Abstract
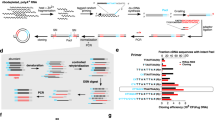
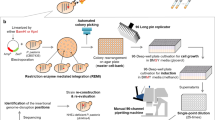
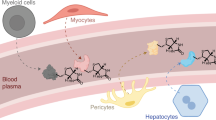
Secreted and cell surface proteins play essential roles in numerous essential biological processes in eukaryotic organisms, but are often more difficult to isolate and identify than proteins that are localized in intracellular compartments. However, several high-throughput 'gene-trap' techniques have been developed to characterize these 'secretomes', including the yeast secretion trap (YST) screen. This method involves fusing cDNA libraries from the tissue or cell type of interest to a yeast (Saccharomyces cerevisiae) invertase reporter gene, transforming the resulting fusion library into an invertase-deficient yeast strain and plating the transformants on a medium containing sucrose as the sole carbon source. A yeast cell with a transgene encoding a secreted or cell surface protein can synthesize a secreted invertase fusion protein that can rescue the mutant, and the plasmid DNA can then be sequenced to identify the gene that encodes it. We describe a recently improved version of this screen, which allows the identification of genes encoding secreted proteins in 1–2 months.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Diehn, M., Eisen, M.B., Botstein, D. & Brown, P.O. Large-scale identification of secreted and membrane-associated gene products using DNA microarrays. Nat. Genet. 25, 58–62 (2000).
Clark, H.F. et al. The secreted protein discovery initiative (SPDI), a large-scale effort to identify novel human secreted and transmembrane proteins: a bioinformatics assessment. Genome Res. 13, 2265–2270 (2003).
Lee, S.-J., Saravanan, R.S., Damasceno, C.M.B., Yamane, H., Kim, B.-D. & Rose, J.K.C. Digging deeper into the plant cell wall proteome. Plant Physiol. Biochem. 42, 979–988 (2004).
De Groot, P.W.J., Ram, A.F. & Klis, F.M. Features and functions of covalently linked proteins in fungal cell walls. Fungal Gen. Biol. 42, 657–675 (2005).
Yin, Q.Y. et al. Comprehensive proteomic analysis of Saccharomyces cerevisiae cell walls. J. Biol Chem. 280, 20894–20901 (2005).
Jamet, E., Canut, H., Boudart, G. & Pont-Lezica, R.F. Cell wall proteins: a new insight through proteomics. Trends Plant. Sci. 11, 33–39 (2006).
von Heijne, G. Signal sequences: the limits of variation. J. Mol. Biol. 184, 99–105 (1985).
Martoglio, B. & Dobbertein, B. Signal sequences: more than just greasy peptides. Trends Cell Biol. 8, 410–415 (1998).
Plath, K., Mothes, W., Wilkinson, B.M., Stirling, C.J. & Rapoport, T.A. Signal sequence recognition in posttranslational protein transport across the yeast ER membrane. Cell 94, 795–807 (1998).
Nielsen, H., Brunak, S. & von Heijne, G. Machine learning approaches for the prediction of signal peptides and other protein sorting signals. Protein Eng. 12, 3–9 (1999).
Ladunga, I. Large-scale predictions of secretory proteins from mammalian genomic and EST sequences. Curr. Opin. Biotechnol. 11, 13–18 (2000).
Fariselli, P., Finocchiaro, G. & Casadio, R. SPEPlip: the detection of signal peptide and lipoprotein cleavage sites. Bioinformatics 19, 2498–2499 (2003).
Bendtsen, J.D., Nielsen, H., von Heijne, G. & Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340, 783–795 (2004).
Hiller, K., Grote, A., Scheer, M., Munch, R. & Jahn, D. PrediSi: prediction of signal peptides and their cleavage positions. Nucleic Acids Res. 32, W375–W379 (2004).
Kall, L., Krogh, A. & Sonnhammer, E.L.L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 338, 1027–1036 (2004).
Small, I., Peeters, N., Legeai, F. & Lurin, C. Predotar: a tool for rapidly screening proteomes for N-terminal targeting sequences. Proteomics 4, 1581–1590 (2004).
Levine, T. & Rabouille, C. Endoplasmic reticulum: one continuous network compartmentalized by extrinsic cues. Curr. Opin. Cell Biol. 17, 362–368 (2005).
Robinson, D.G., Oliviusson, P. & Hinz, G. Protein sorting to the storage vacuoles of plants: a critical appraisal. Traffic 6, 615–625 (2005).
Nickel, W. Unconventional secretory routes: direct protein export across the plasma membrane of mammalian cells. Traffic 6, 607–614 (2005).
Nombela, C., Gil, C. & Chaffin, W.L. Non-conventional protein secretion in yeast. Trends Microbiol. 14, 15–21 (2006).
Bendtsen, J.D., Jensen, L.J., Blom, N., von Heijne, G. & Brunak, S. Feature-based prediction of non-classical and leaderless protein secretion. Prot. Eng. Des. Select. 17, 349–356 (2004).
Tashiro, K. et al. Signal sequence trap: a cloning strategy for secreted proteins and type I membrane proteins. Science 261, 600–603 (1993).
Imai, T. et al. Molecular cloning of a novel T cell-directed CC chemokine expressed in thymus by signal sequence trap using Epstein-Barr virus vector. J. Biol. Chem. 271, 21514–21521 (1996).
Skarnes, W.C., Moss, J.E., Hurtley, S.M. & Beddington, R.S.P. Capturing genes encoding membrane and secreted proteins important for mouse development. Proc. Natl. Acad. Sci. USA 92, 6592–6596 (1995).
