Abstract
DNA sequencing has markedly changed the nature of biomedical research, identifying millions of polymorphisms along the human genome that now require further analysis to study the genetic basis of human diseases. Among the DNA-sequencing platforms available, Pyrosequencing has become a useful tool for medium-throughput single nucleotide polymorphism (SNP) genotyping, mutation detection, copy-number studies and DNA methylation analysis. Its 96-well genotyping format allows reliable results to be obtained at reasonable costs in a few minutes. However, a specific biotinylated primer is usually required for each SNP under study to allow the capture of single-stranded DNA template for the Pyrosequencing assay. Here, we present an alternative to the standard labeling of PCR products for analysis by Pyrosequencing that circumvents the requirement of specific biotinylated primers for each SNP of interest. This protocol uses a single biotinylated primer that is simultaneously incorporated into all M13-tagged PCR products during the amplification reaction. The protocol covers all steps from the PCR amplification and capture of single-stranded template, its preparation, and the Pyrosequencing assay itself. Once the correct primer stoichiometry has been determined, the assay takes around 2 h for PCR amplification, followed by 15–20 min (per plate) to obtain the genotypes.
Similar content being viewed by others
Introduction
During the past decade, a large number of single SNPs have been annotated in public and private databases. Owing to the tremendous amount of information available, low-cost genotyping methods have been developed to provide researchers with the ability to read genome information. The main applied objective is to determine the role of genetic variations in the development of human diseases and to predict drug inefficiency among patients. As each population usually harbors a different genetic background, structural geneticists face enormous work during the following decades. Many different genotyping technologies have been developed, based on mass spectrometry (i.e., MALDI-TOF1), fluorescence (i.e., FRET-based methods2,3) or luminescence (i.e., Pyrosequencing4). Selection of the most appropriate genotyping platform will depend on the specific requirements of each project, taking into account the relative sensitivities and throughputs of the available methods1,2,3,4.
Since its first publication in 1998, Pyrosequencing has significantly changed structural genomics4. Pyrosequencing is a real-time sequencing technique based on the detection of released pyrophosphate during DNA elongation. This procedure starts once the PCR products are converted into single-stranded DNA (ssDNA) fragments and one strand is isolated (usually through labeling with biotin) for use as template in the Pyrosequencing reaction. Then, a sequencing primer is incorporated into the reaction and a specific nucleotide is dispensed. When pyrophosphate molecules are released upon incorporation of the nucleotide in the elongating strand, ATP-sulfurylase converts it to ATP, thus providing the energy for luciferase to generate visible light peaks that can be recorded using an LCC camera5. The molecular basis of the Pyrosequencing reaction is illustrated in Figure 1. The presence of Apyrase in the reaction eliminates the excess of nucleotides, leaving the media prepared for the addition of a second nucleotide. With this technology, researchers are capable of reading up to 100 nucleotides of a DNA fragment in a quantitative manner. Recently, the reading and assembly of tens of thousands of simultaneous Pyrosequencing reactions has been reported to sequence up to 20 Mb on every single run6. Given the linear correlation between the nucleotides incorporated and the light emitted, Pyrosequencing can also be used to calculate allele frequencies in DNA pools7, and has been extremely helpful in performing allele quantification assays, analyzing regions undergoing loss of heterozygosity8,9.
DNA polymerase elongates the single-stranded template DNA from the sequencing primer, which anneals near the mutation to be genotyped. dNTPs are sequentially added, and their incorporation results in the release of pyrophosphate molecules that are converted to ATP by the sulfurylase. These ATP molecules are converted into a detectable light by the luciferase, while the remaining nucleotides are degraded by the apyrase. The relationship between the dNTP incorporation and the released light is proportional, which allows quantitative genotyping.
