Abstract
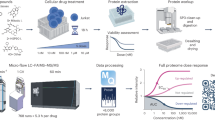
Although differences in protein staining intensity can often be visualized by difference gel electrophoresis, abundant proteins can obscure less abundant proteins, and quantification of post-translational modifications is difficult. We present a protocol for quantifying changes in the abundance of a specific protein or changes in specific modifications of a protein using in-gel stable isotope labeling. In this protocol protein extracts from any source treated under two experimental conditions are resolved in two separate lanes by gel electrophoresis. Parallel gel regions of interest are reacted separately with either light or heavy isotope-labeled reagents, and the gel slices are then combined and digested with proteases. The resulting peptides are then analyzed by liquid chromatography/mass spectrometry (LC/MS) to determine relative abundance of light- and heavy-isotope lysine-containing peptide pairs and analyzed by LC/MS/MS for identification of sequence and modifications. This protocol should take approximately 24–26 h to complete, including the incubation time for proteolytic digestion. Additional time will be needed for data analysis and interpretation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Mann, M. & Jensen, O.N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 21, 255–261 (2003).
Lill, J. Proteomic tools for quantitation by mass spectrometry. Mass Spectrom. Rev. 22, 182–194 (2003).
Lilley, K.S. & Friedman, D.B. All about DIGE: quantification technology for differential-display 2D-gel proteomics. Expert Rev. Proteomics 4, 401–409 (2004).
Kolkman, A., Dirksen, E.H., Slijper, M. & Heck, A.J. Double standards in quantitative proteomics—direct comparative assessment of difference in gel electrophoresis and metabolic stable isotope labeling. Mol. Cell. Proteomics 3, 255–266 (2005).
Gygi, S.O. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Ji, J. et al. Strategy for qualitative and quantitative analysis in proteomics based on signature peptides. J. Chromatogr. A 745, 197–210 (2000).
Ross, P.L., Huang, Y.L., Marchese, J.N. & Pappin, D.J. Multiplexed protein quantification in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Mirgorodskaya, O.A. et al. Quantification of peptides and proteins by matrix-assisted laser desorption/ionization mass spectrometry using 18O-labeled internal standards. Rapid Comm. Mass Spectrom. 14, 1226–1232 (2000).
Schmidt, A., Kellermann, J. & Lottspeich, F. A novel strategy for quantitative proteomics using isotope-coded protein labels. Proteomics 5, 4–15 (2005).
Asara, J.M. et al. In-gel stable isotope labeling (ISIL): a strategy for mass spectrometry-based relative quantification. J. Proteome Res. 5, 155–163 (2006).
Chakraborty, A. & Regnier, F.E. Global internal standard technology for comparative proteomics. J. Chromatogr. A 949, 173–184 (2002).
Zhang, X. et al. An automated method for the analysis of stable isotope labeling data in proteomics. J. Am. Soc. Mass Spectrom. 16, 1181–1191 (2005).
Acknowledgements
The authors thank Takeda Chemical Industries for providing startup funds for the BIDMC Mass Spectrometry and Proteomics Core and NIH Grants GM56203 and CA089021 for providing support for this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Asara, J., Zhang, X., Zheng, B. et al. In-gel stable isotope labeling for relative quantification using mass spectrometry. Nat Protoc 1, 46–51 (2006). https://doi.org/10.1038/nprot.2006.7
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.