Abstract
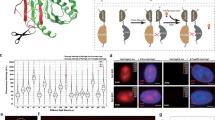
Bimolecular fluorescence complementation (BiFC) analysis enables direct visualization of protein interactions in living cells. The BiFC assay is based on the discoveries that two non-fluorescent fragments of a fluorescent protein can form a fluorescent complex and that the association of the fragments can be facilitated when they are fused to two proteins that interact with each other. BiFC must be confirmed by parallel analysis of proteins in which the interaction interface has been mutated. It is not necessary for the interaction partners to juxtapose the fragments within a specific distance of each other because they can associate when they are tethered to a complex with flexible linkers. It is also not necessary for the interaction partners to form a complex with a long half-life or a high occupancy since the fragments can associate in a transient complex and un-associated fusion proteins do not interfere with detection of the complex. Many interactions can be visualized when the fusion proteins are expressed at levels comparable to their endogenous counterparts. The BiFC assay has been used for the visualization of interactions between many types of proteins in different subcellular locations and in different cell types and organisms. It is technically straightforward and can be performed using a regular fluorescence microscope and standard molecular biology and cell culture reagents.
*Note: In the version of this article initially published online, the article’s page numbers should have been 1278–1286. In addition, the numbered items in Step 2 did not correspond to the format of the HTML version. In Steps 2–13, the locations of the TROUBLESHOOTING headings did not correspond to the Troubleshooting table. These errors have been corrected in the PDF version of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
30 November 2006
In the version of this article initially published online, the article’s page numbers should have been 1278–1286. In addition, the numbered items in Step 2 did not correspond to the format of the HTML version. In Steps 2–13, the locations of the TROUBLESHOOTING headings did not correspond to the Troubleshooting table. These errors have been corrected in the PDF version of the article.
References
Förster, T. 10th Spiers Memorial Lecture. Transfer mechanisms of electronic excitation. Discuss. Faraday Soc. 27, 7–17 (1959).
Hu, C.D., Chinenov, Y. & Kerppola, T.K. Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol. Cell 9, 789–798 (2002).
Brock, R. & Jovin, T.M. Fluorescence correlation microscopy (FCM)-fluorescence correlation spectroscopy (FCS) taken into the cell. Cell. Mol. Biol. 44, 847–856 (1998).
Baudendistel, N., Muller, G., Waldeck, W., Angel, P. & Langowski, J. Two-hybrid fluorescence cross-correlation spectroscopy detects protein-protein interactions in vivo . Chemphyschem. 6, 984–990 (2005).
Petersen, N.O., Hoddelius, P.L., Wiseman, P.W., Seger, O. & Magnusson, K.E. Quantitation of membrane receptor distributions by image correlation spectroscopy: concept and application. Biophys. J. 65, 1135–1146 (1993).
Johnsson, N. & Varshavsky, A. Split ubiquitin as a sensor of protein interactions in vivo . Proc. Natl. Acad. Sci. USA 91, 10340–10344 (1994).
Rossi, F., Charlton, C.A. & Blau, H.M. Monitoring protein-protein interactions in intact eukaryotic cells by beta-galactosidase complementation. Proc. Natl Acad. Sci. USA 94, 8405–8410 (1997).
Pelletier, J.N., Campbell-Valois, F.X. & Michnick, S.W. Oligomerization domain-directed reassembly of active dihydrofolate reductase from rationally designed fragments. Proc. Natl. Acad. Sci. USA 95, 12141–12146 (1998).
Kerppola, T.K. Visualization of molecular interactions by fluorescence complementation. Nat. Rev. Mol. Cell Biol. 7, 449–456 (2006).
Grinberg, A.V., Hu, C.D. & Kerppola, T.K. Visualization of Myc/Max/Mad family dimers and the competition for dimerization in living cells. Mol. Cell. Biol. 24, 4294–4308 (2004).
Hu, C.D. & Kerppola, T.K. Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis. Nat. Biotechnol. 21, 539–545 (2003).
Fang, D.Y. & Kerppola, T.K. Ubiquitin-mediated fluorescence complementation reveals that Jun ubiquitinated by Itch/AIP4 is localized to lysosomes. Proc. Natl Acad. Sci. USA 101, 14782–14787 (2004).
Zamyatnin, A.A. et al. Assessment of the integral membrane protein topology in living cells. Plant J. 46, 145–154 (2006).
MacDonald, M.L. et al. Identifying off-target effects and hidden phenotypes of drugs in human cells. Nat. Chem. Biol. 2, 329–337 (2006).