Hoffman, C.S. & Wright, A. Fusions of secreted proteins to alkaline phosphatase: an approach for studying protein secretion. Proc. Natl. Acad. Sci. USA 82, 5107–5111 (1985).
Chen, H. & Leder, P. A new signal sequence trap using alkaline phosphatase as a reporter. Nucleic Acids Res. 27, 1219–1222 (1999).
Kojima, T. & Kitamura, T. A signal sequence trap based on a constitutively active cytokine receptor. Nat. Biotechnol. 17, 487–490 (1999).
Klein, R.D., Gu, Q., Goddard, A. & Rosenthal, A. Selection for genes encoding secreted proteins and receptors. Proc. Natl. Acad. Sci. USA 93, 7108–7113 (1996).
Jacobs, K.A. et al. A genetic selection for isolating cDNAs encoding secreted proteins. Gene 198, 289–296 (1997).
Kristoffersen, P., Teichmann, T., Stracke, R. & Palme, K. Signal sequence trap to clone cDNAs encoding secreted or membrane-associated plant proteins. Anal. Biochem. 243, 127–132 (1996).
Goo, J.H., Park, A.R., Park, W.J. & Park, O.K. Selection of Arabidopsis genes encoding secreted and plasma membrane proteins. Plant Mol. Biol. 41, 415–423 (1999).
Belanger, K.D., Wyman, A.J., Sudol, M.N., Singla-Pareek, S.L. & Quatrano, R.S. A signal peptide secretion screen in Fucus distichus embryos reveals expression of glucanase, EGF domain-containing, and LRR receptor kinase-like polypeptides during asymmetric cell growth. Planta 217, 931–950 (2003).
Hugot, K. et al. Coordinated regulation of genes for secretion in tobacco at late developmental stages: association with resistance against oomycetes. Plant Physiol. 134, 1–13 (2004).
Yamane, H., Lee, S.-J., Kim, B.-D., Tao, R. & Rose, J.K.C. A coupled yeast signal sequence trap and transient plant expression strategy to identify genes encoding secreted proteins from peach pistils. J. Exp. Bot. 56, 2229–2238 (2005).
Lee, S.-J., Kelley, B., Damasceno, C.M.B., St. John, B., Kim, B.-S., Kim, B.-D. & Rose, J.K.C. A functional screen to characterize the secretomes of eukaryotic phytopathogens and their hosts in planta. Mol. Plant Microbe Interact. 12, 1368–1377 (2006).
Ito, H., Fukuda, Y., Murata, K. & Kimura, A. Transformation of intact yeast cells treated with alkali cations. J. Bacteriol. 153, 163–168 (1983).
Gietz, R.D. & Woods, R.A. Transformation of yeast by the LiAc/SS carrier DNA/PEG method. Methods Enzymol. 350, 87–96 (2002).
Hoffman, C.S. & Winston, F. A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene. 57, 267–272 (1987).
Akada, R., Murakane, T. & Nishizawa, Y. DNA extraction method for screening yeast clones by PCR. BioTechniques 28, 668–674 (2000).
Galliciotti, G. et al. Signal-sequence trap in mammalian and yeast cells: a comparison. J. Membr. Biol. 183, 175–182 (2001).
Heazlewood, J.L., Tonti-Filippini, J., Verboom, R.E. & Millar, A.H. Combining experimental and predicted datasets for determination of the subcellular location of proteins in Arabidopsis. Plant Physiol. 139, 598–609 (2005).
Yates, J.R., Gilchrist, A., Howell, K.E. & Bergeron, J.J.M. Proteomics of organelles and large cellular structures. Nat. Rev. Mol. Cell Biol. 6, 702–714 (2005).
Perco, P., Rapberger, R., Siehs, C., Lukas, A., Oberbauer, R., Mayer, G. & Mayer, B. Transforming omics data into context: bioinformatics on genomics and proteomics raw data. Electrophoresis 27, 2659–2675 (2006).
Ausubel, F.M. et al. (eds.) Current Protocols in Molecular Biology (John Wiley & Sons, Hoboken, New Jersey, USA, 2006).
Miller, E.M. & Nickoloff, J.A. Escherichia coli electrotransformation. Methods Mol. Biol. 47, 105–114 (1995).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual 2nd edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 1989).
Acknowledgements
Research in this area was supported by grants from the US National Science Foundation Plant Genome Program (DBI-0606595) and the New York State Office of Science, Technology and Academic Research (NYSTAR). S.-J.L. and B.-D.K. of the Center for Plant Molecular Genetics and Breeding Research were also supported by a grant from KOSEF/MOST. The authors would like to thank O.K. Park (Graduate School of Biotechnology, Korea University, Seoul, Korea) for the generous gift of the yeast strain DBYα2445 and T. Isaacson for careful reading of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lee, SJ., Kim, BD. & Rose, J. Identification of eukaryotic secreted and cell surface proteins using the yeast secretion trap screen. Nat Protoc 1, 2439–2447 (2006). https://doi.org/10.1038/nprot.2006.373
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.373
This article is cited by
-
Populus trichocarpa encodes small, effector-like secreted proteins that are highly induced during mutualistic symbiosis
Scientific Reports (2017)
-
A Magnaporthe Avr-pita gene orthologous in Rhizoctonia solani AG1-IA shows characteristics of an effector protein
Australasian Plant Pathology (2015)
-
A screen for over-secretion of proteins by yeast based on a dual component cellular phosphatase and immuno-chromogenic stain for exported bacterial alkaline phosphatase reporter
Microbial Cell Factories (2013)
-
A proteomic analysis of the Pichia pastoris secretome in methanol-induced cultures
Applied Microbiology and Biotechnology (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.