Abstract
DNA sensing at a single nucleotide resolution is achieved using a hairpin-shaped, unmodified (unlabeled) RNA probe or the precursor double-stranded DNA (dsDNA) in a prokaryotic cell–free translation medium. The molecular-beacon-like probe consists of a loop region that is complementary to the target sequence and a stem composed of a ribosome-binding site (RBS) and its docking domain; the RBS is followed by the gene for a reporter protein such as luciferase or β-galactosidase. Target binding at the loop opens the hairpin to make RBS accessible by the ribosome to start translation of the reporter protein. This sensing system is signal amplifying by virtue of catalytic DNA-to-RNA transcription when using a dsDNA probe, catalytic RNA-to-protein translation, catalytic signal transduction by the enzymatic reaction of the translated reporter protein and, in the presence of RNase H, catalytic or even irreversible translation-activation of the target-probe heteroduplex. Preparation of a probe takes 1–3 d and gene sensing using the probe takes 1–3 h.
Similar content being viewed by others
Introduction
Coupling of molecular beacon strategy to cis-acting RNA-controlled protein translation
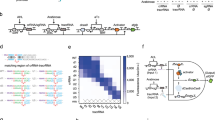
As the number of identified genes continues to grow, the need increases for rapid and simple gene sensing with high sequence selectivity1,2. Gene sensing of any type is based on selective target-probe hybridization and its output, which should be easily monitored. Molecular beacons (MBs) with a fluorescence resonance energy transfer (FRET) pair (Fig. 1a)3 utilize hybridization-induced conformational change of the probe from the light-off hairpin structure to a light-on open form. Much recent attention has also been paid to catalytic gene sensing4,5,6,7,8,9,10,11,12,13 through coupling to a signal-amplifying enzymatic4,5,6,7,8, chemical9,10, electrochemical11 or magnetochemical12 process. On the other hand, we reported a new strategy with MB-mRNA systems for selective genotyping13. This is based on naturally occurring14 or engineered15 hairpin-shaped or MB-like RNAs capable of riboregulator-dependent conformational change to control the translation activity, so that the sensing of the target oligodeoxynucleotide (ODN) as a regulator can be performed using a genetically encodable unmodified RNA as a probe in a typical prokaryotic translation system.
Design of MB-mRNA probes
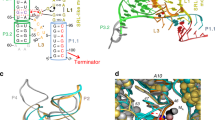
The artificial translation regulation system (MB-mRNA; Fig. 1b) is composed of a cis-acting MB-like RNA structure, wherein the loop region (blue) is complementary to the target (green) and the stem consists of sequences for a ribosome-binding site (RBS; red) and the anti-RBS or RBS-docking domain complementary thereto (pink)13. The RBS is followed by a reporter gene sequence starting with an AUG start codon. The anti-RBS domain is preceded by another hairpin structure16. This is to endow an anti-RNase stability. Binding of the target (green) at the loop is expected to result in the opening of the MB structure, making the RBS domain accessible by the ribosome and hence initiating the translation of the reporter gene, such as luciferase.
The balance of the loop and the stem in terms of base-pair (bp) length is critical for sensitivity and selectivity. If the stem is long, the hairpin structure will be more stabilized and will be opened only by a longer target capable of more extensive target-probe hybridization. This will give a high sensitivity—that is, high target-on/target-off signal ratio—but low sequence selectivity. If, on the other hand, the stem is short, the hairpin structure will be less stabilized and easily opened even by a shorter target. This will give high sequence selectivity but a low sensitivity. Actually, we chose the commonly encountered 6-base AGGAGA sequence as RBS and set, after trial and error, an 8-bp stem and 18- or 16-bp target-probe hybridization at the loop to optimize sensitivity and selectivity.
Advantage of the method
The construction of MB-mRNA probes is apparently complex. The actual sensing takes time because it includes many steps of enzymatic conversion. The sensitivity or detection limit, 9 fmol (3.6 nM) as shown below, is not very high compared with those achieved recently by other methods17. Nevertheless, the present system has a number of unique aspects. (i) The unmodified (unlabeled) probe can be provided not only as RNA (MB-mRNA) but also in the form of much more stable and directly PCR-amplifiable double-stranded DNA (dsDNA), as the former is transcribed in situ from the latter in the cell-free translation system. (ii) The essential transcription and translation are natural processes of the cells. (iii) The sensing is multiply catalytic or signal amplifying, as all of the respective steps—that is, transcription of DNA to RNA, translation of RNA to protein and transduction of signal (chemiluminescence, for example) from the enzymatic reaction of the translated reporter protein—are catalytic. These aspects suggest that the present system is potentially applicable to in-cell gene sensing, as there are established protocols for the cellular uptake of dsDNA and expression therein of the encoded gene into protein. PCR-based methods, on the other hand, cannot be used in cells. The applicability of artificial signal-amplifying systems in cells is highly questionable. The usefulness of labeled probes in cells also remains to be elucidated as regards to the efficacy of cellular uptake, extent of toxicity and compatibility of the sensing reaction, such as FRET, with cellular environments.
