Abstract
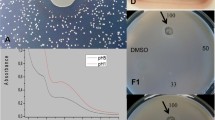
The fungal cell wall is an essential organelle and represents a considerable metabolic investment. Its macromolecular composition, molecular organization and thickness can vary greatly depending on environmental conditions. Its construction is also tightly controlled in space and time. Many genes are therefore involved in building the fungal cell wall. Here we present a simple approach for detecting these genes. The method is based on the observation that cell wall mutants are generally more sensitive to two related anionic dyes, Calcofluor white (CFW) and Congo red (CR), both of which interfere with the construction and stress response of the cell wall. CFW-based and CR-based susceptibility assays identify cell wall mutants not only in ascomycetous yeasts (such as Saccharomyces cerevisiae and Candida albicans) but also in mycelial ascomycetes (such as Aspergillus fumigatus and Aspergillus niger), basidiomycetous species (Cryptococcus neoformans) and probably also zygomycetous fungi. The protocol can be completed in 4–6 h (excluding the incubation time required for fungal growth).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Roncero, C., Valdivieso, M.H., Ribas, J.C. & Duran, A. Effect of calcofluor white on chitin synthases from Saccharomyces cerevisiae. J. Bacteriol. 170, 1945–1949 (1988).
Kopecka, M. & Gabriel, M. The influence of Congo red on the cell wall and (1-3-beta-D-glucan microfibril biosynthesis in Saccharomyces cerevisiae. Arch. Microbiol. 158, 115–126 (1992).
Wood, P.J. Specificity in the interaction of direct dyes with polysaccharides. Carb. Res. 85, 271–287 (1980).
Herth, W. Calcofluor white and Congo red inhibit chitin microfibril assembly of Poterioochromonas: evidence for a gap between polymerization and microfibril formation. J. Cell Biol. 87, 442–450 (1980).
Pringle, J.R. Staining of bud scars and other cell wall chitin with calcofluor. Methods Enzymol. 194, 732–735 (1991).
Vannini, G.L., Poli, P., Donini, A. & Pancaldi, S. Effects of Congo red on wall synthesis and morphogenesis in Saccharomyces cerevisiae. Plant Sci. Lett. 31, 9–17 (1983).
Roncero, C. & Duran, A. Effect of calcofluor white and Congo red on fungal cell wall morphogenesis: in vivo activation of chitin polymerization. J. Bacteriol. 163, 1180–1185 (1985).
Pancaldi, S., Poli, F., Dall'Olio, G. & Vannini, G.L. Morphological anomalies induced by Congo red in Aspergillus niger. Arch. Microbiol. 137, 185–187 (1984).
Damveld, R.A. et al. Expression of agsA, one of five 1,3-alpha-D-glucan synthase-encoding genes in Aspergillus niger, is induced in response to cell wall stress. Fungal Genet. Biol. 42, 165–177 (2005).
Damveld, R.A. et al. The Aspergillus niger MADS-box transcription factor RlmA is required for cell wall reinforcement in response to cell wall stress. Mol. Microbiol. 58, 305–319 (2005).
Levin, D.E. Cell wall integrity signaling in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 69, 262–291 (2005).
Boorsma, A. et al. Characterization of the transcriptional response to cell wall stress in Saccharomyces cerevisiae. Yeast 21, 413–427 (2004).
Garcia, R. et al. The global transcriptional response to transient cell wall damage in Saccharomyces cerevisiae and its regulation by the cell integrity signaling pathway. J. Biol. Chem. 279, 15183–15195 (2004).
Lagorce, A. et al. Genome-wide analysis of the response to cell wall mutations in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 278, 20345–20357 (2003).
Ram, A.F., Wolters, A., Ten Hoopen, R. & Klis, F.M. A new approach for isolating cell wall mutants in Saccharomyces cerevisiae by screening for hypersensitivity to calcofluor white. Yeast 10, 1019–1030 (1994).
