Abstract
Physiological modifications in near weightlessness, as experienced by astronauts during space flight, have been the subject of numerous studies. Various animal models have been used on space missions or in microgravity simulation on ground to understand the effects of gravity on living animals. Here, we used the zebrafish larvae as a model to study the effect of microgravity simulation on bone formation and whole genome gene expression. To simulate microgravity (sim-μg), we used two-dimensional (2D) clinorotation starting at 5 days post fertilization to assess skeletal formation after 5 days of treatment. To assess early, regulatory effects on gene expression, a single day clinorotation was performed. Clinorotation for 5 days caused a significant decrease of bone formation, as shown by staining for cartilage and bone structures. This effect was not due to stress, as assessed by measuring cortisol levels in treated larvae. Gene expression results indicate that 1-day simulated microgravity affected musculoskeletal, cardiovascular, and nuclear receptor systems. With free-swimming model organisms such as zebrafish larvae, the 2D clinorotation setup appears to be a very appropriate approach to sim-μg. We provide evidence for alterations in bone formation and other important biological functions; in addition several affected genes and pathways involved in bone, muscle or cardiovascular development are identified.
Similar content being viewed by others
Introduction
Physiological modifications in near weightlessness, as experienced by astronauts during space flight, have been the subject of numerous studies. A compilation of human responses to prolonged exposure to space conditions indicates a loss of bone mineral density of about 1% per month.1 Prolonged bed rest studies confirmed this bone loss without any substantial gender differences.2 Animal models have also been used to gain deeper insight into the molecular mechanisms of adaptation to microgravity,3 among which are various fish species.4–6 Fishes directly respond to microgravity by displaying a kinetotic swimming behavior,5 probably due to conflicting sensory input reminiscent of motion sickness in humans. After some time, this behavior disappears, but further studies have extensively illustrated the effect of decreased gravity on the growth of the gravity-sensing organs, the otoliths.5,7
Until now, it was, however, unclear whether aquatic organisms, such as fishes, would elicit a physiological response to altered gravity conditions, in addition to sensing the modification and adapting the sensing organs. Recently, we chose to use small fish models, such as medaka or zebrafish, to study physiological changes in altered gravity conditions3,4,8,9 and we showed that zebrafish larvae experiencing hypergravity (3 g) between 1 and 5 days post fertilization (dpf) or between 5 and 10 dpf presented a significantly increased bone formation.9 Here, we used clinorotation to simulate microgravity (sim-μg) and studied its effect on bone formation, as well as the effect on whole genome gene expression during the first day of sim-μg treatment. We show for the first time that clinorotation between 5 and 10 dpf significantly decreases bone formation in zebrafish larvae. Further, we also analyzed the modifications in gene expression caused by exposure to sim-μg for 1 day and we identified several genes and pathways whose expression is altered after clinorotation.
Results
Effects of sim-μg on cartilage and bone formation: 5 days in clinorotation versus controls
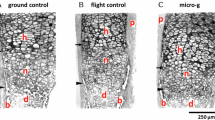
Larvae of 5 dpf were maintained in clinorotation for another 5 days. At the end (10 dpf), controls and sim-μg-exposed larvae were stained with Alcian blue to observe the cartilage and Alizarin red to visualize calcium deposition by osteogenic cells (see Figure 1a,b). The cartilage structures are well-formed, complete and morphologically similar to the respective controls (Figure 1c,d). In contrast, bone formation was clearly decreased in the larvae submitted to clinorotation relative to their controls (Figure 1e,f) and several bone structures, such as anguloarticular, branchiostegal ray2, ceratohyal, and dentary are absent.
Effects in 10 dpf zebrafish larvae after 5 days sim-μg in clinorotation (a) Schematic representation of the different cranial bone elements revealed by alizarin red staining in 9–10 dpf zebrafish larvae and (b) the landmarks used for morphometric analysis. The landmarks used in morphometric analysis are anguloarticular (aa), anterior (an), branchiostegal ray1 (br1), entopterygoid (en), maxilla (m), notochord (n), opercle (o), and parasphenoid (p). Note that the parasphenoid is a triangular bone defined by its anterior summit (a) and two posterior summits (b,c). Additional bone elements that were evaluated for extent of ossification are: branchiostegal ray2 (br2), cleithrum (c), ceratobranchial 5 (cb), ceratohyal (ch), dentary (d), and hyomandibular (hm) (from ref. 9). (c,d) Alcian blue staining of cartilage. (e,f) Alizarin red staining of bone structures. Compared with controls (C), no effect was observed on cartilage development after 5 days clinorotation (D). In contrast, relative to control (E), a clear decrease of bone formation was seen after 5 days in clinostat (F). Scale bars=250 μm. (g) Morphometry. A significant increase of the distance between the anterior part of the larvae and the parasphenoid summit is observed. The distances are measured in pixels. Mean±s.d. and t-test analysis were calculated for each measure on at least 20 individuals. (h) The area covered by the parasphenoid is decreased on clinostat exposure. (i) Extent of bone formation: Bone development is classified for each element into different categories: Absent (no structure present; red), early ossification (beginning of the bone ossification; yellow), advanced ossification (the structure is present and already developed as the control; green), and over ossification (the structure is more developed compared with the control; purple). Cumulated frequencies in % are represented for each element. As no significant difference was observed for paired structures between left and right (up and down), their scores have been combined. Statistical analysis was performed by X2 of Pearson and a logistic regression. (i) Cumulated frequency after 5 days in clinostat. (j) To obtain global scores, the scores attributed to each element were added up for each individual larva. Mean±s.d. and t-test analysis was obtained on at least 20 individuals. **P<0.005 and ***P<0.001. dpf, days post fertilization.