Pyrosequencing has been compared with other SNP genotyping platforms such as single-stranded conformation assays or 5′ nuclease (Taqman) assays, obtaining reliable results at reasonable costs per genotype10,11,12. However, to generate the ssDNA, one of each primer pair must be labeled on its 5′ site (typically with biotin). Depending on the number of SNPs under study and the sample size, the costs per assay may significantly increase. Some authors have previously proposed the use of an indirect labeling protocol13,14. For instance, Pacey-Miller and Henry14 developed a protocol using a primer with an 11-base GC tag; this is used in a first PCR round and then is targeted in a second PCR employing a GC tag-specific biotinylated primer. The inefficiency of these three-primer PCRs, however, necessitated the use of a post-PCR-labeling step, reducing the flexibility of the technique13. Other examples showing different alternatives to generate single-stranded PCR products in a feasible fashion also exist, using different tags15,16,17,18. For example, Groth et al.16 developed a method that uses a ligation of the amplicon into a plasmid vector, thereby facilitating subsequent PCR using a universal vector-specific biotinylated primer. However, no systematic comparison has been performed yet using the different indirect labeling protocols available; thus, no rigorous conclusion regarding the best tag of choice can currently be drawn. In our experience, the M13 sequence has been traditionally used in our laboratory as a 5′ primer tag for direct sequencing and enzymatic mutation detection. For this reason, the M13 sequence was our tag of choice. However, it must be stated that if the region containing the SNP of interest exhibits any structural incompatibility with the M13 sequence, researchers should consider either changing the tag or ordering the directly labeled primer. Here we describe a simple alternative to generate labeled PCR products using M13-tagged primers and a unique M13 biotinylated primer that is incorporated during the PCR, which was originally presented in ref. 19. We have found that setting up the primer stoichiometry can be easily performed, which may be useful for some researchers, as the implementation of this protocol makes unnecessary the use of several biotinylated primers19.
Experimental design
Primer design.
The goal of this procedure is to allow the incorporation of an M13 biotinylated primer in the PCR fragment while the amplification takes place. This is accomplished by adding the M13 sequence (5′-CGCCAGGGTTTTCCCAGTCACGAC-3′) at the 5′ site of one of the amplification primers. A biotinylated M13 primer is also then included in the reaction, which will match its complementary sequence in the amplification primer and so will be incorporated into the PCR product during the amplification reaction. A diagrammatic representation of the entire procedure can be seen in Figure 2.
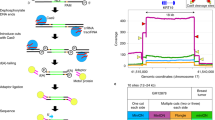
(a) The PCR involves three primers: two amplification primers (forward and reverse) that are specific to the sequence of interest, one of which (in this case the forward primer) is tagged with the M13 sequence and a biotinylated M13 primer. (b) During the PCR, the M13 tail is incorporated into the amplicon sequence. Consequently, the biotinylated M13 primer can anneal to its complementary sequence and be extended, thus labeling the amplicon with biotin. (c) The ratio of labeled and unlabeled amplicons will mainly depend on (i) the stoichiometry of the reaction, (ii) the annealing temperature and (iii) the salt concentration. (d) Biotin-labeled amplicons are captured by binding to streptavidin-coated Sepharose beads, and DNA is denatured to produce ssDNA template for the Pyrosequencing assay. (e) Finally, the ssDNA is released and is combined with the sequencing primer, which is extended during the Pyrosequencing reaction to provide the sequence of the template DNA; the sequential incorporation of every nucleotide is converted to light, which is detected by the PSQ 96 instrument, enabling the sequence of the template strand to be determined and SNPs to be genotyped. (f) Pyrosequencing data are produced in the form of pyrograms, and genotypes are assigned by Pyrosequencing software, as shown here for the three possible genotypes of the human CAV-1 rs3807990 T/C SNP.
Optimization of primer stoichiometry.
