Abstract
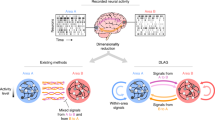
Understanding how the brain operates requires understanding how large sets of neurons function together. Modern recording technology makes it possible to simultaneously record the activity of hundreds of neurons, and technological developments will soon allow recording of thousands or tens of thousands. As with all experimental techniques, these methods are subject to confounds that complicate the interpretation of such recordings, and could lead to erroneous scientific conclusions. Here we discuss methods for assessing and improving the quality of data from these techniques and outline likely future directions in this field.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Buzsáki, G. Large-scale recording of neuronal ensembles. Nat. Neurosci. 7, 446–451 (2004).
Gold, C., Henze, D.A., Koch, C. & Buzsáki, G. On the origin of the extracellular action potential waveform: a modeling study. J. Neurophysiol. 95, 3113–3128 (2006).
Quiroga, R.Q. Spike sorting. Curr. Biol. 22, R45–R46 (2012).
Berényi, A. et al. Large-scale, high-density (up to 512 channels) recording of local circuits in behaving animals. J. Neurophysiol. 111, 1132–1149 (2014).
Lewicki, M.S. A review of methods for spike sorting: the detection and classification of neural action potentials. Network 9, R53–R78 (1998).
Rey, H.G., Pedreira, C. & Quian Quiroga, R. Past, present and future of spike sorting techniques. Brain Res. Bull. 119, Pt B, 106–117 (2015).
Einevoll, G.T., Franke, F., Hagen, E., Pouzat, C. & Harris, K.D. Towards reliable spike-train recordings from thousands of neurons with multielectrodes. Curr. Opin. Neurobiol. 22, 11–17 (2012).
Henze, D.A. et al. Intracellular features predicted by extracellular recordings in the hippocampus in vivo. J. Neurophysiol. 84, 390–400 (2000).
Rey, H.G. et al. Single-cell recordings in the human medial temporal lobe. J. Anat. 227, 394–408 (2015).
Quiroga, R.Q. Concept cells: the building blocks of declarative memory functions. Nat. Rev. Neurosci. 13, 587–597 (2012).
Quiroga, R.Q., Reddy, L., Koch, C. & Fried, I. Decoding visual inputs from multiple neurons in the human temporal lobe. J. Neurophysiol. 98, 1997–2007 (2007).
Hill, D.N., Mehta, S.B. & Kleinfeld, D. Quality metrics to accompany spike sorting of extracellular signals. J. Neurosci. 31, 8699–8705 (2011).
Ludwig, K.A., Uram, J.D., Yang, J., Martin, D.C. & Kipke, D.R. Chronic neural recordings using silicon microelectrode arrays electrochemically deposited with a poly(3,4-ethylenedioxythiophene) (PEDOT) film. J. Neural Eng. 3, 59–70 (2006).
Gerstein, G.L. & Clarke, W.A. Simultaneous studies of firing patterns in several neurons. Science 143, 1325–1327 (1964).
Robinson, D.A. The electrical properties of metal microelectrodes. Proc. IEEE 56, 1065–1071 (1968).
Camuñas-Mesa, L.A. & Quiroga, R.Q. A detailed and fast model of extracellular recordings. Neural Comput. 25, 1191–1212 (2013).
Pedreira, C., Martinez, J., Ison, M.J. & Quian Quiroga, R. How many neurons can we see with current spike sorting algorithms? J. Neurosci. Methods 211, 58–65 (2012).
Quian Quiroga, R. & Panzeri, S. Principles of Neural Coding (Taylor & Francis/CRC Press, 2013).
Buzsáki, G. Neural syntax: cell assemblies, synapsembles, and readers. Neuron 68, 362–385 (2010).
Brown, E.N., Kass, R.E. & Mitra, P.P. Multiple neural spike train data analysis: state-of-the-art and future challenges. Nat. Neurosci. 7, 456–461 (2004).
