 2275–2289 (2001).
Qu, W. et al. PCR detection of human papillomavirus: comparison between MY09/MY11 and GP5+/GP6+ primer systems. J. Clin. Microbiol. 35, 1304–1310 (1997).
Heinze, B. A database of PCR primers for the chloroplast genomes of higher plants. Plant Methods 3, 4 (2007).
Pineau, P. et al. A universal primer set for PCR amplification of nuclear histone H4 genes from all animal species. Mol. Biol. Evol. 22, 582–588 (2005).
Zietkiewicz, E., Rafalski, A. & Labuda, D. Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20, 176–183 (1994).
Southern, E.M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 98, 503–517 (1975).
Drmanac, R., Labat, I., Brukner, I. & Crkvenjakov, R. Sequencing of megabase plus DNA by hybridization: theory of the method. Genomics 4, 114–128 (1989).
Moiseev, L., Unlu, M.S., Swan, A.K., Goldberg, B.B. & Cantor, C.R. DNA conformation on surfaces measured by fluorescence self-interference. Proc. Natl. Acad. Sci. USA. 103, 2623–2628 (2006).
Gharizadeh, B. et al. Viral and microbial genotyping by a combination of multiplex competitive hybridization and specific extension followed by hybridization to generic tag arrays. Nucleic Acids Res. 31, e146 (2003).
Brukner, I., Tremblay, G.A. & Paquin, B. Generation of amplifiable genome-specific oligonucleotide probes and libraries. Biotechniques 33, 874–876 878, 880 passim (2002).
Pozhitkov, A. et al. Tests of rRNA hybridization to microarrays suggest that hybridization characteristics of oligonucleotide probes for species discrimination cannot be predicted. Nucleic Acids Res. 34, e66 (2006).
Brukner, I., El-Ramahi, R., Gorska-Flipot, I., Krajinovic, M. & Labuda, D. An in vitro selection scheme for oligonucleotide probes to discriminate between closely related DNA sequences. Nucleic Acid Res. 35, e66 (2007).
Brukner, I. et al. Hybridization assay performed at ambient temperature for typing high-risk human papillomaviruses. J. Clin. Virol. 39, 113–118 (2007).
Robertson, D.L. & Joyce, G.F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature 344, 467–468 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Evans, M.F., Adamson, C.S., Simmons-Arnold, L. & Cooper, K. Touchdown general primer (GP5+/GP6+) PCR and optimized sample DNA concentration support the sensitive detection of human papillomavirus. BMC Clin. Pathol. 5, 10 (2005).
Acknowledgements
Work reviewed in this article is supported by the Canadian Institutes of Health Research (NTA-71859) and Research Center of Sainte-Justine Hospital. M.K. is a scholar of the Fonds de la Recherche en Santeé du Québec. We thank our colleagues Dr. Izabella Gorska, Jacob Sawicki (Acrobat Illustrator wizard) and Razan El-Ramahi for the contribution to the HPV project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brukner, I., Krajinovic, M., Dascal, A. et al. A protocol for the in vitro selection of specific oligonucleotide probes for high-resolution DNA typing. Nat Protoc 2, 2807–2814 (2007). https://doi.org/10.1038/nprot.2007.398
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.398
This article is cited by
-
Oncogenic HPV infection resembling pseudocondyloma of vulvae: a descriptive study of 31 cases
Archives of Gynecology and Obstetrics (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.