Abstract
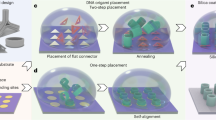
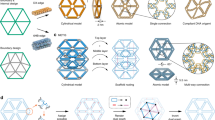
The aim of nanotechnology is to put specific atomic and molecular species where we want them, when we want them there. Achieving such dynamic and functional control could lead to programmable chemical synthesis and nanoscale systems that are responsive to their environments. Structural DNA nanotechnology offers a powerful route to this goal by combining stable branched DNA motifs1 with cohesive ends to produce programmed nanomechanical devices2 and fixed3,4,5 or modified6,7 patterned lattices. Here, we demonstrate a dynamic form of patterning8 in which a pattern component is captured between two independently programmed DNA devices. A simple and robust error-correction protocol has been developed that yields programmed targets in all cases. This capture system can lead to dynamic control either on patterns or on programmed elements; this capability enables computation or a change of structural state as a function of information in the surroundings of the system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Seeman, N. C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Seeman, N. C. & Lukeman, P. S. Nucleic acid nanostructures. Rep. Prog. Phys. 68, 237–270 (2005).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Rothemund, P. W. K. Scaffolded DNA origami for nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of Sierpinski triangles. PLoS Biol. 2, 2041–2053 (2004).
Liu, F., Sha, R. & Seeman, N. C. Modifying the surface features of two-dimensional DNA crystals. J. Am. Chem. Soc. 121, 917–922 (1999).
Garibotti, A. V., Knudsen, S. M., Ellington, A. D. & Seeman, N. C. Functional DNAzymes organized into 2D arrays. Nano Lett. 6, 1505–1507 (2006).
Carbone, A. & Seeman, N. C. Circuits and programmable self-assembling DNA structures. Proc. Natl Acad. Sci. USA 99, 12577–12582 (2002).
Yan, H., Zhang, X., Shen, Z. & Seeman, N. C. A robust DNA mechanical device controlled by hybridization topology. Nature 415, 62–65 (2002).
Ding, B. & Seeman, N. C. Operation of a DNA robot arm inserted into a 2D DNA crystalline substrate. Science 314, 1583–1585 (2006).
Rinker, S., Ke, Y., Liu, Y., Chhabra, R. & Yan, H. Self-assembled DNA nanostructures for distance-dependent multivalent ligand–protein binding. Nature Nanotech. 3, 418–422 (2008).
Mao, C., LaBean, T., Reif, J. H. & Seeman, N. C. Logical computation using algorithmic self-assembly of DNA triple crossover molecules. Nature 407, 493–496 (2000).
Zheng, J. et al. 2D nanoparticle arrays show the organizational power of robust DNA motifs. Nano Lett. 6, 1502–1504 (2006).
Ke, Y., Lindsay, S., Chang, Y., Liu, Y. & Yan, H. Self-assembled water-soluble nucleic acid probe molecules for label-free RNA hybridization. Science 319, 180–183 (2008).
Acknowledgements
We are grateful to A. Carbone, H. Yan, N. Jonoska and C. Mao for comments on this manuscript. This research has been supported by grants to N.C.S. from the National Institute of General Medical Sciences, the National Science Foundation, the Army Research Office, the NYNBIT program of the Department of Energy and the W.M. Keck Foundation and to S.J.X. from the National Basic Research Program of China (no. 2007CB925101) and NSFC (no. 20721002). J.C. thanks the Chinese Scholarship Council for a research fellowship.
Author information
Authors and Affiliations
Supplementary information
Supplementary Information
Supplementary Information (PDF 3839 kb)
Rights and permissions
About this article
Cite this article
Gu, H., Chao, J., Xiao, SJ. et al. Dynamic patterning programmed by DNA tiles captured on a DNA origami substrate. Nature Nanotech 4, 245–248 (2009). https://doi.org/10.1038/nnano.2009.5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2009.5
This article is cited by
-
DNA nanotechnology
Nature Reviews Materials (2017)
-
A device that operates within a self-assembled 3D DNA crystal
Nature Chemistry (2017)
-
DNA hydrogel-based supercapacitors operating in physiological fluids
Scientific Reports (2013)