It must be stated that the addition of a third primer (in our case, the M13 biotinylated one) in a PCR may interfere with the PCR. Therefore, the key to this process is to determine the adequate primer ratios, to obtain sufficient labeled amplicon yield. This is performed in two steps. First, it is necessary to set up and optimize the PCR without the biotinylated primer, during which we fix the concentration of the amplification primers (between 0.2 and 0.5 mM each). This greatly simplifies the setting up of the labeling reaction. Once these parameters are defined, four decreasing amounts of the biotinylated M13 primer are tested. The right ratio will therefore be empirically determined, and this will depend on the PCR annealing temperature and the Mg2+ content. Thus, with an adjusted primer stoichiometry, we can standardize the labeling of every amplicon with a unique biotinylated universal primer.
Materials
Reagents
-
Biotinylated M13 primer (biotin-CGCCAGGGTTTTCCCAGTCACGAC-3′)
-
Forward and reverse PCR-amplification primers and sequencing primers (Sigma-Genosis)
Critical
Note that one of the amplification primers must be designed with the M13 sequence at the 5′ end (see Experimental design). Remember that the M13-tagged primer will define the strand complementary to the sequencing primer.
-
Streptavidin Sepharose HP (Amersham Biosciences, cat. no. 17-5113-01)
-
Binding buffer (10 mM Tris-HCl, 2 M NaCl, 1 mM EDTA, 0.1% (vol/vol) Tween 20, pH 7.6; Biotage, cat. no. 40-0033)
-
Annealing buffer (20 mM Tris-acetate, 2 mM Mg-acetate, pH 7.6; Biotage, cat. no. 40-0036)
-
Denaturation solution (0.2 M NaOH; Panreac, cat. no. 1416871210)
-
Washing buffer (10 mM Tris-acetate, pH 7.6; Amresco, cat. no. J838)
-
High-purity water (Milli-Q)
-
Ethanol 70% (Panreac, cat. no. 1410851214)
-
PSQ reagent kit (Isogen, cat. no. 40-0044)
-
PCR buffer (Roche, cat. no. 3560597001)
-
Mg2+ (Roche, cat. no. 3560597001)
-
dNTPs (Invitrogen, cat. no. R72501)
-
Taq polymerase (Roche, cat. no. 3560597001)
-
Agarose (Ecogen, cat. no. AG0220)
-
Ethidium bromide (Sigma, cat. no. E1510)
-
Gel running buffer (Applichem, cat. no. A14161000)
-
DNA loading dye (Bioline, cat. no. 37045)
Equipment
-
PSQ 96 instrument (Biotage)
-
Vacuum Prep Tool (Biotage, cat. no. 60-0112)
-
Vacuum source (water jet or vacuum pump)
-
PSQ 96 plate (Biotage, cat. no. 40-0002)
-
PCR 96-well plate (Cultek Spain, cat. no. 872619)
-
Orbital shaker for microtiter plates
-
Heating block (80 °C)
Procedure
PCR design and optimization of primer stoichiometry
-
1
Design PCR primers to amplify the sequence region containing the mutation of interest according to the information given by specialized primer software (i.e., UCSC Genome Bioinformatics: http://www.genome.ucsc.edu/cgi-bin/hgPcr), with a recommended amplicon length between 150 and 200 bp.
-
2
Once each primer pair is designed, submit the amplicon to the Biotage web tool (http://techsupport.pyrosequencing.com/) to select an appropriate sequencing primer, following the manufacturer's instructions.
Critical Step
Remember to include the M13 tail in the 5′ site of one of the primers. Remember that the M13-tagged primer will define the strand complementary to the sequencing primer.
-
3
To initially optimize the PCR amplification of the target DNA (without the addition of the universal biotinylated M13 primer to label the product), prepare a PCR mix in a final volume of 20–40 μl, according to standard PCR protocols, as follows: 25 ng of genomic DNA, forward and reverse amplification primers at final concentrations of 0.2–0.5 mM each, 1 × PCR buffer, 1.25 mM Mg2+, 0.2 mM of dNTPs and 1 U of Taq polymerase.
Critical Step
Add only the forward and reverse primers at this stage. Leave the biotinylated M13 primer until Step 6.