Another merit of the present system is that the reporter protein can, in principle, be of any type. This is particularly important in terms of single nucleotide polymorphism (SNP) detection, since we can easily construct a set of allele-sensitive probes using a class of otherwise closely related reporter proteins that can be distinguished from each other.
Outline of protocol
Although our ultimate goal is to establish in vivo gene sensing using the present system, the scope of this protocol is to describe its in vitro performance at a quantitative level, thus providing a basis for its in-cell application on one hand and, on the other hand, a guideline for the in vitro use of RNA systems for selection/sensing of translation-controlling small molecules (riboswitches)18. The targets are ODNs (Fig. 2) copying the 18-nt (620–637) or 16-nt (620–635) sequence (green) of the HIV 1–related human CC chemokine receptor 5 (CCR5) with a mutation at position 627 (ref. 19). The actual targets with variation at the corresponding position shown in red are designated 18X and 16X (X = A, T, C or G) (Fig. 2). We also used a doubly mutated target 18TA.
Luciferase was our first choice as sensing output because of its high sensitivity, the good linearity of the related chemiluminescence assay and the ready color tuning (see below). Although this system requires luciferin as an external assay reagent, the latter is readily taken up into the cell and can thus be used for in vivo sensing20. A prototypical 18C-targeting MB-mRNA probe, designated G-RNABEST (Fig. 3), was obtained by sequential PCRs on the luciferase gene (luc) encoded in plasmid vector pBESTluc and subsequent transcription (Fig. 4). The first PCR added the RBS (red), an 18-nt target-binding site (blue) with a target-complementary G base, and an intervening spacer. The second PCR added an anti-RBS domain (pink) and an additional hairpin structure preceded by the T7 promoter region. The primer sequences used are summarized in Figure 5. For sequence-selective production of multicolor reporter proteins, we took advantage of the MultiReporter Assay System. Three plasmid vectors, pSLG-test, pSLO-test and pSLR-test, encode green-luminescing wide-type Rhagophthalmus ohbai firefly luciferase (G-luc; emission maximum, 550 nm), orange-luminescing mutated (T226N) R. ohbai firefly luciferase (O-luc; 580 nm) and red-luminescing wild-type Phrixothrix hirtus firefly luciferase (R-luc; 630 nm), respectively21,22,23. They were PCR-amplified as above to incorporate an A-sensitive, T-sensitive or G-sensitive target-binding site, respectively, to give U-RNASLG, A-RNASLO and C-RNASLR (Fig. 3). We also prepared an 18C-targeting probe, G-DNAgal (Fig. 3), encoding β-galactosidase (β-gal). The latter cleaves o-nitrophenyl β-galactoside as a substrate to give colored o-nitrophenol for colorimetric or visual assay.
To further enhance catalytic performance, we also use RNase H. The translation-activated mRNA probe reversibly formed from stoichiometric binding of the target to the probe (structure on the right-hand side of Fig. 1b) is, at best, equimolar to the target. This would lead to low signal intensity, especially when the target &1QJ;is present in a tiny amount. RNase H is an endonuclease that specifically cleaves RNA in the DNA/RNA heteroduplex24. It thus irreversibly digests the target-bound loop region of the probe to release the anti-RBS domain from the mRNA body, thus allowing catalytic use of target DNA (RNase H–coupled MB-mRNA; Fig. 1c) with improved selectivity and enhanced sensitivity25, although, given the nature of RNase H, it can be used only in vitro for DNA targets and not for in-cell gene sensing, where the targets should be mRNAs.
The translation- or transcription/translation-coupled sensing of ODN targets can be carried out in a normal Escherichia coli S30 extract system such as RTS HY100 (Roche), but we use a reconstituted prokaryotic cell–free transcription/translation system (Pure System; Post Genome Institute)26 throughout this work. Colorimetric assay of β-galactosidase cannot be conducted in the E. coli extract, which itself contains the enzyme. The effect of RNase H can also be most clearly evaluated in the reconstituted medium, which is otherwise free from RNase H.
Materials
Reagents
-
Agarose for 50–800-bp fragments (Nacalai Tesque, cat. no. 01147-96) for gel electrophoresis (see REAGENT SETUP)
-
β-Galactosidase enzyme assay system (Promega, cat. no. E2000) accompanied by the reporter lysis buffer
-
DNA oligomers as primers in PCR (made to order by Gene Design Inc.)
-
DNA stable marker 1-kbp ladder (Sigma Genosys, cat. no. MBMA1KBP-S)
-
DNeasy tissue kit (Qiagen, cat. no. 69504) for purification of total DNAs
-
dNTPs (accompanied by respective polymerase samples)
-
E. coli K-12 MG1655 strain (ATCC, for example)
-
Ethidium bromide solution (Nippon Gene, cat. no. 315-90051) for gel electrophoresis
Caution
Toxic.