The extent of bone formation was analyzed more precisely by using qualitative and quantitative descriptions of the images of the stained larvae by applying two approaches.9
The morphometric approach
Each image was manually annotated by defining specific landmarks indicating the positions of the different skeletal structures. Symmetric structures, such as the maxilla (m), entopterygoid (en), branchiostegal ray (br), or opercle (o) were distinguished by the suffix “up” or “down” (see Figure 1a,b). The software then calculates all the distances between the selected landmarks to obtain a morphometric description of the head skeleton. In the cartilage skeleton, larvae subjected to clinorotation did not reveal any significant modifications (data not shown). In contrast, larvae stained with alizarin red revealed a clearly increased distance of the parasphenoid summit (pa) and the anterior (an) part of the larvae (Figure 1g, Supplementary Table S2), probably due to the significant decrease of the parasphenoid (p) area (Figure 1h, Supplementary Table S2). Note that not all of the measures were possible due to the absence of several structures in >60% larvae such as the anguloarticular (aa) and ceratohyal (ch) (see also below).
The staining intensity evaluation approach
According to its developmental status (absent, early, advanced, or over ossification), a score, from 0 for absent to 3 for advanced ossification, is given to each structure. The frequency distribution of these scores reveals that exposure to clinorotation lead to a significant decrease in ossification of all the structures (Figure 1i, Supplementary Table S3). The branchiostegal ray1 and the entopterygoid structures were absent in ~25 % of the treated larvae, while they presented advanced ossification in 100% of the control larvae (Figure 1i). The anguloarticular and the ceratohyal were absent in about 60% of the treated larvae, compared with 25% or 15%, respectively, in the controls. Absence of the branchiostegal ray2 switched from 25% in controls to about 80% after clinorotation. For all structures, the frequency of larvae presenting advanced ossification significantly decreased on clinorotation. Assigning a global score for all structures within each larva confirmed that 5 days of clinorotation treatment produced a significantly decreased ossification (Figure 1j).
Sim-μg and stress in larvae
Stress can induce bone loss;10–13 therefore we decided to evaluate the stress status of 6 dpf larvae after 1-day exposure to sim-μg. To this end, we determined the whole body cortisol levels in 15 larvae directly after sacrifice. This analysis has been performed through subsequent adjustments:
(i) To collect and kill the larvae, we compared two different methods, shortly defined as low-tricaine and high-tricaine. Low-tricaine: to cause a rapid anesthesia followed after 2–3 min by death by adding tricaine to a final concentration of 0.04 g/l;14 high-tricaine: to cause immediate death by collecting larvae in a reduced volume of E3 medium and adding 1.6 g/l of tricaine. Since we observed significantly higher cortisol levels in larvae killed with low-tricaine (Figure 2a), probably due to the acute stress induced by anesthesia, we applied the high-tricaine method in all experiments.
(ii) To induce acute stress in zebrafish larvae, we performed positive control experiments by using two different known procedures: the first consists in intense agitation for 30 s in 5 ml medium followed by a 5-min resting period15 before sacrifice at high-tricaine, while the second exposes the larvae to a 1.75 g/l NaCl solution for 5 min before leaving them for 5 min recovery in E3 (ref. 16) and then sacrifice. As expected, both stress conditions lead to a significant increase in cortisol levels, which interestingly were similar to the levels observed under low-tricaine conditions (Figure 2a).
(iii) Finally, careful measurements of the cortisol levels indicated no differences between the larvae subjected to clinorotation for 1 day as compared with their respective controls (Figure 2b). These results demonstrate that the larvae placed into this microgravity simulator are not stressed compared with their respective controls. Thus, any modification observed is most likely related to the effect of sim-μg and not stress.
Effects of sim-μg on gene expression: 1 day clinorotation versus controls
To obtain a global view of the physiological changes caused by sim-μg, we performed a microarray whole genome expression analysis. We compared 6 dpf control larvae with larvae growing in clinorotation for 1 day, i.e., from 5 dpf to 6 dpf, to capture early-regulatory events rather than secondary regulations, leading ultimately to the observed modulations of bone formation at 10 dpf. Four independent experiments were carried out, using 60 larvae for each experimental condition. Due to the small volume available in the rotating tubes in clinorotation, only 15 larvae were run in parallel in three tubes; thus each control or rotated sample consisted of a pool from 4 different experiments to reach a sample size of 60 larvae. Total RNA was extracted from 6 dpf control larvae and larvae that experienced sim-μg, reverse transcribed into complementary DNA, and used for gene expression microarray analysis.
A list of genes affected more than 1.4-fold (|log2 fold change|>0.49) was extracted and introduced into the Ingenuity Pathway Analysis software (IPA) for further analysis. In total, 208 genes were significantly affected in the clinorotation experiment, of which 66 genes were annotated in IPA. The full list of these genes is given in Supplementary Table S4. Validation by reverse transcription-quantitative PCR of five genes selected from the list demonstrated the reliability of the microarray data (Figure 3a).