Quian Quiroga, R. & Panzeri, S. Extracting information from neuronal populations: information theory and decoding approaches. Nat. Rev. Neurosci. 10, 173–185 (2009).
Harris, K.D. Neural signatures of cell assembly organization. Nat. Rev. Neurosci. 6, 399–407 (2005).
Spira, M.E. & Hai, A. Multi-electrode array technologies for neuroscience and cardiology. Nat. Nanotechnol. 8, 83–94 (2013).
Lambacher, A. et al. Identifying firing mammalian neurons in networks with high-resolution multitransistor array (MTA). Appl. Phys. A. 102, 1–11 (2011).
Frey, U., Egert, U., Heer, F., Hafizovic, S. & Hierlemann, A. Microelectronic system for high-resolution mapping of extracellular electric fields applied to brain slices. Biosens. Bioelectron. 24, 2191–2198 (2009).
Litke, A. et al. What does the eye tell the brain? Development of a system fo the large-scale recording of retinal output activity. IEEE Trans. Nucl. Sci. 51, 1434–1440 (2004).
Lopez, C.M. et al. A 966-electrode neural probe with 384 configurable channels in 0.13 μm SOI CMOS. IEEE Int. Solid State Circuits Conf. 392–393 (2016).
Nicolelis, M.A., Ghazanfar, A.A., Faggin, B.M., Votaw, S. & Oliveira, L.M. Reconstructing the engram: simultaneous, multisite, many single neuron recordings. Neuron 18, 529–537 (1997).
Hochberg, L.R. et al. Neuronal ensemble control of prosthetic devices by a human with tetraplegia. Nature 442, 164–171 (2006).
McNaughton, B.L., O'Keefe, J. & Barnes, C.A. The stereotrode: a new technique for simultaneous isolation of several single units in the central nervous system from multiple unit records. J. Neurosci. Methods 8, 391–397 (1983).
Gray, C.M., Maldonado, P.E., Wilson, M. & McNaughton, B. Tetrodes markedly improve the reliability and yield of multiple single-unit isolation from multi-unit recordings in cat striate cortex. J. Neurosci. Methods 63, 43–54 (1995).
Blanche, T.J., Spacek, M.A., Hetke, J.F. & Swindale, N.V. Polytrodes: high-density silicon electrode arrays for large-scale multiunit recording. J. Neurophysiol. 93, 2987–3000 (2005).
Drake, K.L., Wise, K.D., Farraye, J., Anderson, D.J. & BeMent, S.L. Performance of planar multisite microprobes in recording extracellular single-unit intracortical activity. IEEE Trans. Biomed. Eng. 35, 719–732 (1988).
Bai, Q. & Wise, K.D. Single-unit neural recording with active microelectrode arrays. IEEE Trans. Biomed. Eng. 48, 911–920 (2001).
Csicsvari, J. et al. Massively parallel recording of unit and local field potentials with silicon-based electrodes. J. Neurophysiol. 90, 1314–1323 (2003).
Harris, K.D., Henze, D.A., Csicsvari, J., Hirase, H. & Buzsáki, G. Accuracy of tetrode spike separation as determined by simultaneous intracellular and extracellular measurements. J. Neurophysiol. 84, 401–414 (2000).
Rossant, C. et al. Spike sorting for large, dense electrode arrays. Nat. Neurosci. 19, 634–641 (2016).
Briggman, K.L., Helmstaedter, M. & Denk, W. Wiring specificity in the direction-selectivity circuit of the retina. Nature 471, 183–188 (2011).
Dhawale, A.K. et al. Automated long-term recording and analysis of neural activity in behaving animals. bioRxiv http://dx.doi.org/10.1101/033266 (2015).
Anastassiou, C.A., Perin, R., Buzsáki, G., Markram, H. & Koch, C. Cell-type- and activity-dependent extracellular correlates of intracellular spiking. J. Neurophysiol. 114, 608–623 (2015).
Wehr, M., Pezaris, J. & Sahani, M. Simultaneous paired intracellular and tetrode recordings for spike sorting algorithms. Neurocomputing 26, 1061–1068 (1999).