-
Ethanol (Wako Pure Chemical Industries, cat. no. 053-06531)
Caution
Highly flammable.
-
GFX PCR DNA and gel band purification kit (GE Healthcare, cat. no. 27-9602-01) for purification of PCR products
-
Luciferase assay kit (Promega, cat. no. E1501)
-
Luria broth base powder (Invitrogen, cat. no. 12795-027)
-
MEGAShortScript T7 (Ambion, cat. no. 1354) or MEGAScript T7 Kit (Ambion, cat. no. 1333) for transcription
-
pBESTluc vector (Promega)
-
pSLG-test vector (Toyobo, cat. no. MRV-101)
-
pSLO-test vector (Toyobo, cat. no. MRV-102)
-
pSLR-test vector (Toyobo, cat. no. MRV-103)
-
PfuUltra high-fidelity DNA polymerase (Stratagene, cat. no. 600380) for PCR
-
Pure system standard classic II (Post Genome Institute, cat. no. PURE2030C) or pure system classic I (Post Genome Institute) as a reconstituted prokaryotic transcription/translation medium
-
Pyrobest DNA polymerase (Takara, cat. no. R005A) for PCR
-
RNase-free water (molecular biology reagent) (Sigma, cat. no. W4502)
Critical
RNase-free water should be used throughout the procedure.
-
RNeasy MinElute Cleanup kit (Qiagen, cat. no. 74204) for purification of RNA
-
TaKaRa Ex Taq hot start version (TaKaRa, cat. no. RR006A) for PCR
-
Tris-Acetate-EDTA (TAE) buffer (50×) (Nacalai Tesque, cat. no. 32666-81)
-
Tth RNase H (Toyobo, cat. no. RNH-201)
-
Buffer RPE (see REAGENT SETUP)
-
LB medium (see REAGENT SETUP)
-
Buffer AW1 (see REAGENT SETUP)
-
Buffer AW2 (see REAGENT SETUP)
Equipment
-
96-well assay plate (Costar)
-
Bio-shaker (Taitec, cat. no. BR-300LF)
-
Centrifuge (Waken, model 2320)
-
Densitograph (ATTO, cat. no. AE-6920V-FX)
-
GeneQuant pro (GE Healthcare)
-
iCycler thermalcycler dual block (Bio-Rad) for PCR
-
Lumat LB9507 luminometer (Berthold Detection System)
-
i-Mupid (COSMO BIO) for electrophoresis
-
Multilabel counter (Wallac, cat. no. 1420) for luciferase/galactosidase assay
-
25-ml reagent reservoir (VistaLab) for multichannel pipetting
-
Surgical stainless steel blades and handles (Feather Safety Razor) for gel cutting
Reagent setup
-
1% Agarose gel Weigh 0.5 g agarose, and dissolve in 50 ml MilliQ-grade deionized hot water. When the solution has cooled to 50 °C, add 1 ml 50× TAE buffer and 5 μl 10 μg μl−1 ethidium bromide solution. Mix thoroughly, and pour into gel tray.
-
Buffer RPE Combine 11 ml of the supplied concentrated buffer RPE (RNeasy MinElute Cleanup kit) and 44 ml ethanol.
-
Buffer AW1 Combine 19 ml of the supplied concentrated buffer AW1 (DNeasy Tisssue kit) and 25 ml ethanol.
-
Buffer AW2 Combine 13 ml of the supplied concentrated buffer AW2 (DNeasy Tisssue kit) and 30 ml ethanol.
-
LB medium Combine 7.5 g Luria broth base powder (Invitrogen) and 300 ml MilliQ-grade deionized water, and heat the mixture in an autoclave at 121 °C for 20 min.
Procedure
Preparation of template dsDNA (G-DNABEST)
Timing 5 h
-
1
Prepare a first PCR mix containing the following:
Table 1 Table 2 Critical Step
To prevent contamination, use nuclease-free filter tips.
-
2
Add 1 μl pBESTluc solution (5 ng μl−1) or 1 μl water (for negative control) to each mix in a PCR tube.
-
3
Run the following PCR cycles:
Table 2 Table 3 -
4
Check the PCR by running 5 μl of each reaction mixture on a 1% agarose gel with a 1-kbp ladder as a marker.
Pause point
The PCR product can be stored at −80 °C for 3 months.
-
5
Dilute the PCR product with water to give a 1:99 dilution.
-
6
Prepare a second PCR mix containing the following:
Table 3 Table 4 Critical Step
To prevent contamination, use nuclease-free filter tips.
-
7
Add 1 μl of the diluted first PCR solution or 1 μl water (for negative control) to each mix in a PCR tube.
-
8
Perform the PCR in the same manner as in Step 3, and check as in Step 4 to ensure the formation of the desired, allele C-targeting dsDNA template (G-DNABEST).
Critical Step
If the purity (as judged by gel electrophoresis) is not satisfactory (more than one spot), purify the PCR product with a GFX PCR DNA and gel band purification kit for subsequent transcription to give a single spot.