Imai, K., Noda, Y., Adachi, H. & Yoda, K. A novel endoplasmic reticulum membrane protein Rcr1 regulates chitin deposition in the cell wall of Saccharomyces cerevisiae. J. Biol. Chem. 280, 8275–8284 (2005).
Ram, A.F. et al. The cell wall stress response in Aspergillus niger involves increased expression of the glutamine: fructose-6-phosphate amidotransferase-encoding gene (gfaA) and increased deposition of chitin in the cell wall. Microbiology 150, 3315–3326 (2004).
Roncero, C., Valdivieso, M.H., Ribas, J.C. & Duran, A. Isolation and characterization of Saccharomyces cerevisiae mutants resistant to calcofluor white. J. Bacteriol. 170, 1950–1954 (1988).
Lussier, M. et al. Large scale identification of genes involved in cell surface biosynthesis and architecture in Saccharomyces cerevisiae. Genetics 147, 435–450 (1997).
Popolo, L. & Vai, M. Defects in assembly of the extracellular matrix are responsible for altered morphogenesis of a Candida albicans phr1 mutant. J. Bacteriol. 180, 163–166 (1998).
Ruiz-Herrera, J., Garcia-Maceira, P., Castillo-Barahona, L.C., Valentin, E. & Sentandreu, R. Cell wall composition and structure of Yarrowia lipolytica transposon mutants affected in calcofluor sensitivity. Antonie Van Leeuwenhoek 84, 229–238 (2003).
Shaw, B.D. & Momany, M. Aspergillus nidulans polarity mutant swoA is complemented by protein O-mannosyltransferase pmtA. Fungal Genet. Biol. 37, 263–270 (2002).
Oka, T. et al. Protein O-mannosyltransferase A of Aspergillus awamori is involved in O-mannosylation of glucoamylase I. Microbiology 151, 3657–3667 (2005).
Gerik, K.J. et al. Cell wall integrity is dependent on the PKC1 signal transduction pathway in Cryptococcus neoformans. Mol. Microbiol. 58, 393–408 (2005).
De Groot, P.W. et al. A genomic approach for the identification and classification of genes involved in cell wall formation and its regulation in S. cerevisiae. Comp. Funct. Genom. 2, 124–142 (2001).
Damveld, R.A. et al. Characterisation of CwpA, a putative glycosylphosphatidylinositol-anchored cell wall mannoprotein in the filamentous fungus Aspergillus niger. Fungal Genet. Biol. 42, 873–885 (2005).
Acknowledgements
We thank A. Franken and M. Arentshorst for their excellent technical assistance, and S. Langeveld for her help with preparing the figures. F.K. acknowledges financial support from the European Union Specific Targeted Research or Innovation Projects (STREP) FungWall program (1540260).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ram, A., Klis, F. Identification of fungal cell wall mutants using susceptibility assays based on Calcofluor white and Congo red. Nat Protoc 1, 2253–2256 (2006). https://doi.org/10.1038/nprot.2006.397
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.397
This article is cited by
-
Secretory expression of β-1,3-glucomannanase in the oleaginous yeast Rhodosporidium toruloides for improved lipid extraction
Bioresources and Bioprocessing (2023)
-
Regression modelling of conditional morphogene expression links and quantifies the impact of growth rate, fitness and macromorphology with protein secretion in Aspergillus niger
Biotechnology for Biofuels and Bioproducts (2023)
-
Magnetotropism: a tropic response of Candida guillemondii by the effect of the oscillating magnetic field of extremely low frequency
Air Quality, Atmosphere & Health (2023)
-
Saccharomyces cerevisiae survival against heat stress entails a communication between CCT and cell wall integrity pathway
Biologia Futura (2023)
-
Sensitivity and control efficacy of a novel complex III inhibitor pyribencarb against Sclerotinia sclerotiorum
Journal of Plant Diseases and Protection (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.