Delescluse, M. & Pouzat, C. Efficient spike-sorting of multi-state neurons using inter-spike intervals information. J. Neurosci. Methods 150, 16–29 (2006).
Neto, J.P. et al. Validating silicon polytrodes with paired juxtacellular recordings: method and dataset. bioRxiv http://dx.doi.org/10.1101/037937 (2016).
Lindén, H. et al. LFPy: a tool for biophysical simulation of extracellular potentials generated by detailed model neurons. Front. Neuroinform. 7, 41 (2014).
Menne, K., Folkers, A., Malina, T., Maex, R. & Hofmann, U. Test of spike-sorting algorithms on the basis of simulated network data. Neurocomputing 44–46, 1119–1126 (2002).
Petersen, K.E. & Einevoll, G.T. Amplitude variability and extracellular low-pass filtering of neuronal spikes. Biophys. J. 94, 784–802 (2008).
Martinez, J., Pedreira, C., Ison, M.J. & Quian Quiroga, R. Realistic simulation of extracellular recordings. J. Neurosci. Methods 184, 285–293 (2009).
Navajas, J. et al. Minimum requirements for accurate and efficient real-time on-chip spike sorting. J. Neurosci. Methods 230, 51–64 (2014).
Barnett, A.H., Magland, J.F. & Greengard, L.F. Validation of neural spike sorting algorithms without ground-truth information. J. Neurosci. Methods 264, 65–77 (2016).
Franke, F., Natora, M., Boucsein, C., Munk, M.H. & Obermayer, K. An online spike detection and spike classification algorithm capable of instantaneous resolution of overlapping spikes. J. Comput. Neurosci. 29, 127–148 (2010).
Pillow, J.W., Shlens, J., Chichilnisky, E.J. & Simoncelli, E.P. A model-based spike sorting algorithm for removing correlation artifacts in multi-neuron recordings. PLoS One 8, e62123 (2013).
Ekanadham, C., Tranchina, D. & Simoncelli, E.P. A unified framework and method for automatic neural spike identification. J. Neurosci. Methods 222, 47–55 (2014).
Pachitariu, M., Steinmetz, N.A., Kadir, S.N., Carandini, M. & Harris, K.D. Kilosort: realtime spike-sorting for extracellular electrophysiology with hundreds of channels. bioRxiv http://dx.doi.org/10.1101/061481 (2016).
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A.D. Quantitative measures of cluster quality for use in extracellular recordings. Neuroscience 131, 1–11 (2005).
Pouzat, C., Mazor, O. & Laurent, G. Using noise signature to optimize spike-sorting and to assess neuronal classification quality. J. Neurosci. Methods 122, 43–57 (2002).
Harris, K.D., Hirase, H., Leinekugel, X., Henze, D.A. & Buzsáki, G. Temporal interaction between single spikes and complex spike bursts in hippocampal pyramidal cells. Neuron 32, 141–149 (2001).
Fee, M.S., Mitra, P.P. & Kleinfeld, D. Variability of extracellular spike waveforms of cortical neurons. J. Neurophysiol. 76, 3823–3833 (1996).
Quirk, M.C. & Wilson, M.A. Interaction between spike waveform classification and temporal sequence detection. J. Neurosci. Methods 94, 41–52 (1999).
Cohen, M.R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Pazienti, A. & Grün, S. Robustness of the significance of spike synchrony with respect to sorting errors. J. Comput. Neurosci. 21, 329–342 (2006).
Ventura, V. & Gerkin, R.C. Accurately estimating neuronal correlation requires a new spike-sorting paradigm. Proc. Natl. Acad. Sci. USA 109, 7230–7235 (2012).
Ganguly, K., Dimitrov, D.F., Wallis, J.D. & Carmena, J.M. Reversible large-scale modification of cortical networks during neuroprosthetic control. Nat. Neurosci. 14, 662–667 (2011).
