Abstract
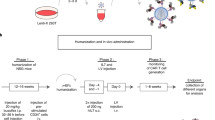
T-cell receptor (TCR) transgenic (Tg) mice have revolutionized our understanding of many aspects of T-cell biology. Whereas they provide an almost unlimited source of T cells with a single specificity, breeding them onto different backgrounds and/or new knockout/knock-in mouse models is often time-consuming (6 months to several years), which can make the process costly and can significantly delay research. This protocol describes a new method for expressing defined TCR-α and TCR-β proteins from a single 2A peptide–linked multicistronic retroviral vector in mice, using retrovirus-mediated stem cell gene transfer. We refer to these as 'retrogenic' (Rg) mice ('retro' from retrovirus and 'genic' from Tg) to avoid confusion with traditional transgenic mice. We have successfully used this approach to express over 50 different TCRs on several different mouse backgrounds in as little as 6 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
de Felipe, P. Polycistronic viral vectors. Curr. Gene Ther. 2, 355–378 (2002).
de Felipe, P. Skipping the co-expression problem: the new 2A “CHYSEL” technology. Genet. Vaccines Ther. 2, 13 (2004).
de Felipe, P. et al. E unum pluribus: multiple proteins from a self-processing polyprotein. Trends Biotechnol. 24, 68–75 (2006).
Szymczak, A.L. & Vignali, D.A. Development of 2A peptide-based strategies in the design of multicistronic vectors. Expert Opin. Biol. Ther. 5, 627–638 (2005).
Ryan, M.D. & Drew, J. Foot-and-mouth disease virus 2A oligopeptide mediated cleavage of an artificial polyprotein. EMBO J. 13, 928–933 (1994).
Donnelly, M.L. et al. Analysis of the aphthovirus 2A/2B polyprotein 'cleavage' mechanism indicates not a proteolytic reaction, but a novel translational effect: a putative ribosomal 'skip'. J. Gen. Virol. 82, 1013–1025 (2001).
Szymczak, A.L. et al. Correction of multi-gene deficiency in vivo using a single 'self-cleaving' 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Miller, J.F. & Flavell, R.A. T-cell tolerance and autoimmunity in transgenic models of central and peripheral tolerance. Curr. Opin. Immunol. 6, 892–899 (1994).
Benoist, C. & Mathis, D. Positive and negative selection of the T cell repertoire in MHC class II transgenic mice. Semin. Immunol. 1, 117–124 (1989).
Arnold, P.Y., Burton, A.R. & Vignali, D.A. Diabetes incidence is unaltered in glutamate decarboxylase 65-specific TCR retrogenic nonobese diabetic mice: generation by retroviral-mediated stem cell gene transfer. J. Immunol. 173, 3103–3111 (2004).
Holst, J., Vignali, K.M., Burton, A.R. & Vignali, D.A.A. Rapid analysis of T-cell selection in vivo using T cell-receptor retrogenic mice. Nat. Methods 3, 191–197 (2006).
Baldwin, T.A., Sandau, M.M., Jameson, S.C. & Hogquist, K.A. The timing of TCR alpha expression critically influences T cell development and selection. J. Exp. Med. 202, 111–121 (2005).
Szymczak-Workman, A.L., Vignali, K.M. & Vignali, D.A.A. Generation of 2A peptide-linked multicistronic vectors. in DNA Delivery/Gene Transfer Lab Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, in the press).
Szymczak, A.L. et al. The CD3-ε proline-rich sequence, and its interaction with Nck, is not required for T cell development and function. J. Immunol. 175, 270–275 (2005).
Persons, D.A., Mehaffey, M.G., Kaleko, M., Nienhuis, A.W. & Vanin, E.F. An improved method for generating retroviral producer clones for vectors lacking a selectable marker gene. Blood Cells Mol. Dis. 24, 167–182 (1998).
Persons, D.A. et al. Retroviral-mediated transfer of the green fluorescent protein gene into murine hematopoietic cells facilitates scoring and selection of transduced progenitors in vitro and identification of genetically modified cells in vivo. Blood 90, 1777–1786 (1997).
Hawley, R.G., Lieu, F.H., Fong, A.Z. & Hawley, T.S. Versatile retroviral vectors for potential use in gene therapy. Gene Ther. 1, 136–138 (1994).
Persons, D.A. et al. Use of the green fluorescent protein as a marker to identify and track genetically modified hematopoietic cells. Nat. Med. 4, 1201–1205 (1998).
Rees, W.A., Yager, T.D., Korte, J. & von Hippel, P.H. Betaine can eliminate the base pair composition dependence of DNA melting. Biochemistry 32, 137–144 (1993).
Varadaraj, K. & Skinner, D.M. Denaturants or cosolvents improve the specificity of PCR amplification of a G + C-rich DNA using genetically engineered DNA polymerases. Gene 140, 1–5 (1994).
Baskaran, N. et al. Uniform amplification of a mixture of deoxyribonucleic acids with varying GC content. Genome Res. 6, 633–638 (1996).
Henke, W., Herdel, K., Jung, K., Schnorr, D. & Loening, S.A. Betaine improves the PCR amplification of GC-rich DNA sequences. Nucleic Acids Res. 25, 3957–3958 (1997).
Acknowledgements
We are grateful to everyone in the Vignali lab for helping this protocol evolve into its current form. This work was supported by the NIH (AI52199, AI39480), the Juvenile Diabetes Research Foundation International (1-2004-141), a Cancer Center Support CORE grant (CA-21765) and the American Lebanese Syrian Associated Charities (ALSAC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Holst, J., Szymczak-Workman, A., Vignali, K. et al. Generation of T-cell receptor retrogenic mice. Nat Protoc 1, 406–417 (2006). https://doi.org/10.1038/nprot.2006.61
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.61
This article is cited by
-
Celastrol attenuates hepatitis C virus translation and inflammatory response in mice by suppressing heat shock protein 90β
Acta Pharmacologica Sinica (2023)
-
EventDNA: a dataset for Dutch news event extraction as a basis for news diversification
Language Resources and Evaluation (2023)
-
Newborn and child-like molecular signatures in older adults stem from TCR shifts across human lifespan
Nature Immunology (2023)
-
A cardioimmunologist’s toolkit: genetic tools to dissect immune cells in cardiac disease
Nature Reviews Cardiology (2022)
-
Covalent TCR-peptide-MHC interactions induce T cell activation and redirect T cell fate in the thymus
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.