Demidov, V.V. et al. Fast complementation of split fluorescent protein triggered by DNA hybridization. Proc. Natl. Acad. Sci. USA 103, 2052–2056 (2006).
Ghosh, I., Hamilton, A.D. & Regan, L. Antiparallel leucine zipper-directed protein reassembly: application to the green fluorescent protein. J. Am. Chem. Soc 122, 5658–5659 (2000).
Magliery, T.J. et al. Detecting protein-protein interactions with a green fluorescent protein fragment reassembly trap: Scope and mechanism. J. Am. Chem. Soc. 127, 146–157 (2005).
Guo, Y.J., Rebecchi, M. & Scarlata, S. Phospholipase C beta(2) binds to and inhibits phospholipase C delta(1). J. Biol. Chem. 280, 1438–1447 (2005).
Schmidt, C. et al. Mechanisms of proinflammatory cytokine-induced biphasic NF-kappa B activation. Mol. Cell 12, 1287–1300 (2003).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Shyu, Y.J., Liu, H., Deng, X. & Hu, C.D. Identification of new fluorescent protein fragments for bimolecular fluorescence complementation analysis under physiological conditions. Biotechniques 40, 61–66 (2006).
von der Lehr, N. et al. The F-Box protein Skp2 participates in c-Myc proteosomal degradation and acts as a cofactor for c-Myc-regulated transcription. Mol. Cell 11, 1189–1200 (2003).
Deppmann, C.D., Thornton, T.M., Utama, F.E. & Taparowsky, E.J. Phosphorylation of BATF regulates DNA binding: a novel mechanism for AP-1 (activator protein-1) regulation. Biochem. J. 374, 423–431 (2003).
Rajaram, N. & Kerppola, T.K. Synergistic transcription activation by Maf and Sox and their subnuclear localization are disrupted by a mutation in Maf that causes cataract. Mol. Cell. Biol. 24, 5694–5709 (2004).
Zhu, L.Q. et al. Inhibition of Mist1 homodimer formation induces pancreatic acinar-to-ductal metaplasia. Mol. Cell. Biol. 24, 2673–2681 (2004).
Kanno, T. et al. Selective recognition of acetylated histones by bromodomain proteins visualized in living cells. Mol. Cell 13, 33–43 (2004).
Farina, A. et al. Bromodomain protein Brd4 binds to GTPase-activating SPA-1, modulating its activity and subcellular localization. Mol. Cell. Biol. 24, 9059–9069 (2004).
Jang, M.K. et al. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol. Cell 19, 523–534 (2005).
Remy, I., Montmarquette, A. & Michnick, S.W. PKB/Akt modulates TGF-beta signalling through a direct interaction with Smad3. Nat. Cell Biol. 6, 358–365 (2004).
Remy, I. & Michnick, S.W. Regulation of apoptosis by the Ft1 protein, a new modulator of protein kinase B/Akt. Mol. Cell. Biol. 24, 1493–1504 (2004).
Laricchia-Robbio, L. et al. Partner-regulated interaction of IFN regulatory factor 8 with chromatin visualized in live macrophages. PNAS 102, 14368–14373 (2005).
Hoff, B. & Kuck, U. Use of bimolecular fluorescence complementation to demonstrate transcription factor interaction in nuclei of living cells from the filamentous fungus Acremonium chrysogenum . Curr. Genet. 47, 132–138 (2005).
Blondel, M. et al. Degradation of Hof1 by SCFGrr1 is important for actomyosin contraction during cytokinesis in yeast. EMBO J. 24, 1440–1452 (2005).
Stolpe, T. et al. In planta analysis of protein-protein interactions related to light signaling by bimolecular fluorescence complementation. Protoplasma 226, 137–146 (2005).
Marrocco, K. et al. Functional analysis of EID1, an F-box protein involved in phytochrome A-dependent light signal transduction. Plant J. 45, 423–438 (2006).
Atmakuri, K., Ding, Z.Y. & Christie, P.J. VirE2, a type IV secretion substrate, interacts with the VirD4 transfer protein at cell poles of Agrobacterium tumefaciens . Mol. Microbiol. 49, 1699–1713 (2003).
Tzfira, T., Vaidya, M. & Citovsky, V. Involvement of targeted proteolysis in plant genetic transformation by Agrobacterium . Nature 431, 87–92 (2004).