Fraser, G.W., Chase, S.M., Whitford, A. & Schwartz, A.B. Control of a brain-computer interface without spike sorting. J. Neural Eng. 6, 055004 (2009).
Redish, A.D. et al. Independence of firing correlates of anatomically proximate hippocampal pyramidal cells. J. Neurosci. 21, RC134 (2001).
Shoham, S., O'Connor, D.H. & Segev, R. How silent is the brain: is there a “dark matter” problem in neuroscience? J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 192, 777–784 (2006).
Moffitt, M.A. & McIntyre, C.C. Model-based analysis of cortical recording with silicon microelectrodes. Clin. Neurophysiol. 116, 2240–2250 (2005).
Claverol-Tinture, E. & Nadasdy, Z. Intersection of microwire electrodes with proximal CA1 stratum-pyramidale neurons at insertion for multiunit recordings predicted by a 3-D computer model. IEEE Trans. Biomed. Eng. 51, 2211–2216 (2004).
Buzsáki, G. & Mizuseki, K. The log-dynamic brain: how skewed distributions affect network operations. Nat. Rev. Neurosci. 15, 264–278 (2014).
de Kock, C.P., Bruno, R.M., Spors, H. & Sakmann, B. Layer- and cell-type-specific suprathreshold stimulus representation in rat primary somatosensory cortex. J. Physiol. (Lond.) 581, 139–154 (2007).
Sakata, S. & Harris, K.D. Laminar structure of spontaneous and sensory-evoked population activity in auditory cortex. Neuron 64, 404–418 (2009).
Niell, C.M. & Stryker, M.P. Highly selective receptive fields in mouse visual cortex. J. Neurosci. 28, 7520–7536 (2008).
O'Connor, D.H., Peron, S.P., Huber, D. & Svoboda, K. Neural activity in barrel cortex underlying vibrissa-based object localization in mice. Neuron 67, 1048–1061 (2010).
Hubel, D.H. Tungsten microelectrode for recording from single units. Science 125, 549–550 (1957).
Kentros, C. et al. Abolition of long-term stability of new hippocampal place cell maps by NMDA receptor blockade. Science 280, 2121–2126 (1998).
Okun, M., Carandini, M. & Harris, K.D. Long term recordings with immobile silicon probes in the mouse cortex. PLoS One http://dx.doi.org/10.1371/journal.pone.0151180 (2015).
Spruston, N., Schiller, Y., Stuart, G. & Sakmann, B. Activity-dependent action potential invasion and calcium influx into hippocampal CA1 dendrites. Science 268, 297–300 (1995).
Buzsáki, G. & Kandel, A. Somadendritic backpropagation of action potentials in cortical pyramidal cells of the awake rat. J. Neurophysiol. 79, 1587–1591 (1998).
Tolias, A.S. et al. Recording chronically from the same neurons in awake, behaving primates. J. Neurophysiol. 98, 3780–3790 (2007).
Swindale, N.V. & Spacek, M.A. Spike sorting for polytrodes: a divide and conquer approach. Front. Syst. Neurosci. 8, 6 (2014).
Bar-Hillel, A., Spiro, A. & Stark, E. Spike sorting: Bayesian clustering of non-stationary data. J. Neurosci. Methods 157, 303–316 (2006).
Jackson, A. & Fetz, E.E. Compact movable microwire array for long-term chronic unit recording in cerebral cortex of primates. J. Neurophysiol. 98, 3109–3118 (2007).
Fee, M.S., Mitra, P.P. & Kleinfeld, D. Automatic sorting of multiple unit neuronal signals in the presence of anisotropic and non-Gaussian variability. J. Neurosci. Methods 69, 175–188 (1996).
Williams, J.C., Rennaker, R.L. & Kipke, D.R. Stability of chronic multichannel neural recordings: implications for a long-term neural interface. Neurocomputing 26–27, 1069–1076 (1999).
Greenberg, P.A. & Wilson, F.A. Functional stability of dorsolateral prefrontal neurons. J. Neurophysiol. 92, 1042–1055 (2004).