Pause point
The PCR product can be stored at −80 °C for 3 months.
Preparation of RNA probe G-RNABEST
Timing 15 h
-
9
Prepare the following MEGAshortscript T7 transcription mix in a PCR tube (MEGAscript T7 can be used in place of MEGAshortscript):
Table 4 Table 5 -
10
Mix briefly, spin down and tap the sample.
Pause point
Incubate the mixture overnight at 37 °C.
-
11
To decompose the DNA template, add 1 μl TURBO DNase solution, mix well and incubate at 37 °C for 15 min.
-
12
Purify the transcribed RNA probe (G-RNABEST) with an RNeasy MinElute Cleanup Kit according to the manufacturer's manual.
-
13
Determine the concentration of the purified probe by the absorbance at 260 nm, and dilute to give a stock solution of G-RNABEST (1 μg μl−1).
Critical Step
Avoid repeated freezing and thawing of the sample, because this results in decomposition of the RNA. For repeated use, divide the stock solution into 10-μl aliquots and freeze separately.
Pause point
The transcription product (G-RNABEST) can be stored at −80 °C for 2 weeks.
RNase H–uncoupled sensing of 18-nt targets with probe G-RNABEST
Timing 1.5 h
-
14
Prepare a reconstituted prokaryotic translation medium (solution AB) by mixing 35 μl of solution A and 5 μl of solution B of pure system classic I.
Critical Step
A normal E. coli S30 extract system (RTS HY100, Roche, cat no. 3 186 148) can be used with similar results in place of the reconstituted translation system.
-
15
Mix probe G-RNABEST and 18-nt full-match target (18C) in a PCR tube, as follows:
Table 5 Table 6 For negative control, add 1 μl water in place of target 18C.
Critical Step
Negative control without target is critical, as the target-on chemiluminescence intensity should always be evaluated in reference to that in the absence of the target.
-
16
Spin down and tap the mixture gently, and incubate for 1 h at 37 °C.
-
17
Add 10 μl water into each sample in the tube.
-
18
Take three 5-μl aliquots from each sample, and put them separately in a 96-well assay plate set in a multilabel counter (Wallac 1420).
-
19
Add 100 μl luciferase assay reagent to each well, and read the chemiluminescence intensity (I). Confirm that the presence of the probe gives rise to approximately threefold higher intensity than is seen in its absence (Ion/Ioff ≅ 3).
Critical Step
To standardize conditions for all samples, it is critical to use a multichannel pipette and reagent reservoir and to read the chemiluminescence intensity immediately after mixing the sample with the reagent by pipetting. Chemiluminescence intensity gradually decreases in a time-dependent manner, giving approximately 50% loss of intensity in 10 min.
RNase H–coupled sensing of 18-nt and 16-nt targets with probe G-RNABEST
Timing 2 h for each set of targets
-
20
For RNase H-coupled sensing, mix probe G-RNABEST, 18-nt target (fully matched 18C, singly A-mismatched 18A, singly T-mismatched 18T or doubly TA-mismatched 18TA) and RNase H in a PCR tube, as follows:
Table 6 Table 7 For negative control, add 1 μl water in place of target ODN.
Critical Step
Negative control without target is essential, as the target-on chemiluminescence intensity should always be evaluated in reference to that in the absence of the target.
-
21
Follow the translation (Step 16), workup (Steps 17 and 18) and assay (Step 19) procedures as above with 9 μl water in place of 10 μl in Step 17. Confirm that the Ion/Ioff ratio (target-on to target-off signal ratio) for the full-match target 18C is enhanced from 3 (in the absence of RNase H) to 6 (in its presence) with more (for 18A and 18TA) or less (for 18T) efficient base discrimination.
Critical Step
To standardize conditions for all samples, it is critical to use a multichannel pipette and reagent reservoir and to read the chemiluminescence intensity immediately after mixing the sample with the reagent by pipetting. Chemiluminescence intensity gradually decreases in a time-dependent manner, giving approximately 50% loss of intensity in 10 min.
Critical Step
To see the clear effect of RNase H, it is critical to use the reconstituted translation system, which is free from RNase H.
-
22
To check the effect of base-pair length in target-probe hybridization, use 16-nt targets (fully matched 16C and singly T-mismatched 16T) in place of the 18-nt counterparts under RNase H–uncoupled (Steps 15–19) and RNase H–coupled (Steps 20 and 21) conditions. Confirm that a significant on/off ratio (Ion/Ioff ≅ 6) is achieved only for the full-match target 16C, with a satisfactory T-allele discrimination capacity, under the RNase H–coupled conditions.
Critical Step
To standardize conditions for all samples, it is critical to use a multichannel pipette and reagent reservoir and to read the chemiluminescence intensity immediately after mixing the sample with the reagent by pipetting. Chemiluminescence intensity gradually decreases in a time-dependent manner, giving approximately 50% loss of intensity in 10 min.