-
4
Run the PCR according to a conventional amplification protocol, as follows: initial 5 min denaturation step followed by 35 amplification cycles of 95 °C for 30 s, 55–62 °C (use annealing temperature as optimal for each specific amplification primer set) for 30 s and 72 °C for 30 s. End the program with a final elongation of 5 min at 72 °C.
Pause point
PCR products can be stored at −20 °C for a month.
-
5
Analyze the entire volume of the PCR product in an agarose gel stained with ethidium bromide (or use another standard DNA visualization method) to check that a single product of the expected size is obtained.
Critical Step
If nonspecific amplicons also result from the PCR, further optimization of PCR conditions is required, by testing either different primer-annealing temperatures or Mg2+ concentrations.
-
6
Once the PCR amplification of the target DNA is optimized, empirically determine the correct primer stoichiometry with which to label the PCR product with biotin during amplification using the universal biotinylated M13 primer, by preparing four PCR tubes according to Step 3. Also include universal biotinylated M13 primer at the following decreasing concentrations, one concentration per tube: 0.5, 0.375, 0.25, 0.125 mM.
Critical Step
Keep identical final volume and salt concentrations.
-
7
Run the PCR as in Step 4.
Pause point
Biotinylated PCR products can be stored at 4 °C overnight or at −20 °C for a week.
-
8
To assess which concentration of universal biotinylated M13 primer is optimal for use with a particular amplification primer set, use the PCR products obtained in Step 7 as template for Pyrosequencing assays, following the procedure detailed below (Steps 10–29). The optimal concentration of universal biotinylated M13 primer will result in the PSQ software directly giving a genotype without any further analysis.
Critical Step
Steps 1–8 need to be undertaken only the first time a new primer set is used in order to optimize the conditions for PCR product amplification and labeling. Once optimal PCR components and conditions have been determined using these steps, the procedure can just be followed from Step 9 onwards to use the optimized assay on experimental samples.
PCR amplification of Pyrosequencing template
-
9
Once the optimal PCR components and conditions for an assay have been established (using Steps 1–8), the PCR amplification of Pyrosequencing template is carried out by following just Steps 3 (including also biotinylated M13 primer at the optimal concentration as determined by Steps 6–8) and 4. It is highly recommended to include an additional sample without DNA template that will act as a negative control.
Sequencing primer annealing plate preparation
-
10
Prepare a PSQ 96 plate by adding 0.4 μM sequencing primer in 40 μl annealing buffer per well, one for each sample to be tested.
Immobilization of PCR product to streptavidin beads
-
11
Before starting, let the buffers reach room temperature (22–24°C).
-
12
Transfer the required total amount of streptavidin Sepharose beads (3 μl per sample) to a tube, together with 37 μl of binding buffer (per sample).
-
13
Add high-purity water to the PCR samples to a final volume of 40 μl and transfer each sample to a well of a 96-well PCR plate (if PCRs were prepared in separate tubes).
-
14
Add an equal volume of the mixture of binding buffer and streptavidin Sepharose beads (from Step 12) to each PCR sample from Step 13 (final volume of 80 μl per well). Seal the plate.
-
15
To allow biotinylated PCR product capture on streptavidin-coated beads, shake the mixture constantly at 800–1,000 r.p.m. for 10 min at room temperature.
Critical Step
Weak mixing may lead to bead sedimentation and reduce PCR product binding.
Preparation of single-stranded Pyrosequencing template DNA
-
16
Place the Vacuum Prep Tool in a water trough. Connect the vacuum pump and switch on the vacuum and let the water flush through the filters for 20 s.
-
17
Remove the PCR plate from the shaker and capture the beads containing immobilized templates on the filter probes by lowering the Vacuum Prep Tool into the PCR plate.
Critical Step
Beads sediment quickly, so this step must be carried out immediately after the plate is removed from the shaker.
-
18
Move the Vacuum Prep Tool (with the captured beads) to the trough containing 70% ethanol and let the solution flush through the filters for 5 s.