Ohki, K., Chung, S., Ch'ng, Y.H., Kara, P. & Reid, R.C. Functional imaging with cellular resolution reveals precise micro-architecture in visual cortex. Nature 433, 597–603 (2005).
Stosiek, C., Garaschuk, O., Holthoff, K. & Konnerth, A. In vivo two-photon calcium imaging of neuronal networks. Proc. Natl. Acad. Sci. USA 100, 7319–7324 (2003).
Theis, L. et al. Benchmarking spike rate inference in population calcium imaging. Neuron 90, 471–482 (2016).
Rose, T., Goltstein, P.M., Portugues, R. & Griesbeck, O. Putting a finishing touch on GECIs. Front. Mol. Neurosci. 7, 88 (2014).
Ghosh, K.K. et al. Miniaturized integration of a fluorescence microscope. Nat. Methods 8, 871–878 (2011).
Keller, P.J., Ahrens, M.B. & Freeman, J. Light-sheet imaging for systems neuroscience. Nat. Methods 12, 27–29 (2015).
Chen, T.W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Ohki, K. et al. Highly ordered arrangement of single neurons in orientation pinwheels. Nature 442, 925–928 (2006).
Dombeck, D.A., Harvey, C.D., Tian, L., Looger, L.L. & Tank, D.W. Functional imaging of hippocampal place cells at cellular resolution during virtual navigation. Nat. Neurosci. 13, 1433–1440 (2010).
Yasuda, R. et al. Imaging calcium concentration dynamics in small neuronal compartments. Sci. STKE 2004, pl5 (2004).
Podgorski, K. & Ranganathan, G.N. Brain heating induced by near infrared lasers during multi-photon microscopy. J.Neurophysiol. http://dx.doi.org/10.1152/jn.00275.2016 (2016).
Valmianski, I. et al. Automatic identification of fluorescently labeled brain cells for rapid functional imaging. J. Neurophysiol. 104, 1803–1811 (2010).
Sadovsky, A.J. et al. Heuristically optimal path scanning for high-speed multiphoton circuit imaging. J. Neurophysiol. 106, 1591–1598 (2011).
Cotton, R.J., Froudarakis, E., Storer, P., Saggau, P. & Tolias, A.S. Three-dimensional mapping of microcircuit correlation structure. Front. Neural Circuits 7, 151 (2013).
Hamel, E.J., Grewe, B.F., Parker, J.G. & Schnitzer, M.J. Cellular level brain imaging in behaving mammals: an engineering approach. Neuron 86, 140–159 (2015).
Wilt, B.A., Fitzgerald, J.E. & Schnitzer, M.J. Photon shot noise limits on optical detection of neuronal spikes and estimation of spike timing. Biophys. J. 104, 51–62 (2013).
Young, M.D., Field, J.J., Sheetz, K.E., Bartels, R.A. & Squier, J. A pragmatic guide to multiphoton microscope design. Adv. Opt. Photonics 7, 276–378 (2015).
Kerr, J.N., Greenberg, D. & Helmchen, F. Imaging input and output of neocortical networks in vivo. Proc. Natl. Acad. Sci. USA 102, 14063–14068 (2005).
Lecoq, J. et al. Visualizing mammalian brain area interactions by dual-axis two-photon calcium imaging. Nat. Neurosci. 17, 1825–1829 (2014).
Stirman, J.N., Smith, I.T., Kudenov, M.W. & Smith, S.L. Wide field-of-view, twin-region two-photon imaging across extended cortical networks. Nat. Biotechnol. http://dx.doi.org/10.1038/nbt.3594 (2016).
Helmchen, F. & Denk, W. Deep tissue two-photon microscopy. Nat. Methods 2, 932–940 (2005).
Yang, W. et al. Simultaneous multi-plane imaging of neural circuits. Neuron 89, 269–284 (2016).
Dunn, A.K., Wallace, V.P., Coleno, M., Berns, M.W. & Tromberg, B.J. Influence of optical properties on two-photon fluorescence imaging in turbid samples. Appl. Opt. 39, 1194–1201 (2000).