Genes whose expression is affected by clinorotation for 24 h starting at 5 dpf. (a) Fold change values on clinorotation treatment for selected genes. The fold change after clinorotation between 5 and 6 dpf is given as deduced from the microarray data and the RT–qPCR confirmation experiments. For microarray data, only significant fold-change values are shown, while for RT–qPCR bold type indicates significant changes in expression (P<0.05). (b) Diseases and Biological functions affected by clinorotation. IPA analysis of the list of genes affected at 6 dpf after 1-day clinorotation compared with controls. The columns represent the −log(P-value) for significance that the list of genes affects the indicated biological function. (c) Upstream regulators, as suggested by the lists of genes affected at 6 dpf after 1-day clinorotation as compared with the corresponding controls. Potential upstream regulators known to modify expression of genes in the gene list were identified, the numbers given are the z-scores indicating significance of the upstream regulator, negative z-scores correspond to predicted decreased activity, while positive z-scores correspond to a predicted increased activity of the proposed regulator. (d) Genes connected to the upstream regulator CREB1 as predicted by Ingenuity Pathway Analysis. CREB1 activity is predicted to be downregulated in clinorotation condition (blue color). Blue or orange lines indicate, respectively, inhibition or activation of expression consistent with the prediction, gray arrow indicates that no information is available (inconsistent findings would be in yellow). Arrows indicate an interaction activating, while stop-lines indicate an interaction inhibiting expression of the target gene. Red overlay color indicates increased gene expression, while green overlay indicates decreased gene expression in the corresponding experiment.
In general, it appears that the number of significantly affected genes is relatively low, indicating that sim-μg has no major immediate impact on general physiology. The most highly affected gene in clinorotation (Supplementary Table S4) is AXIN2 (log2 fold −3.48), indicating a downregulation of the Wnt pathway, while the most highly upregulated gene is HES1 that is involved in NOTCH signaling.17 Other affected genes are IGF2R, a component of the insulin-like pathway, or the ryanodin receptor RYR2. Affected transcriptional regulator genes are also represented, such as E2F2, FOSB, or the bone-specific factor HOXB9.
IPA was used to identify the biological functions and regulatory pathways that were affected by sim-μg. Among the ‘canonical pathways’ affected (Supplementary Table S5), the retinoid X receptor RXR is prominent in its common role for FXR/RXR, VDR/RXR, and FXR/RXR signaling, while other pathways regulated by IL-6, Notch, VDR, Hif1ß, LPS/IL-1, and Notch were also affected. Classification of the list of affected genes according to their involvement in specific ‘Disease and Biological Functions’ revealed their role in morphology, size, and resorption of bone, as well as the quantity of osteoclasts, bone, and blood cells. When considering the more generic ‘Disease and Biological Function’ terms according to their relevance (Figure 3b), ‘Connective Tissue Disorders’, Skeletal and Muscular Disorders, and ‘Connective Tissue Development and Function’ ranked on position 2, 5, and 11, respectively. Finally, we analyzed the data set for putative upstream regulators and the predicted change in activity of these regulators (Figure 3c) and we identified CREB1 and CREM as putative affected regulators, but also TCR, IL-12, IL-6, and the bone-specific transcription factor RUNX2. The observed changes in gene expression are indeed consistent with a decrease in CREB1 activity in clinorotation (Figure 3d).
One important aspect of our study is the fact that we investigated gene expression using mRNA from the entire larvae. We first filtered the gene list according to their involvement in musculoskeletal or cardiovascular function or disease (Supplementary Table S7), we identified 8 genes common to both systems, 9 genes related to the musculoskeletal function, and 10 genes more specific to the cardiovascular system. When we focused on individual organ systems by filtering the affected gene set against available databases of genes involved in specific functions (gene ontology annotations of human or mouse gene orthologs using the IPA knowledge base), networks of regulatory interactions could be constructed for each system. Specific to bone, a small pathway centered around FGF2 (increased expression) and including FOSB, GAB2, and E2F2 (decreased expression) was identified. In muscle, FOSB is absent, but FGF2 is additionally connected to MCL1 through GAB2. Further, the increased expression of CYP19A1, required for estrogen synthesis, and its effect on lipoprotein metabolism was also pointed out. In the cardiovascular system, all these genes and pathways are represented
Discussion
We used zebrafish larvae as a model system to explore changes in skeletal formation and gene expression under sim-μg conditions. Due to their small size, a relatively large number of larvae can be maintained in reasonably small volumes of medium. Thus, devices traditionally used for studying the effects of sim-μg on microorganisms or cell cultures can be adapted to the study of this vertebrate model system. In this study, we used clinorotation to sim-μg.