RNase H–coupled sensing with G-DNABEST as a probe
Timing 1.5 h
-
23
Mix dsDNA template G-DNABEST (Step 8), 18-nt full-match target 18C and RNase H in a PCR tube, as follows:
Table 7 Table 8 For negative control, add 1 μl water in place of target 18C.
Critical Step
Negative control without target is essential, as the target-on chemiluminescence intensity should always be evaluated in reference to that in the absence of the target.
-
24
Follow the translation (Step 16), workup (Steps 17 and 18) and assay (Step 19) procedures as above with 9 μl water in place of 10 μl in Step 17. Confirm that the on/off ratio is much enhanced, to approximately 13.
Critical Step
To standardize conditions for all samples, it is critical to use a multichannel pipette and reagent reservoir and to read the chemiluminescence intensity immediately after mixing the sample with the reagent by pipetting. Chemiluminescence intensity gradually decreases in a time-dependent manner, giving approximately 50% loss of intensity in 10 min.
Preparation of template dsDNAs of multicolor reporter proteins (T-DNASLG, A-DNASLO and C-DNASLR)
Timing 6 h for each template
-
25
For the preparation of T-DNASLG, which is transcribable into A-targeting U-RNASLG, prepare a first PCR mix containing the following:
Table 8 Table 9 Critical Step
To prevent contamination, use nuclease-free filter tips.
-
26
Add 2 μl solution of green luciferase–encoding vector pSLG-test (5 ng μl−1) or 2 μl water (for negative control) to each mix in a PCR tube.
-
27
Run the following PCR cycles:
Table 9 Table 10 -
28
Check the PCR by running 5 μl of reaction mixture on a 1% agarose gel with a 1-kbp ladder as a marker. If the purity (as judged by gel electrophoresis) is not satisfactory (more than one spot), purify the PCR product with a GFX PCR DNA and gel band purification kit according to the manufacturer's manual.
Pause point
The PCR product can be stored at −80 °C for 3 months.
-
29
Dilute the purified PCR product with water to give a 1:199 dilution.
-
30
Prepare a second PCR mix containing the following:
Table 10 Table 11 Critical Step
To prevent contamination, use nuclease-free filter tips.
-
31
Add 1 μl of the diluted purified first PCR solution or 1 μl water (for negative control) to each mix in a PCR tube.
-
32
Perform the PCR in exactly the same manner as in Step 27.
-
33
Check the purity and, when it is not satisfactory, purify the product as in Step 28.
Pause point
The PCR product can be stored at −80 °C for 3 months.
-
34
Dilute the purified second PCR product as in Step 29.
-
35
Prepare a third PCR mix containing the following:
Table 11 Table 12 -
36
Add 2 μl of the diluted purified second PCR solution or 2 μl water (for negative control) to each mix in a PCR tube.
-
37
Perform the PCR in exactly the same manner as in Step 27.
-
38
Check the purity, and purify the product dsDNA, that is, T-DNASLG, as in Step 28.
Pause point
The PCR product can be stored at −80 °C for 3 months.
-
39
For the preparation of A-DNASLO transcribable into T-targeting A-RNASLO, follow Steps 25–38 using orange luciferase–encoding vector pSLO-test and A-SLO relevant primers (1st-fw-A-SLO, 1st-rv-A-SLO, 2nd-fw-A-SLO, 2nd-rv-A-SLO, 3rd-fw-A-SLO and 3rd-rv-A-SLO) in place of pSLG-test (Step 26) and T-SLG relevant ones (1st-fw-U-SLG, 1st-rv-U-SLG, 2nd-fw-U-SLG and 2nd-rv-U-SLG, 3rd-fw-U-SLG and 3rd-rv-U-SLG) (Steps 25, 30 and 35).
-
40
For the preparation of C-DNASLR transcribable into G-targeting C-RNASLR, follow Steps 25–38 using red luciferase–encoding vector pSLR-test and C-SLR relevant primers (1st-fw-C-SLR, 1st-rv-C-SLR, 2nd-fw-C-SLR, 2nd-rv-C-SLR, 3rd-fw-C-SLR and 3rd-rv-C-SLR) in place of pSLG-test (Step 26) and U-SLG relevant ones (1st-fw-U-SLG, 1st-rv-U-SLG, 2nd-fw-U-SLG, 2nd-rv-U-SLG, 3rd-fw-U-SLG and 3rd-rv-U-SLG) (Steps 25, 30 and 35).
Preparation of RNA probes, U-RNASLG, A-RNASLO and C-RNASLR
Timing 15 h for each probe
-
41
For U-RNASLG, prepare the following MEGAshortscript T7 transcription mix in a PCR tube (MEGAscript T7 can be used in place of MEGAshortscript):
Table 12 Table 13 -
42
Carry out transcription as in Steps 10–12. Determine the concentration of purified product by the absorbance at 260 nm, and dilute to give a stock solution of probe U-RNASLG (1 μg μl−1).
Critical Step
Avoid repeated freezing and thawing of the sample, because this results in decomposition of the RNA. For repeated use, divide the stock solution into 10-μl aliquots and freeze.