Loyter, A. et al. The plant VirE2 interacting protein 1. A molecular link between the Agrobacterium T-complex and the host cell chromatin? Plant Physiol. 138, 1318–1321 (2005).
Li, J.X., Krichevsky, A., Vaidya, M., Tzfira, T. & Citovsky, V. Uncoupling of the functions of the Arabidopsis VIN protein in transient and stable plant genetic transformation by Agrobacterium . Proc. Natl. Acad. Sci. USA 102, 5733–5738 (2005).
Lacroix, B., Vaidya, M., Tzfira, T. & Citovsky, V. The VirE3 protein of Agrobacterium mimics a host cell function required for plant genetic transformation. EMBO J. 24, 428–437 (2005).
Hynes, T.R., Mervine, S.M., Yost, E.A., Sabo, J.L. & Berlot, C.H. Live cell imaging of G(s) and the beta(2)-adrenergic receptor demonstrates that both alpha(s) and beta(1)gamma(7) internalize upon stimulation and exhibit similar trafficking patterns that differ from that of the beta(2)-adrenergic receptor. J. Biol. Chem. 279, 44101–44112 (2004).
Hynes, T.R. et al. Visualization of g protein beta gamma dimers using bimolecular fluorescence complementation demonstrates roles for both beta and gamma in subcellular targeting. J. Biol. Chem. 279, 30279–30286 (2004).
Takahashi, Y. et al. Loss of Bif-1 suppresses Bax/Bak conformational change and mitochondrial apoptosis. Mol. Cell. Biol. 25, 9369–9382 (2005).
Blumenstein, A. et al. The Aspergillus nidulans phytochrome FphA represses sexual development in red light. Curr. Biol. 15, 1833–1838 (2005).
Tsuchisaka, A. & Theologis, A. Heterodimeric interactions among the 1-amino-cyclopropane-1-carboxylate synthase polypeptides encoded by the Arabidopsis gene family. Proc. Natl. Acad. Sci. USA 101, 2275–2280 (2004).
Ozalp, C., Szczesna-Skorupa, E. & Kemper, B. Bimolecular fluorescence complementation analysis of cytochrome P4502C2, 2E1, and NADPH-cytochrome P450 reductase molecular interactions in living cells. Drug Metabolism and Disposition 33, 1382–1390 (2005).
Szczesna-Skorupa, E. & Kemper, B. BAP31 is involved in the retention of cytochrome P4502C2 in the endoplasmic reticulum. J. Biol. Chem. 281, 4142–4148 (2006).
deVirgilio, M., Kiosses, W.B. & Shattil, S.J. Proximal, selective, and dynamic interactions between integrin alpha II beta 3 and protein tyrosine kinases in living cells. J. Cell Biol. 165, 305–311 (2004).
Niu, T.K., Pfeifer, A.C., Lippincott-Schwartz, J. & Jackson, C.L. Dynamics of GBF1, a brefeldin A-sensitive Arf1 exchange factor at the Golgi. Mol. Biol. Cell 16, 1213–1222 (2005).
Nyfeler, B., Michnick, S.W. & Hauri, H.P. Capturing protein interactions in the secretory pathway of living cells. Proc. Natl. Acad. Sci. USA 102, 6350–6355 (2005).
Giese, B. et al. Dimerization of the cytokine receptors gp130 and LIFR analysed in single cells. J. Cell Sci. 118, 5129–5140 (2005).
Rackham, O. & Brown, C.M. Visualization of RNA-protein interactions in living cells: FMRP and IMP1 interact on mRNAs. EMBO J. 23, 3346–3355 (2004).
Ye, H.H., Choi, H.J., Poe, J. & Smithgall, T.E. Oligomerization is required for HIV-1 nef-induced activation of the Src family protein-tyrosine kinase, Hck. Biochemistry 43, 15775–15784 (2004).
Schmidt, U., Richter, K., Berger, A.B. & Lichter, P. In vivo BiFC analysis of Y14 and NXF1 mRNA export complexes: preferential localization within and around SC35 domains. J. Cell Biol. 172, 373–381 (2006).
Bracha-Drori, K. et al. Detection of protein-protein interactions in plants using bimolecular fluorescence complementation. Plant J. 40, 419–427 (2004).
Walter, M. et al. Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. Plant J. 40, 428–438 (2004).
Hackbusch, J., Richter, K., Muller, J., Salamini, F. & Uhrig, J.F. A central role of Arabidopsis thaliana ovate family proteins in networking and subcellular localization of 3-aa loop extension homeodomain proteins. Proc. Natl. Acad. Sci. USA 102, 4908–4912 (2005).