-
19
Move the Vacuum Prep Tool to the trough containing denaturation solution and let it flush through the filters for 5 s.
-
20
Move the Vacuum Prep Tool to the trough containing washing buffer and let it flush through the filters for 5 s.
-
21
Hold the Vacuum Prep Tool at a 90° angle and let it dry for a few seconds. Return it to the horizontal position.
-
22
Switch off the vacuum pump, then place the Vacuum Prep Tool into the wells of the PSQ 96 plate (prepared in Step 10).
Critical Step
Make sure to switch off the pump before lowering the Vacuum Prep Tool into the wells, because if the pump is still on, it will aspirate the annealing buffer and the sequencing primer.
-
23
Shake the Vacuum Prep Tool and let the filters rest for a few seconds on the bottom of the wells while the beads are released.
-
24
Move the Vacuum Prep Tool to the water trough to wash the filters.
Pyrosequencing reaction
-
25
Heat the plate containing the ssDNA template (from Step 23) at 80 °C for 2 min to denature the template and let the sequencing primer anneal during cooling to room temperature.
-
26
Select the instrument parameters according to the manufacturer's instructions and load the cartridge with the determined amount of PSQ reagents.
-
27
Place the plate in the PSQ 96 instrument and insert the cartridge carefully.
-
28
Start the run.
-
29
Pyrograms are produced and genotypes are assigned to each sample at the end of the run by the Pyrosequencing software, which compares the obtained patterns of nucleotide addition to those expected for each of the theoretical genotypes for a certain target sequence containing the mutation of interest (known as the 'sequence to analyze'), which is entered into the software to set up the Pyrosequencing assay (see Fig. 3).
Figure 3: Comparison between the expected and obtained pyrograms to genotype two independent mutations. Theoretical pyrograms for the three possible genotypes at each site are shown on the left-hand side. Pyrograms obtained by Pyrosequencing analysis of samples representing each possible genotype are shown on the right-hand side. Key nucleotides have been delimited. (a) Genotyping human CYP19A1 rs11575899 (a TCT insertion) was performed using 0.125 mM of the biotinylated M13 primer. (b) Genotyping human CAPN10 rs2975760 (a T/C SNP) was performed using 0.5 mM of the biotinylated M13 primer.
Troubleshooting
Troubleshooting advice can be found in Table 1.
Timing
Unlabeled PCR setup and analysis (Steps 3–5): 3–6 h
M13 biotin-labeling PCR (Steps 6 and 7): 2.5 h
Preparing samples for Pyrosequencing (Steps 10–27): 20 min per plate
Pyrosequencing reaction (Steps 28 and 29): 15 min per plate
Anticipated results
We have found that typical results are obtained using 0.125–0.25 μM of the biotinylated M13 primer for those PCRs with annealing temperatures between 58 and 64 °C. However, the effect of the different primer stoichiometries on the pyrograms produced must be analyzed carefully to determine the optimal primer stoichiometry. If the annealing temperature is significantly higher (i.e., 68 °C), an adequate labeling will probably require 0.5 μM of biotinylated M13 primer19. In Figure 3, we show the pyrograms obtained from two independent mutations (rs11575899 and rs2975760), using indirect M13 tags and the indirect labeling process.
References
Tang, K. et al. Matrix-assisted laser desorption/ionization mass spectrometry of immobilized duplex DNA probes. Nucleic Acids Res. 23, 3126–3131 (1995).
Hardenbol, P. et al. Multiplexed genotyping with sequence-tagged molecular inversion probes. Nat. Biotechnol. 21, 673–678 (2003).
Tyagi, S., Bratu, D.P. & Kramer, F.R. Multicolor molecular beacons for allele discrimination. Nat. Biotechnol. 16, 49–53 (1998).
Ronaghi, M., Uhlen, M. & Nyren, P. A sequencing method based on real-time pyrophosphate. Science 281, 363–365 (1998).