Pause point
The transcription product (U-RNASLG) can be stored at −80 °C for 2 weeks.
-
43
For the preparation of A-RNASLO, follow Steps 41 and 42 using A-DNASLO in place of T-DNASLG.
-
44
For the preparation of C-RNASLR, follow Steps 41 and 42 using C-DNASLR in place of T-DNASLG.
Sensing of 16-nt targets with allele-selective RNA probes
Timing 2 h for each set of targets
-
45
Prepare a reconstituted prokaryotic translation medium (solution AB) by mixing 25 μl solution A and 10 μl solution B of pure system classic II.
-
46
Mix probe U-RNASLG, target (16A, 16T, 16C or 16G) and RNase H in a PCR tube, as follows:
Table 13 Table 14 For negative control, add 1 μl water in place of target ODN.
Critical Step
Negative control without target is essential, as the target-on chemiluminescence intensity should always be evaluated in reference to that in the absence of the target.
-
47
Carry out target-dependent translation of green luciferase as in Step 16. Follow the workup and assay procedures as in Steps 17–19, but by adding 11 μl water in place of 10 μl in Step 17, taking 5.5-μl aliquots in place of 5-μl aliquots in Step 18 and using a luminometer in place of a multilabel counter. Confirm that only A-allele target 16A gives rise to a significant increase in the chemiluminescence intensity, that is, on/off ratio.
Critical Step
To standardize conditions for all samples, it is critical to read the chemiluminescence intensity immediately after mixing the sample with the reagent by pipetting. Chemiluminescence intensity gradually decreases in a time-dependent manner, giving approximately 50% loss of intensity in 10 min.
-
48
Mix probe A-RNASLO, target (16A, 16T, 16C or 16G) and RNase H in a PCR tube as in Step 46. Follow the translation and assay procedures in Step 47 for orange luciferase. Confirm that only T-allele target 16T gives rise to a significant increase in the chemiluminescence intensity, that is, on/off ratio.
-
49
Mix probe C-RNASLR, target (16A, 16T, 16C or 16G) and RNase H in a PCR tube as in Step 46. Follow the translation and assay procedures in Step 47 for red luciferase. Confirm that only G-allele target 16G gives rise to a significant increase in the chemiluminescence intensity, that is, on/off ratio.
Critical Step
For multicolor SNP detection in a single tube, it is critical to optimize the quantities of the three probes or use Ion/Ioff criteria for evaluation, as the translation efficiencies and emission coefficients for the three types (green, orange and red) of luminescence are different.
Preparation of dsDNA template G-DNAgal for a colorimetric assay based on a β-galactosidase reporter protein
Timing 25 h
-
50
Inoculate E. coli K-12 MG1655 strain in 5 ml LB medium in a plastic tube.
-
51
Incubate the mixture for 16 h at 37 °C with vigorous shaking.
-
52
Purify total DNAs in the strain with a DNeasy tissue kit according to the manufacturer's manual.
-
53
Determine the concentration of the purified total DNAs by the absorbance at 260 nm.
Pause point
The DNAs can be stored at −80 °C for 3 months.
-
54
Prepare the first PCR mix containing the following:
Table 14 Table 15 Critical Step
To prevent contamination, use nuclease-free filter tips.
-
55
Add 1 μl of the total DNA solution obtained above (5 ng μl−1) or 1 μl water (for negative control) to each mix in a PCR tube.
-
56
Run the following PCR cycles:
Table 15 Table 16 -
57
Check the PCR by running 5 μl of each reaction on a 1% agarose gel with a 1-kbp ladder as a marker.
Pause point
The PCR product can be stored at −80 °C for 3 months.
-
58
Dilute the PCR product with water to give a 1:99 dilution.
-
59
Prepare the second PCR mix containing the following:
Table 16 Table 17 Critical Step
To prevent contamination, use nuclease-free filter tips.
-
60
Add 1 μl of the diluted first PCR solution or 1 μl water (for negative control) to each mix in a PCR tube.
-
61
Run the following PCR cycles:
Table 17 Table 18 -
62
Check as in Step 57 to make sure of the formation of the desired dsDNA template, that is, G-DNAgal.
Pause point
The PCR product can be stored at −80 °C for 3 months.
RNase H–coupled sensing with G-DNAgal as a probe
Timing 2.5 h
-
63
Mix full-match 18-nt target 18C, dsDNA probe G-DNAgal and RNase H in a PCR tube, as follows:
Table 18 Table 19 For negative control, add 1 μl water in place of target 18C.
Critical Step
Negative control without target is essential, as the target-on chemiluminescence intensity should always be evaluated in reference to that in the absence of the target.
Critical Step
It is critical to use the reconstituted translation system. A normal E. coli extract cannot be used, because it contains β-galactosidase.
-
64
Spin down and tap the mixture gently, and incubate for 1 h at 37 °C.