We first concentrated on the effects of sim-μg on cartilage/bone formation. In zebrafish, since ossification starts in the head skeleton at 3 dpf, with extensive bone mineralization observed beyond 4–5 dpf, we chose 5 dpf as starting point for the sim-μg experiments. Simulation was continued for some days to observe the effect on skeletal formation at 10 dpf. Changes were assessed using the previously validated morphometric and ‘extent of ossification’ methods. In the cartilage skeleton, no changes in morphology due to sim-μg were observed (Figure 1c,d). This absence of major effects on head cartilage formation during this period was previously also shown in hypergravity,9 indicating that the cranial cartilage morphology is not affected by mechanical constraints, at least past a certain stage. In this context, it is interesting to note that also inhibition of the Fgf or Bmp signaling pathways in zebrafish larvae older than 2 dpf did not affect cartilage formation.18–20
In contrast, the morphology of the head bone skeleton was affected by clinorotation causing a significant decrease of the parasphenoid area, in line with the general decrease in ossification in these conditions (see below). Such a narrowing of the head skeleton was not observed for the cartilage elements, suggesting that the dermal skeleton might be more responsive to changes in environment at these stages. Clinorotation for 5 days caused a significant decrease in ossification of individual bone elements, and on the global score. The continuous rotation of the water column during clinorotation allowed a natural, mainly resting behavior. Indeed, we observed that the mostly immobile larvae were following the rotating movement, while swimming larvae tended to compensate the rotating environment to keep their level. We observed that clinorotation starting at 5 dpf, when ossification is ongoing, causes a significant decrease of the calcified parasphenoid area and a general decrease of calcification in all major cranial bones at 10 dpf, without major changes in the general morphology. This result is consistent with previous experiments exposing mouse fetal long bones for 4 days to space conditions21 that revealed decreased mineralization, but no change in growth or collagen synthesis. Taken together, we conclude that clinorotation is an appropriate approach to study the effects of sim-μg on free-swimming aquatic organisms. However, the final proof will require comparing the simulation results against real in-flight data.
The observation that bone formation was decreased on clinorotation raised the question concerning a possible involvement of systemic stress in this effect. Loss of bone mineral density was shown in US military during combat missions10 or during glucocorticoid treatment of inflammatory diseases.11,13 Similarly, increased bone resorption, which was partly attributed to stress, was shown in Atlantic bluefin tuna (Thynnus thynnus) when reared in captivity.12 On the other hand, the level of systemic stress during space missions was assessed by performing cortisol measurements before, during, and post flight. The cortisol levels did not significantly change in the cosmonauts.22 Another study found identical cortisol levels in animals in space versus Earth control.23 These results suggest that the bone loss observed in astronauts is not due to stress, as further substantiated by in-flight skeletal in vitro mineralization studies using mouse metatarsal long bones.24 Nevertheless, we could not exclude that the zebrafish larvae would experience stress when placed into the microgravity simulation device. No increase in cortisol levels was, however, observed in larvae after undergoing a 1-day simulation in clinorotation, strongly arguing against the possibility that the decreased bone formation in clinorotation may be due to systemic stress.
The clinorotation results are also in line with the well-established bone loss experienced by astronauts in space or bed rest studies,1,2 and with the space experiments performed on rats25–28 or mice.29 Microgravity caused effects ranging from decreased trabecular numbers27,29 and thickness27 in tibia to a decreased mineral content and number of osteoblasts.26,28,30 In astronauts, bone formation markers (type I procollagen propeptide and bone alkaline phosphatase) decreased, while bone resorption markers such as the procollagen C-telopeptide increased during space flight.22,31 The decreased bone formation due to sim-μg, as well as the increased bone formation due to hypergravity that was previously reported,9 strongly indicate that skeleton formation in zebrafish between 5 and 10 dpf is a good model to study gravitational effects on bone metabolism. One important difference is, however, evident: only weight-bearing bones are significantly affected by microgravity in humans and rodents,1,27,29 while most cranial bone elements appear to be affected by gravitational changes in zebrafish larvae. It is unlikely that these effects result from changes in muscle strain, as is generally accepted for the mammalian weight-bearing bones, suggesting the existence of a response to microgravity independent of mechanical strain; further experiments will be required to better understand this general sensitivity to gravitational conditions of the developing zebrafish bones.
Finally, we chose to analyze the changes in gene expression caused by sim-μg in clinorotation after only 1 day of treatment, as we were mainly interested in early-regulatory events rather than in secondary effects. In general, the number of affected genes was low. Among the genes affected by clinorotation, a small molecular network could be constructed containing the FOSB, FGF2, E2F2, GAB2, and MCL1 genes (Figure 4). Compared with a network that was previously described of genes affected by 3 g hypergravity relative to 1 g,9 the transcription factor complex AP1 attracted our attention. This complex, formed by members of the JUN and FOS families, is well-known to control cell proliferation, differentiation, and transformation. It is regulated by the GADD45B factor that mediates cellular stress response through the p38/JNK pathway32 and is involved in hematopoiesis and immune response.33 On the other hand, compared with 1 g, microgravity decreases the expression of FOSB, while moderate 3 g hypergravity leads to increased expression of FOSB and its homolog FOS.9 Expression of truncated versions of FosB was shown in mice to cause osteosclerosis and increased expression of the osteoblast marker gene Runx2.24 Decreased expression of c-Fos in microgravity was shown in osteoblast cells,34,35 while in murine carcinoma cells decreased induction of c-Fos and c-Jun was shown in microgravity.36,37 Taken together, these observations point to a central network comprising members of the FOS–JUN factors whose global expression may serve as an indicator for gravity conditions. Another pathway that was highlighted by the study of affected genes was the signaling through CREB1/CREM transcription factors, typically involved in response to cAMP. Other genes appear to be more specifically regulated by different gravity conditions. Here, we show that the bone-related HES1 (ref. 38) gene is strongly upregulated (Supplementary Table S4), while its homolog HES5 was upregulated in hypergravity. Both genes, coding for helix–loop–helix transcription factors, are involved in the control of neural stem cell differentiation,39 HES5 is regulated during cartilage differentiation,40 while HES1 is involved in digestive system development.41 Other genes specifically affected by clinorotation are GAB2, a GRB2-associated-binding protein involved in signal transduction through tyrosine kinase (RTK) or non-RTK receptors42 and required for allergic reactions, mast cell growth in bone marrow, bone homeostasis,43 and heart function,42 the BCL-2-related MCL1 involved in control of cell survival,44 and E2F2 controlling the cell cycle.45,46 Finally, the sex steroid aromatase gene CYP19A1, converting androgens into estrogens,47 is upregulated after clinorotation.
