Abstract
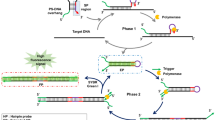
Here we describe a protocol for the amplified detection of a target DNA using a DNA/FokI-based replicating cutting machine. The protocol is based on the design of a sensing hairpin oligonucleotide that is opened upon hybridization with the analyte DNA. The endonuclease FokI binds to the double-stranded complex and cleaves it to a “cutter” unit. The “cutter” unit reacts with a fuel oligonucleotide to generate and amplify the signal. The fuel molecule is an oligonucleotide in a hairpin configuration with a fluorophore/quencher pair attached to the 5′ and 3′ ends. Formation of the duplex between the cutter and the fuel leads to the scission of the duplex by FokI, leading to a second, replicated “cutter”, a fluorescent waste product, and to the regeneration of the original “cutter” unit. The autonomous replication of the “cutter” unit, as a result of the primary recognition of the analyte DNA, leads to the amplified fluorescent detection of the analyte DNA with a sensitivity limit of 1 × 10−14 M. The operation of the machine and the sensing process are monitored by the fluorescence generated by the waste product. Here we apply the protocol, which takes about 2 h to complete, to analyze a Tay-Sachs genetic disorder mutant DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Weizmann, Y., Cheglakov, Z., Pavlov, V. & Willner, I. Autonomous fueled mechanical replication of nucleic acid templates for the amplified optical detection of DNA. Angew. Chem. Int. Edn. Engl. 45, 2238–2242 (2006).
Szybalski, W. Universal restriction endonucleases: designing novel cleavage specificities by combining adaptor oligodeoxynucleotide and enzyme moieties. Gene 40, 169–173 (1985).
Lyamichev, V. et al. Polymorphism identification and quantitative detection of genomic DNA by invasive cleavage of oligonucleotide probes. Nat. Biotechnol. 17, 292–296 (1999).
Notomi, T. et al. Loop-mediated isothermal amplification. Nucleic Acids Res. 28, E63 (2000).
Lizardi, P.M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Solinas, A. et al. Duplex Scorpion primers in SNP analysis and FRET applications. Nucleic Acids Res. 29, E96 (2001).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Weizmann, Y., Cheglakov, Z., Pavlov, V. et al. An autonomous fueled machine that replicates catalytic nucleic acid templates for the amplified optical analysis of DNA. Nat Protoc 1, 554–558 (2006). https://doi.org/10.1038/nprot.2006.78
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.78
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.