-
65
Mix the transcription/translation solution with 39 μl 1× reporter lysis buffer and 50 μl assay 2× buffer containing o-nitrophenyl β-galactoside of the β-galactosidase assay system.
-
66
Incubate the resulting solution in a 96-well plate for 1 h at 37 °C.
-
67
Read the absorbance at 405 nm with a multilabel counter (Wallac 1420), and take a photographic image of each sample.
Troubleshooting
Troubleshooting advice can be found in Table 1.
Anticipated results
Steps 19, 21, 22 and 24
As translation of reporter protein is not intended to be completely suppressed in the absence of target (see INTRODUCTION), it is most appropriate to refer to the on/off ratio of the chemiluminescence intensity (Ion) in the presence of a particular target to that (Ioff) in its absence. Anticipated Ion/Ioff ratios are summarized in Figure 6. In the absence of RNase H, target 18C can be sensed by the full-match probe G-RNABEST with Ion/Ioff = 3.1 (lane 1). In the presence of RNase H, the sensitivity becomes higher (Ion/Ioff = 6.2; lane 2) with good discrimination of doubly mutated (18TA; Ion/Ioff = 1.3; lane 3) and singly A-mutated (18A; Ion/Ioff = 1.7; lane 4) targets. This is, however, not the case for singly T-mutated target (18T; Ion/Ioff = 4.3; lane 5), which would form a relatively stable GT mismatch in the target-probe heteroduplex. The on/off ratio for target 18C becomes even more pronounced when using G-DNABEST (Ion/Ioff = 12.5; lane 6) in place of G-RNABEST (Ion/Ioff = 6.2; lane 2). The advantage of the RNase H–coupled system can also be clearly demonstrated when using a smaller amount of target. Target 18C in 45 fmol in 2.5 μl of the medium (18 nM) can scarcely be detected (Ion/Ioff = 1.4; lane 7) in the absence of RNase H, but in its presence it is readily sensed, with Ion/Ioff = 3.8 (lane 8), when a sensitive luminometer (Lumat LB 9507) is used. The detection limit with respect to full-match target 18C seems to lie in the range of 9 fmol (Ion/Ioff = 1.7).
In the case of short 16-nt targets, even 16C (Ion/Ioff = 1.4; lane 9), in addition to 16T (Ion/Ioff = 0.94; lane 10), cannot be sensed effectively in the absence of RNase H. However, in the presence of the latter, the full-match C-allele target gives rise to a significant signal enhancement (Ion/Ioff = 5.7; lane 11), whereas the T-mismatch remains inactive (Ion/Ioff = 1.4;lane 12).
Steps 47–49
The chemiluminescence intensity (Ion) for any mismatch target-probe combination—that is, 16A/A-RNASLO, 16A/C-RNASLR, 16T/U-RNASLG, 16T/C-RNASLR, 16C/A-RNASLO, 16C/U-RNASLG, 16C/C-RNASLR, 16G/U-RNASLG and 16G/A-RNASLO—is hardly distinguishable from that (Ioff) in the absence of target (Ion/Ioff ≅ 1 in most cases). On the other hand, matched target/probe combinations, that is, 16A/U-RNASLG and 16T/A-RNASLO (Ion/Ioff ≅ 4) and 16G/C-RNASLR (Ion/Ioff ≅ 3), give rise to higher signal intensities, as shown in Figure 7. Thus, any probe sensitively responds to its partner target on a complementarity basis.
Step 67
The anticipated target-on/target-off ratio of the absorbance at 405 nm with RNase H is Ion/Ioff = 12.6 (Fig. 6, lane 13). o-Nitrophenol formed can be easily detected even visually (Fig. 8).
References
Nakatani, K. Chemistry challenges in SNP typing. Chembiochem 5, 1623–1633 (2004).
Silverman, A.P. & Kool, E.T. Quenched probes for highly specific detection of cellular RNAs. Trends Biotechnol. 23, 225–230 (2005).
Tyagi, S. & Kramer, F.R. Molecular beacons: probes that fluoresce upon hybridization. Nat. Biotechnol. 14, 303–308 (1996).
Stojanovic, M.N., de Prada, P. & Landry, D.W. Catalytic molecular beacons. Chembiochem 2, 411–415 (2001).
Hartig, J.S., Grune, I., Najafi-Shoushtari, S.H. & Famulok, M. Sequence-specific detection of MicroRNAs by signal-amplifying ribozymes. J. Am. Chem. Soc. 126, 722–723 (2004).
Sando, S., Sasaki, T., Kanatani, K. & Aoyama, Y. Amplified nucleic acid sensing using programmed self-cleaving DNAzyme. J. Am. Chem. Soc. 125, 15720–15721 (2003).
Saghatelian, A., Guckian, K.M., Thayer, D.A. & Ghadiri, M.R. DNA detection and signal amplification via an engineered allosteric enzyme. J. Am. Chem. Soc. 125, 344–345 (2003).