Networks of genes involved in bone, muscular, or cardiovascular systems and affected by clinorotation for 1 day. The list of genes affected by clinorotation was filtered according to the described function for their mammalian homologs in the different physiological system using IPA. The color overlay indicates the fold change relative to the respective controls (upregulated genes in red, downregulated genes in green) observed, respectively in (a) bone, (b) muscle, and (c) cardiovascular system. IPA, Ingenuity Pathway Analysis software.
Few data have been published concerning whole genome gene expression studies in microgravity.48–51 Comparing the gene list derived from zebrafish larvae submitted to clinorotation for 1 day, no gene commonly affected was found in human adipose tissue-derived mesenchymal stem cells kept for 14 days in random positioning machine,50 human umbilical vascular endothelial cells exposed to space conditions for 7 days51 or in the skin of mice that spent 3 months in space.48 Murine 2T3 osteoblast precursor cells cultured for 3 days on an random positioning machine revealed decreased expression of IGF-1 and IGF-2,49 possibly related to the decreased expression of IGF2R observed here. Other genes whose expression was affected in 2T3 cells, such as decreased expression of BMPs, PTHR or RUNX2, may reflect secondary events after 3 days in microgravity, rather than the regulatory events that were investigated here.
In conclusion, we show here for the first time that zebrafish larvae experiencing sim-μg by clinorotation in early-life stages between 5 and 10 dpf exhibit decreased bone formation in the head, in sharp contrast to the previously described increase in bone formation after exposure to hypergravity.9 This decrease in skeleton ossification was not preceded by decreased cartilage formation, and was not due to increased stress. Finally, we analyzed whole genome gene expression after 1 day sim-μg in a clinostat and propose a regulatory gene network centred around the FOS–JUN transcription factor complex as a general indicator for gravity conditions. The cAMP-responsive CREB1/CREM pathway was also affected. The relevance of the sim-μg paradigm can only be conclusively shown by performing such experiments in real, near weightlessness conditions in an orbital spaceflight.
Materials and methods
Animals procedures
Zebrafish (Danio rerio) were maintained and treated under standard conditions14 in the GIGA zebrafish facility (Liège, Belgium; licence LA2610359) as described previously.9 Wild-type embryos were used and staged according to.52 All protocols for experiments were evaluated by the Institutional Animal Care and Use Committee of the University of Liège and approved under the file numbers 568, 1074, and 1264 (licence LA 1610002).
Microgravity simulation experiments using clinorotation
The clinostat device (benchtop 2D clinostat)53 allows parallel positioning of six horizontal tubes, three of which are rotating at a precisely controlled constant speed of 60 r.p.m., while three others are kept immobile to serve as control. As the tubes have to be hermetically closed during the experiment, we carried out preliminary experiments to determine a density of 1 larva per ml as maximal to avoid effects on health and behavior (slow movements). In each tube, five larvae of 5 dpf were placed into 5 ml of freshly prepared and oxygenated E3 medium (5 mmol/l NaCl, 0.17 mmol/l KCl, 0.33 mmol/l CaCl2, 0.33 mmol/l MgSO4, 0.00001 % Methylene Blue). The medium was changed every 24 h to renew the oxygen level. The clinostat was placed in the zebrafish facility (room temperature 26 °C) and covered by an aluminum foil to keep the larvae in the dark, isolated from possible visual clues concerning the rotation. This procedure resulted in an increase of the temperature to 28 °C. Visual inspection just after setup revealed that the water column within the rotating tubes followed the movement without turbulence, as well as the immobile larvae. In an attempt to generate an additional control experiment, we also used the same device in which the rotating tubes were vertically oriented, to test the effect of rotation without the sim-μg effect. Unfortunately, we observed that in these experiments all the larvae were staying in the bottom of the tube, thus possibly creating a hypoxic environment. Therefore we decided that the best controls for the clinorotation experiments were the horizontal non-rotating tubes run in parallel; we did not further follow up vertical rotation.
Cortisol measurement
Cortisol was quantified using a cortisol enzyme immunoassay (EIA) kit (Cayman Chemical, Ann Arbor, MI, USA). For each condition, 15 larvae at 6 dpf were used in triplicates in E3 at 28 °C. After treatment, the larvae were killed, washed in cold PBS (pH=7.4) and frozen in liquid nitrogen for storage at −80 °C. The larvae were thawed and homogenized in cold PBS using a Potter (Thomas Scientific, Swedesboro, NJ, USA). Five hundred microliters of homogenate was used for cortisol extraction with 3 ml of diethyl ether, which was repeated three times before evaporating the ether to dryness in a bath at 45 °C under a nitrogen flow.15 Eluates were dissolved in 500 μl EIA buffer. Normalization to the protein concentration of each homogenate was performed using the Micro BCA Protein Assay kit (Thermo Scientific, Pierce Biotechnology, Rockford, IL, USA). The data were adjusted to the cortisol extraction efficiency (=93%); the detection limit was 12 pg cortisol per ml.15,16
Staining methods
Acid-free protocols were adapted54 to perform Alcian blue (8 GX Sigma-Aldrich, Diegem, Belgium) staining of cartilage structures 55 and Alizarin red S (Sigma-Aldrich) staining of calcified structures. Images of stained larvae (n=20–30 larvae) were obtained on a binocular (cell B software, Olympus, Berchem, Belgium).
