Abstract
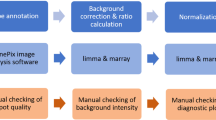
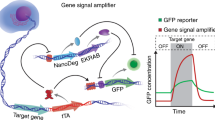
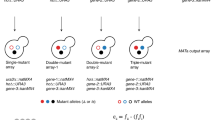
Gene overexpression can be used to investigate the biological pathways that are important in the response to a small molecule or other perturbation. To facilitate the use of gene overexpression in the study of small-molecule mechanisms, we developed a microarray-based protocol for monitoring the growth of a pool of yeast strains, each overexpressing a different protein. In this protocol, yeast harboring a set of ∼3,900 galactose-inducible overexpression plasmids are grown in the absence or presence of a small molecule for multiple generations. The plasmids are then extracted from the two populations, processed and labeled in such a manner that their relative concentrations can be determined by competitive hybridization to a microarray. Although this protocol was developed for monitoring a specific set of overexpression plasmids, it could presumably be adapted to monitor yeast that have been transformed with any set of plasmids for which the gene inserts have been spotted, or otherwise arrayed, in a microarray format. This protocol can be completed in approximately 15 hours of hands-on time over the course of several days.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nature Genet. 21, 278–283 (1999).
Lum, P.Y. et al. Discovering modes of action for therapeutic compounds using a genome-wide screen of yeast heterozygotes. Cell 116, 121–137 (2004).
Giaever, G. et al. Chemogenomic profiling: identifying the functional interactions of small molecules in yeast. Proc. Natl. Acad. Sci. USA 101, 793–798 (2004).
Parsons, A.B. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nature Biotechnol. 22, 62–69 (2004).
Butcher, R.A. et al. Microarray-based method for monitoring yeast overexpression strains reveals small-molecule targets in TOR pathway. Nature Chem. Biol. 2, 103–109 (2006).
Schiestl, R.H. & Gietz, R.D. High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Curr. Genet. 16,339–346 (1989).
Soni, R., Carmichael, J.P. & Murray, J.A. Parameters affecting lithium acetate-mediated transformation of Saccharomyces cerevisiae and development of a rapid and simplified procedure. Curr. Genet. 24, 455–459 (1993).
Gollub, J. et al. The Stanford Microarray Database: data access and quality assessment tools. Nucleic Acids Res. 31, 94–96 (2003).
Heitman, J., Movva, N.R. & Hall, M.N. Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science 253, 905–909 (1991).
Acknowledgements
We acknowledge Chih Long Liu for advice regarding the in vivo transcription amplification strategy. We thank the National Institute of General Medical Sciences (GM38627) for support of this research. S.L.S. is an Investator at the Howard Hughes Medical Institute. R.A.B. was supported by a graduate fellowship from the National Science Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Butcher, R., Schreiber, S. A microarray-based protocol for monitoring the growth of yeast overexpression strains. Nat Protoc 1, 569–576 (2006). https://doi.org/10.1038/nprot.2006.80
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.80
This article is cited by
-
Dosage suppression genetic interaction networks enhance functional wiring diagrams of the cell
Nature Biotechnology (2011)
-
A molecular barcoded yeast ORF library enables mode-of-action analysis of bioactive compounds
Nature Biotechnology (2009)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.