Pavlov, V., Shlyahovsky, B. & Willner, I. Fluorescence detection of DNA by the catalytic activation of an aptamer/thrombin complex. J. Am. Chem. Soc. 127, 6522–6523 (2005).
Abe, H. & Kool, E.T. Destabilizing universal linkers for signal amplification in self-ligating probes for RNA. J. Am. Chem. Soc. 126, 13980–13986 (2004).
Cai, J., Li, X., Yue, X. & Taylor, J.S. Nucleic acid-triggered fluorescent probe activation by the Staudinger reaction. J. Am. Chem. Soc. 126, 16324–16325 (2004).
Fan, C., Plaxco, K.W. & Heeger, A.J. Electrochemical interrogation of conformational changes as a reagentless method for the sequence-specific detection of DNA. Proc. Natl. Acad. Sci. USA 100, 9134–9137 (2003).
Patolsky, F., Weizmann, Y., Katz, E. & Willner, I. Magnetically amplified DNA assays (MADA): sensing of viral DNA and single-base mismatches by using nucleic acid modified magnetic particles. Angew. Chem. Int. Ed. Engl. 42, 2372–2376 (2003).
Sando, S., Narita, A., Abe, K. & Aoyama, Y. Doubly catalytic sensing of HIV-1-related CCR5 sequence in prokaryotic cell-free translation system using riboregulator-controlled luciferase activity. J. Am. Chem. Soc. 127, 5300–5301 (2005).
Brantl, S. Antisense-RNA regulation and RNA interference. Biochim. Biophys. Acta 1575, 15–25 (2002).
Isaacs, F.J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nat. Biotechnol. 22, 841–847 (2004).
Mertens, N., Remaut, E. & Fiers, W. Increased stability of phage T7g10 mRNA is mediated by either a 5′- or a 3′-terminal stem-loop structure. Biol. Chem. 377, 811–817 (1996).
Rosi, N.L. & Mirkin, C.A. Nanostructures in biodiagnostics. Chem. Rev. 105, 1547–1562 (2005).
Breaker, R.R. Natural and engineered nucleic acids as tools to explore biology. Nature 432, 838–845 (2004).
Mummidi, S., Ahuja, S.S., McDaniel, B.L. & Ahuja, S.K. The human CC chemokine receptor 5 (CCR5) gene. Multiple transcripts with 5′-end heterogeneity, dual promoter usage, and evidence for polymorphisms within the regulatory regions and noncoding exons. J. Biol. Chem. 272, 30662–30671 (1997).
Bhaumik, S. & Gambhir, S.S. Optical imaging of Renilla luciferase reporter gene expression in living mice. Proc. Natl. Acad. Sci. USA 99, 377–382 (2002).
Nakajima, Y., Kimura, T., Suzuki, C. & Ohmiya, Y. Improved expression of novel red- and green-emitting luciferases of Phrixothrix railroad worms in mammalian cells. Biosci. Biotechnol. Biochem. 68, 948–951 (2004).
Nakajima, Y., Ikeda, M., Kimura, T., Honma, S., Ohmiya, Y. & Honma, K. Bidirectional role of orphan nuclear receptor RORα in clock gene transcriptions demonstrated by a novel reporter assay system. FEBS Lett. 565, 122–126 (2004).
Viviani, V.R., Bechara, E.J. & Ohmiya, Y. Cloning, sequence analysis and expression of active Phrixothrix railroadworms luciferases: relationship between bioluminescence spectra and primary structures. Biochemistry 38, 8271–8279 (1999).
Duck, P., Alvarado-Urbina, G., Burdick, B. & Collier, B. Probe amplifier system based on chimeric cycling oligonucleotides. BioTechniques 9, 142–148 (1990).
Narita, A., Ogawa, K., Sando, S. & Aoyama, Y. Visible sensing of nucleic acid sequences with a genetically encodable unmodified RNA probe. Angew. Chem. Int. Ed. Engl. 45, 2879–2883 (2006).
Shimizu, Y. et al. Cell-free translation reconstituted with purified components. Nat Biotechnol. 19, 751–755 (2001).
Acknowledgements
We thank Drs. T. Kanamori and Y. Shimizu of the University of Tokyo for their kind advice and technical support. This work was supported by the Industrial Technology Research Grant Program of the New Energy and Industrial Technology Development Organization (NEDO) of Japan (S.S.) and by the Grant-in-Aid for Scientific Research (KAKENHI) in Priority Area “Molecular Nano Dynamics” (no. 17034026 for Y.A.) from the Ministry of Education, Culture, Sports, Science and Technology, Japan. We thank Toyobo Co. Ltd. for technical advice.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare a pending patent application whose value may be affected by publication of this paper.
Rights and permissions
About this article
Cite this article
Narita, A., Ogawa, K., Sando, S. et al. Cis-regulatory hairpin-shaped mRNA encoding a reporter protein: catalytic sensing of nucleic acid sequence at single nucleotide resolution. Nat Protoc 2, 1105–1116 (2007). https://doi.org/10.1038/nprot.2007.140
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.140
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.