Image analysis
Image analysis was performed on the pictures of larvae stained with Alcian blue for cartilage or Alizarin red for bone as previously described.9 Briefly, for morphometric analysis, images were uploaded into the CYTOMINE (Angleur, Belgium) environment56 and manually annotated by positioning landmarks in the cartilage or bone skeleton (Figure 1a,b) as previously defined in the CYTOMINE ontology. The program computes all the distances (in pixels) and angles (in radian) for any two points of interest, which were exported into an Excel file. Statistics were performed using GraphPad Prism5 (La Jolla, CA, USA). A t-test was used for control versus treatment experiments, while a one-way analysis of variance was used for multiple comparisons.
Morphometric analysis did not inform about the extent of ossification within each larva, and was sometimes hindered by the complete absence of one or several elements. Therefore, a systematic analysis was generated by giving each structure a score based on its progress of development: absent, early ossification, advanced ossification and over ossification, or numerical values of, respectively, 0, 1, 2, or 3. A contingency table considered ordinal values distributed among the four classes (from absent to over ossification) or only three classes when one class was not present in the sample, and association between classes and treatment was assessed by X2-test and by an ordinal logistic regression and the odds ratio. Quantitative values were compared between two groups using a Student t-test. Statistical analyses were performed using the Statistica Software (version 10, Dell Software, Paris, France). Results were considered statistically significant at the 5% critical level (P<0.05).
RNA extraction, reverse transcription and real-time PCR
Samples of 60 larvae at 6 dpf, after 24 h treatment, were pooled for RNA extraction, complementary DNA synthesis, and quantitative PCR were performed as previously described.57 Gene-specific oligonucleotide primers were designed using Primer3 software58 to span exon–exon junctions to avoid detection of genomic DNA contamination (see Supplementary Table S1 for primer sequences) and synthesized by Eurogentec (Seraing, Belgium) or Integrated DNA Technology (Leuven, Belgium).
Microarray expression experiments
Microarray expression analysis was performed as previously described,9 the slides were scanned using a GenePix 4000B (Axon instrument, Foster City, CA, USA). Raw data and complete lists of analyzed data are publicly available at Arrayexpress (https://www.ebi.ac.uk/arrayexpress/) under accession number E-MTAB-4476.
Ingenuity pathway analysis
Pathway and biological function analysis of significantly differently expressed genes using IPA (QIAGEN Redwood City; http://www.ingenuity.com) was previously described.9 In some cases, activation z-scores are used in the statistical analysis. This score identifies upstream regulators or pathways that can explain the observed gene expression changes in the dataset, by taking into account the direction (induced or reduced expression) and extent of change, and based on regulations known from the entire IPA database. Z-scores >2 predict activation of the upstream regulator, z-score <−2 predict inhibition.
References
Nagaraja, M. P. & Risin, D. The current state of bone loss research: data from spaceflight and microgravity simulators. J. Cell Biochem. 114, 1001–1008 (2013).
Morgan, J. L. et al. Sex-specific responses of bone metabolism and renal stone risk during bed rest. Physiol. Rep. 2; e12119; doi:10.14814/phy2.12119 (2014).
Horn, E., van Loon JJWA, Aceto, J., Muller, M. Life Sciences: Animal Physiology Laboratory Science with Space Data, (eds). Beysens D., Carotenuto L., van Loon J. J. W. A. & Zell M. ISBN 978-3-642-21143-0, Springer, Verlag Berlin Heidelberg 2011. 2011: 123–9.
Muller, M. et al. Small fish species as powerful model systems to study vertebrate physiology in space. J. Gravit. Physiol. 15, 253–254 (2008).
Rahmann, H. & Anken, R. H. Gravity related research with fishes—Perspectives in regard to the upcoming International Space Station, ISS. Space Life Sci. 30, 697–710 (2002).
Slenzka, K., Appel, R. & Rahmann, H. Development and altered gravity dependent changes in glucose-6-phosphate dehydrogenase activity in the brain of the cichlid fish Oreochromis mossambicus. Neurochem. Int. 26, 579–585 (1995).
Brungs, S., Hauslage, J., Hilbig, R., Hemmersbach, R. & Anken, R. Effects of simulated weightlessness on fish otolith growth: clinostat versus rotating-wall vessel. Adv Space Res 48, 792–798 (2011).
Aceto, J. et al. Microgravity simulation comparison at genome level in Danio rerio and role of Sox4 transcription factors in cranial skeleton development. J. Gravit. Physiol. 16, 103–104 (2009).
Aceto, J. et al. Zebrafish bone and general physiology are differently affected by hormones or changes in gravity. PLoS ONE 10, e0126928 (2015).
Henning, P. C., Park, B. S. & Kim, J. S. Physiological decrements during sustained military operational stress. Military Med. 176, 991–997 (2011).
Feng, X. & McDonald, J. M. Disorders of bone remodeling. Annu. Rev. Pathol. 6, 121–145 (2011).
Santamaria, N. et al. Fin spine bone resorption in atlantic bluefin tuna, Thunnus thynnus, and comparison between wild and captive-reared specimens. PLoS ONE 10, e0121924 (2015).