Tung, C.K. et al. Effects of objective numerical apertures on achievable imaging depths in multiphoton microscopy. Microsc. Res. Tech. 65, 308–314 (2004).
Zheng, G., Ou, X., Horstmeyer, R. & Yang, C. Characterization of spatially varying aberrations for wide field-of-view microscopy. Opt. Express 21, 15131–15143 (2013).
von Diezmann, A., Lee, M.Y., Lew, M.D. & Moerner, W.E. Correcting field-dependent aberrations with nanoscale accuracy in three-dimensional single-molecule localization microscopy. Optica 2, 985–993 (2015).
Wang, C. & Ji, N. Characterization and improvement of three-dimensional imaging performance of GRIN-lens-based two-photon fluorescence endomicroscopes with adaptive optics. Opt. Express 21, 27142–27154 (2013).
Barretto, R.P., Messerschmidt, B. & Schnitzer, M.J. In vivo fluorescence imaging with high-resolution microlenses. Nat. Methods 6, 511–512 (2009).
Ji, N., Sato, T.R. & Betzig, E. Characterization and adaptive optical correction of aberrations during in vivo imaging in the mouse cortex. Proc. Natl. Acad. Sci. USA 109, 22–27 (2012).
Dana, H. et al. Thy1-GCaMP6 transgenic mice for neuronal population imaging in vivo. PLoS One 9, e108697 (2014).
Chen, Q. et al. Imaging neural activity using Thy1-GCaMP transgenic mice. Neuron 76, 297–308 (2012).
Glickfeld, L.L., Andermann, M.L., Bonin, V. & Reid, R.C. Cortico-cortical projections in mouse visual cortex are functionally target specific. Nat. Neurosci. 16, 219–226 (2013).
Peron, S.P., Freeman, J., Iyer, V., Guo, C. & Svoboda, K. A cellular resolution map of barrel cortex activity during tactile behavior. Neuron 86, 783–799 (2015).
Bonin, V., Histed, M.H., Yurgenson, S. & Reid, R.C. Local diversity and fine-scale organization of receptive fields in mouse visual cortex. J. Neurosci. 31, 18506–18521 (2011).
Dombeck, D.A., Khabbaz, A.N., Collman, F., Adelman, T.L. & Tank, D.W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Chen, J.L., Pfäffli, O.A., Voigt, F.F., Margolis, D.J. & Helmchen, F. Online correction of licking-induced brain motion during two-photon imaging with a tunable lens. J. Physiol. (Lond.) 591, 4689–4698 (2013).
Holtmaat, A. et al. Long-term, high-resolution imaging in the mouse neocortex through a chronic cranial window. Nat. Protoc. 4, 1128–1144 (2009).
Dubbs, A., Guevara, J. & Yuste, R. moco: fast motion correction for calcium imaging. Front. Neuroinform. 10, 6 (2016).
Thévenaz, P., Ruttimann, U.E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Greenberg, D.S. & Kerr, J.N. Automated correction of fast motion artifacts for two-photon imaging of awake animals. J. Neurosci. Methods 176, 1–15 (2009).
Wilms, C.D. & Häusser, M. Reading out a spatiotemporal population code by imaging neighbouring parallel fibre axons in vivo. Nat. Commun. 6, 6464 (2015).
Dufour, P., Piché, M., De Koninck, Y. & McCarthy, N. Two-photon excitation fluorescence microscopy with a high depth of field using an axicon. Appl. Opt. 45, 9246–9252 (2006).
Andermann, M.L., Kerlin, A.M. & Reid, R.C. Chronic cellular imaging of mouse visual cortex during operant behavior and passive viewing. Front. Cell. Neurosci. 4, 3 (2010).
Laffray, S. et al. Adaptive movement compensation for in vivo imaging of fast cellular dynamics within a moving tissue. PLoS One 6, e19928 (2011).