Diaz, I., Martinez, M., Isabel-LaMoneda, I., Rubio-Somoza, I. & Carbonero, P. The DOF protein, SAD, interacts with GAMYB in plant nuclei and activates transcription of endosperm-specific genes during barley seed development. Plant J. 42, 652–662 (2005).
Shimizu, H. et al. LIP19, a basic region leucine zipper protein, is a fos-like molecular switch in the cold signaling of rice plants. Plant Cell Physiol. 46, 1623–1634 (2005).
Abe, M. et al. FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309, 1052–1056 (2005).
Maple, J., Aldridge, C. & Moller, S.G. Plastid division is mediated by combinatorial assembly of plastid division proteins. Plant J. 43, 811–823 (2005).
MacDonald, M.L. et al. Identifying off-target effects and hidden phenotypes of drugs in human cells. Nat. Chem. Biol. 2, 329–337 (2006).
Ding, Y.H., Liu, N.Y., Tang, Z.S., Liu, J. & Yang, W.C. Arabidopsis GLUTAMINE-RICH PROTEIN23 is essential for early embryogenesis and encodes a novel nuclear PPR motif protein that interacts with RNA polymerase II subunit III. Plant Cell 18, 815–830 (2006).
Wang, K.Z.Q. et al. TRAF6 activation of PI 3-kinase-dependent cytoskeletal changes is cooperative with Ras and is mediated by an interaction with cytoplasmic Src. J. Cell Sci. 119, 1579–1591 (2006).
Xu, X.M. & Moller, S.G. AtSufE is an essential activator of plastidic and mitochondrial desulfurases in Arabidopsis . EMBO J. 25, 900–909 (2006).
Chen, C.D., Oh, S.Y., Hinman, J.D. & Abraham, C.R. Visualization of APP dimerization and APP-Notch2 heterodimerization in living cells using bimolecular fluorescence complementation. J. Neurochem. 97, 30–43 (2006).
Hu, C.-D. & Kerppola, T.K. in Protein-Protein Interactions (eds. Adams, P. & Golemis, E.) 673–693 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2005).
Liu, H. et al. Mutual regulation of c-Jun and ATF2 by transcriptional activation and subcellular localization. EMBO J. 25, 1058–1069 (2006).
Pazhouhandeh, M. et al. F-box-like domain in the polerovirus protein P0 is required for silencing suppressor function. Proc. Natl. Acad. Sci. USA 103, 1994–1999 (2006).
Cascales, E., Atmakuri, K., Liu, Z., Binns, A.N. & Christie, P.J. Agrobacterium tumefaciens oncogenic suppressors inhibit T-DNA and VirE2 protein substrate binding to the VirD4 coupling protein. Mol. Microbiol. 58, 565–579 (2005).
Stains, C.I., Porter, J.R., Ooi, A.T., Segal, D.J. & Ghosh, I. DNA sequence-enabled reassembly of the green fluorescent protein. J. Am. Chem. Soc. 127, 10782–10783 (2005).
Zhang, J., Campbell, R.E., Ting, A.Y. & Tsien, R.Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 3, 906–918 (2002).
Acknowledgements
I thank C.-D. Hu for his participation in the design and implementation of the BiFC assay in mammalian cells and all members of the Kerppola laboratory for their contributions to the improvement and adaptation of the BiFC approach.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Kerppola, T. Design and implementation of bimolecular fluorescence complementation (BiFC) assays for the visualization of protein interactions in living cells. Nat Protoc 1, 1278–1286 (2006). https://doi.org/10.1038/nprot.2006.201
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.201
This article is cited by
-
Identification of small molecules capable of enhancing viral membrane fusion
Virology Journal (2023)
-
Genome-wide CRISPR–Cas9 Knockout Screening Reveals a TSPAN3-mediated Endo-lysosome Pathway Regulating the Degradation of α-Synuclein Oligomers
Molecular Neurobiology (2023)
-
A Nodal enhanced micropeptide NEMEP regulates glucose uptake during mesendoderm differentiation of embryonic stem cells
Nature Communications (2022)
-
Interaction network among factors involved in heterocyst-patterning in cyanobacteria
Molecular Genetics and Genomics (2022)
-
Nucleus-translocated mitochondrial cytochrome c liberates nucleophosmin-sequestered ARF tumor suppressor by changing nucleolar liquid–liquid phase separation
Nature Structural & Molecular Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.