Weinstein, R. S. Glucocorticoid-induced osteoporosis and osteonecrosis. Endocrinol. Metab. Clin. North Am. 41, 595–611 (2012).
Westerfield, M . The Zebrafish Book: a Guide for the Laboratory Use of Zebrafish (Danio rerio) 5th edn, Eugene, University of Oregon Press, 2007.
Alsop, D. & Vijayan, M. M. Development of the corticosteroid stress axis and receptor expression in zebrafish. Am. J. Physiol. Regul. Integr. Comp. Physiol. 294, R711–R719 (2008).
Alderman, S. L. & Bernier, N. J. Ontogeny of the corticotropin-releasing factor system in zebrafish. Gen. Comp. Endocrinol. 164, 61–69 (2009).
Kageyama, R. & Ohtsuka, T. The Notch-Hes pathway in mammalian neural development. Cell Res. 9, 179–188 (1999).
Windhausen, T., Squifflet, S., Renn, J. & Muller, M. BMP signaling regulates bone morphogenesis in zebrafish through promoting osteoblast function as assessed by their nitric oxide production. Molecules 20, 7586–7601 (2015).
Larbuisson, A., Dalcq, J., Martial, J. A. & Muller, M. Fgf receptors Fgfr1a and Fgfr2 control the function of pharyngeal endoderm in late cranial cartilage development. Differentiation 86, 192–206 (2013).
Dalcq, J. et al. Runx3, Egr1 and Sox9b form a regulatory cascade required to modulate BMP-signaling during cranial cartilage development in zebrafish. PLoS ONE 7, e50140 (2012).
Van Loon, J. J. et al. Decreased mineralization and increased calcium release in isolated fetal mouse long bones under near weightlessness. J. Bone Miner. Res. 10, 550–557 (1995).
Caillot-Augusseau, A. et al. Bone formation and resorption biological markers in cosmonauts during and after a 180-day space flight (Euromir 95). Clin. Chem. 44, 578–585 (1998).
Carmeliet, G., Vico, L. & Bouillon, R. Space flight: a challenge for normal bone homeostasis. Crit. Rev. Eukaryot. Gene Expr. 11, 131–144 (2001).
Sabatakos, G. et al. Doubly truncated FosB isoform (Delta2DeltaFosB) induces osteosclerosis in transgenic mice and modulates expression and phosphorylation of Smads in osteoblasts independent of intrinsic AP-1 activity. J. Bone Miner. Res. 23, 584–595 (2008).
Morey, E. R. & Baylink, D. J. Inhibition of bone formation during space flight. Science 201, 1138–1141 (1978).
Wronski, T. J., Morey-Holton, E. R., Doty, S. B., Maese, A. C. & Walsh, C. C. Histomorphometric analysis of rat skeleton following spaceflight. Am. J. Physiol. 252, R252–R255 (1987).
Vico, L. et al. Trabecular bone remodeling after seven days of weightlessness exposure (BIOCOSMOS 1667). Am. J. Physiol. 255, R243–R247 (1988).
Turner, R. T., Evans, G. L. & Wakley, G. K. Spaceflight results in depressed cancellous bone formation in rat humeri. Aviat. Space Environ. Med. 66, 770–774 (1995).
Tavella, S. et al. Bone turnover in wild type and pleiotrophin-transgenic mice housed for three months in the International Space Station (ISS). PLoS ONE 7, e33179 (2012).
Wronski, T. J. & Morey, E. R. Recovery of the rat skeleton from the adverse effects of simulated weightlessness. Metab. Bone Dis. Relat. Res. 4, 347–352 (1983).
Caillot-Augusseau, A. et al. Space flight is associated with rapid decreases of undercarboxylated osteocalcin and increases of markers of bone resorption without changes in their circadian variation: observations in two cosmonauts. Clin. Chem. 46, 1136–1143 (2000).
Papa, S. et al. Gadd45 beta mediates the NF-kappa B suppression of JNK signalling by targeting MKK7/JNKK2. Nat. Cell Biol. 6, 146–153 (2004).
Lu, B., Ferrandino, A. F. & Flavell, R. A. Gadd45beta is important for perpetuating cognate and inflammatory signals in T cells. Nat. Immunol. 5, 38–44 (2004).
Hughes-Fulford, M., Rodenacker, K. & Jutting, U. Reduction of anabolic signals and alteration of osteoblast nuclear morphology in microgravity. J. Cell Biochem. 99, 435–449 (2006).
Sato, A. et al. Effects of microgravity on c-fos gene expression in osteoblast-like MC3T3-E1 cells. Adv. Space Res. 24, 807–813 (1999).
de Groot, R. P. et al. Microgravity decreases c-fos induction and serum response element activity. J. Cell Sci. 97, 33–38 (1990).
de Groot, R. P. et al. Nuclear responses to protein kinase C signal transduction are sensitive to gravity changes. Exp. Cell Res. 197, 87–90 (1991).
Zanotti, S. & Canalis, E. Notch regulation of bone development and remodeling and related skeletal disorders. Calcif. Tissue Int. 90, 69–75 (2012).
Hatakeyama, J. et al. Hes genes regulate size, shape and histogenesis of the nervous system by control of the timing of neural stem cell differentiation. Development 131, 5539–5550 (2004).
Karlsson, C. et al. Notch and HES5 are regulated during human cartilage differentiation. Cell Tissue Res. 327, 539–551 (2007).