Kerlin, A.M., Andermann, M.L., Berezovskii, V.K. & Reid, R.C. Broadly tuned response properties of diverse inhibitory neuron subtypes in mouse visual cortex. Neuron 67, 858–871 (2010).
Andermann, M.L., Kerlin, A.M., Roumis, D.K., Glickfeld, L.L. & Reid, R.C. Functional specialization of mouse higher visual cortical areas. Neuron 72, 1025–1039 (2011).
Schultz, S.R., Kitamura, K., Post-Uiterweer, A., Krupic, J. & Häusser, M. Spatial pattern coding of sensory information by climbing fiber-evoked calcium signals in networks of neighboring cerebellar Purkinje cells. J. Neurosci. 29, 8005–8015 (2009).
Mukamel, E.A., Nimmerjahn, A. & Schnitzer, M.J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009).
Pnevmatikakis, E.A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Andilla, F.D. & Hamprecht, F.A. Sparse space-time deconvolution for calcium image analysis. Adv. Neural Inf. Process. Syst. 27, 64–72 (2014).
Pachitariu, M. et al. Suite2p: beyond 10,000 neurons with standard two-photon microscopy. bioRxiv http://dx.doi.org/10.1101/061507 (2016).
Pachitariu, M. et al. Extracting regions of interest from biological images with convolutional sparse block coding. Adv. Neural Inf. Process. Syst. 24, 1745–1753 (2013).
Vogelstein, J.T. et al. Fast nonnegative deconvolution for spike train inference from population calcium imaging. J. Neurophysiol. 104, 3691–3704 (2010).
Yaksi, E. & Friedrich, R.W. Reconstruction of firing rate changes across neuronal populations by temporally deconvolved Ca2+ imaging. Nat. Methods 3, 377–383 (2006).
Ganmore, E., Krumin, M., Rossi, L.F., Carandini, M. & Simoncelli, E.P. Direct estimation of firing rates from calcium imaging data. arXivhttp://arxiv.org/abs/1601.00364 (2016).
Harvey, C.D., Coen, P. & Tank, D.W. Choice-specific sequences in parietal cortex during a virtual-navigation decision task. Nature 484, 62–68 (2012).
Smith, S.L. & Häusser, M. Parallel processing of visual space by neighboring neurons in mouse visual cortex. Nat. Neurosci. 13, 1144–1149 (2010).
Harris, K.D., Csicsvari, J., Hirase, H., Dragoi, G. & Buzsáki, G. Organization of cell assemblies in the hippocampus. Nature 424, 552–556 (2003).
Acknowledgements
We thank N. Steinmetz and M. Pachitariu for comments on the manuscript. K.D.H. is supported by the Wellcome Trust (95668, 95669, 100154), EPSRC (K015141, I005102), MRC and Simons foundation (SCGB 325512). S.L.S. is supported by the Human Frontier Science Program (CDA00063/2012, RGP0027/2016), National Science Foundation (1450824), Whitehall Foundation, Klingenstein Foundation, McKnight Foundation, Simons Foundation (SCGB 325407SS) and the National Institutes of Health (R01NS091335, R01EY024294). R.Q.Q. is supported by the Human Frontiers Science Program (RGP0015/2013).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Harris, K., Quiroga, R., Freeman, J. et al. Improving data quality in neuronal population recordings. Nat Neurosci 19, 1165–1174 (2016). https://doi.org/10.1038/nn.4365
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4365
This article is cited by
-
Neuronal K+-Cl- cotransporter KCC2 as a promising drug target for epilepsy treatment
Acta Pharmacologica Sinica (2024)
-
Respiratory entrainment of units in the mouse parietal cortex depends on vigilance state
Pflügers Archiv - European Journal of Physiology (2023)
-
A modular architecture for organizing, processing and sharing neurophysiology data
Nature Methods (2023)
-
SmaRT2P: a software for generating and processing smart line recording trajectories for population two-photon calcium imaging
Brain Informatics (2022)
-
ABOT: an open-source online benchmarking tool for machine learning-based artefact detection and removal methods from neuronal signals
Brain Informatics (2022)