Ronaghi, M. Pyrosequencing sheds light on DNA sequencing. Genome Res. 11, 3–11 (2001).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Lavebratt, C. & Sengul, S. Single nucleotide polymorphism (SNP) allele frequency estimation in DNA pools using Pyrosequencing. Nat. Protoc. 6, 2573–2582 (2006).
Pascual, M.H. et al. Exploring allelic imbalance within paraffin-embedded tumor biopsies using pyrosequencing technology. Clin. Chem. Lab. Med. 44, 1076–1081 (2006).
Asplund, A. et al. PTCH codon 1315 polymorphism and risk for nonmelanoma skin cancer. Br. J. Dermatol. 152, 868–873 (2005).
Sivertsson, A., Platz, A., Hansson, J. & Lundeberg, J. Pyrosequencing as an alternative to single-strand conformation polymorphism analysis for detection of N-ras mutations in human melanoma metastases. Clin. Chem. 48, 2164–2170 (2002).
Aquilante, C.L., Lobmeyer, M.T., Langaee, T.Y. & Johnson, J.A. Comparison of cytochrome P450 2C9 genotyping methods and implications for the clinical laboratory. Pharmacotherapy 24, 720–726 (2004).
Nordfors, L. et al. Large-scale genotyping of single nucleotide polymorphisms by Pyrosequencing trade mark and validation against the 5′ nuclease (Taqman) assay. Hum. Mutat. 19, 395–401 (2002).
Fakhrai-Rad, H., Pourmand, N. & Ronaghi, M. Pyrosequencing: an accurate detection platform for single nucleotide polymorphisms. Hum. Mutat. 19, 479–485 (2002).
Pacey-Miller, T. & Henry, R. Single-nucleotide polymorphism detection in plants using a single-stranded pyrosequencing protocol with a universal biotinylated primer. Anal. Biochem. 317, 166–170 (2003).
Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 18, 233–234 (2000).
Groth, M. et al. Method for preparing single-stranded DNA templates for Pyrosequencing using vector ligation and universal biotinylated primers. Anal. Biochem. 356, 194–201 (2006).
Guo, D.C. & Milewicz, D.M. Methodology for using a universal primer to label amplified DNA segments for molecular analysis. Biotechnol. Lett. 25, 2079–2083 (2003).
Aydin, A., Toliat, M.R., Bahring, S., Becker, C. & Nurnberg, P. New universal primers facilitate Pyrosequencing. Electrophoresis 27, 394–397 (2006).
Royo, J.L. et al. Pyrosequencing protocol requiring a unique biotinylated primer. Clin. Chem. Lab. Med. 44, 435–341 (2006).
Acknowledgements
We thank Maria del Carmen Rivero and Ana Salinas for their excellent technical support. This work was founded by Ministerio de Educación y Ciencias (Spanish Ministry of Education and Sciences) PROFIT 010000-2004-69 and the Fundación de Investigación Biomédica Mutua Madrileña Automovilista (Ref. 2004/20).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Royo, J., Hidalgo, M. & Ruiz, A. Pyrosequencing protocol using a universal biotinylated primer for mutation detection and SNP genotyping. Nat Protoc 2, 1734–1739 (2007). https://doi.org/10.1038/nprot.2007.244
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.244
This article is cited by
-
Chronic oxytocin administration stimulates the oxytocinergic system in children with autism
Nature Communications (2024)
-
Endogenous oxytocin levels in children with autism: Associations with cortisol levels and oxytocin receptor gene methylation
Translational Psychiatry (2023)
-
Perceived Parental Support and Psychological Control, DNA Methylation, and Loneliness: Longitudinal Associations Across Early Adolescence
Journal of Youth and Adolescence (2023)
-
High frequency of benzimidazole resistance alleles in trichostrongyloids from Austrian sheep flocks in an alpine transhumance management system
BMC Veterinary Research (2020)
-
DNA methylation signatures of Prostate Cancer in peripheral T-cells
BMC Cancer (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.