Crosnier, C., Stamataki, D. & Lewis, J. Organizing cell renewal in the intestine: stem cells, signals and combinatorial control. Nat. Rev. Genet. 7, 349–359 (2006).
Wohrle, F. U., Daly, R. J. & Brummer, T. Function, regulation and pathological roles of the Gab/DOS docking proteins. Cell Commun. Signal. 7, 22 (2009).
Simister, P. C. & Feller, S. M. Order and disorder in large multi-site docking proteins of the Gab family--implications for signalling complex formation and inhibitor design strategies. Mol. Biosyst. 8, 33–46 (2012).
Yang-Yen, H. F. Mcl-1: a highly regulated cell death and survival controller. J. Biomed. Sci. 13, 201–204 (2006).
Wu, L. et al. The E2F1-3 transcription factors are essential for cellular proliferation. Nature 414, 457–462 (2001).
Denis, G. V., Vaziri, C., Guo, N. & Faller, D. V. RING3 kinase transactivates promoters of cell cycle regulatory genes through E2F. Cell Growth Differ. 11, 417–424 (2000).
Simpson, E. R. et al. Aromatase—a brief overview. Annu. Rev. Physiol. 64, 93–127 (2002).
Neutelings, T. et al. Skin physiology in microgravity: a 3-month stay aboard ISS induces dermal atrophy and affects cutaneous muscle and hair follicles cycling in mice. NPJ Microgravity 1; doi:10.1038/npjmgrav.2015.1032 (2015).
Pardo, S. J. et al. Simulated microgravity using the Random Positioning Machine inhibits differentiation and alters gene expression profiles of 2T3 preosteoblasts. Am. J. Physiol. Cell Physiol. 288, C1211–C1221 (2005).
Versari, S., Klein-Nulend, J., van Loon, J. & Bradamante, S. Influence of oxygen in the cultivation of human mesenchymal stem cells in simulated microgravity: an explorative study. Microgravity Sci. Technol. 25, 59–66 (2013).
Versari, S., Longinotti, G., Barenghi, L., Maier, J. A. & Bradamante, S. The challenging environment on board the International Space Station affects endothelial cell function by triggering oxidative stress through thioredoxin interacting protein overexpression: the ESA-SPHINX experiment. FASEB J. 27, 4466–4475 (2013).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
van Loon, J. J. W. A. et al. Microgravity research starts on the ground! Apparatuses for long-term ground based hypo-and hypergravity studies in Proceedings of the 2nd European Symposium on the Utilisation of the International Space Station, ESA SP-433, European Space Agency, Noordwijk, the Netherlands. 415–419 (1999).
Walker, M. B. & Kimmel, C. B. A two-color acid-free cartilage and bone stain for zebrafish larvae. Biotechnol. Histochem. 82, 23–28 (2007).
Pruvot, B., Curé, Y., Djiotsa, J., Voncken, A. & Muller, M. Developmental defects in zebrafish for classification of EGF pathway inhibitors. Toxicol. Appl. Pharmacol. 274, 339–349 (2014).
Marée, R. et al. A rich internet application for remote visualization and collaborative annotation of digital slide images in histology and cytology. Diagn. Pathol. 8, p S26–S29 (2013).
Quiroz, Y. et al. The HMG-Box transcription factor Sox4b is required for pituitary expression of gata2a and specification of thyrotrope and gonadotrope cells in zebrafish. Mol. Endocrinol. 26, 1014–1027 (2012).
Andreas, U. et al. Leunissen: Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Research 35, W71–W74; doi:10.1093/nar/gkm306 (2007).
Acknowledgements
We wish to thank the GIGA-R zebrafish facility for providing zebrafish adults and fertilized eggs, the GIGA-R GenoTranscriptomics platform for DNA sequencing and RNA quality control, and the ESA-ESTEC, Prodex, TEC-MMG LIS Lab, and especially Mr Alan Dowson for his support during the study. This work was supported by the ‘Fonds de la Recherche Fondamentale Collective’: 2.4555.99/ 2.4542.00/2.4561.10; the SSTC PAI: P5/35; the University of Liège GAME project; the European Space Agency projects AO-99-LSS-003 and AO-99-LSS-006; the Belgian Space Agency Prodex projects FISH-GSIM and FISH-SIM. M.M. is a ‘Chercheur Qualifié du F.N.R.S.’ J.J.W.A.vL. grant MG-057 from the Netherlands Organisation for Scientific (NWO) Research Earth, and Life Sciences via the Netherlands Space Office (NSO) and an ESA contract 4000107455/12/NL/PA.
Author information
Authors and Affiliations
Contributions
J.A. was involved in performing all experiments; R.N.-L. helped to perform the microarray experiments and analysis; M.M., P.A., J.J.W.A.vL., S.B. and J.M. were involved in planning, interpretation, writing and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Aceto, J., Nourizadeh-Lillabadi, R., Bradamante, S. et al. Effects of microgravity simulation on zebrafish transcriptomes and bone physiology—exposure starting at 5 days post fertilization. npj Microgravity 2, 16010 (2016). https://doi.org/10.1038/npjmgrav.2016.10
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/npjmgrav.2016.10
This article is cited by
-
Comparison of Effects of 2D and 3D Simulated Microgravity Rotation on Ossification in Larval Danio Rerio (Zebrafish)
Microgravity Science and Technology (2023)
-
Acute transcriptional up-regulation specific to osteoblasts/osteoclasts in medaka fish immediately after exposure to microgravity
Scientific Reports (2016)