Abstract
Contrary to the long-held belief that DNA methylation of terminally differentiated cells is permanent and essentially immutable, post-mitotic neurons exhibit extensive DNA demethylation. The cellular function of active DNA demethylation in neurons, however, remains largely unknown. Tet family proteins oxidize 5-methylcytosine to initiate active DNA demethylation through the base-excision repair (BER) pathway. We found that synaptic activity bi-directionally regulates neuronal Tet3 expression. Functionally, knockdown of Tet or inhibition of BER in hippocampal neurons elevated excitatory glutamatergic synaptic transmission, whereas overexpressing Tet3 or Tet1 catalytic domain decreased it. Furthermore, dysregulation of Tet3 signaling prevented homeostatic synaptic plasticity. Mechanistically, Tet3 dictated neuronal surface GluR1 levels. RNA-seq analyses further revealed a pivotal role of Tet3 in regulating gene expression in response to global synaptic activity changes. Thus, Tet3 serves as a synaptic activity sensor to epigenetically regulate fundamental properties and meta-plasticity of neurons via active DNA demethylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Tsankova, N., Renthal, W., Kumar, A. & Nestler, E.J. Epigenetic regulation in psychiatric disorders. Nat. Rev. Neurosci. 8, 355–367 (2007).
Gräff, J., Kim, D., Dobbin, M.M. & Tsai, L.H. Epigenetic regulation of gene expression in physiological and pathological brain processes. Physiol. Rev. 91, 603–649 (2011).
Telese, F., Gamliel, A., Skowronska-Krawczyk, D., Garcia-Bassets, I. & Rosenfeld, M.G. “Seq-ing” insights into the epigenetics of neuronal gene regulation. Neuron 77, 606–623 (2013).
Shin, J., Ming, G.L. & Song, H. DNA modifications in the mammalian brain. Phil. Trans. R. Soc. Lond. B 369 (2014).
Day, J.J., Kennedy, A.J. & Sweatt, J.D. DNA methylation and its implications and accessibility for neuropsychiatric therapeutics. Annu. Rev. Pharmacol. Toxicol. 55, 591–611 (2015).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Law, J.A. & Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Martinowich, K. et al. DNA methylation-related chromatin remodeling in activity-dependent BDNF gene regulation. Science 302, 890–893 (2003).
Miller, C.A. & Sweatt, J.D. Covalent modification of DNA regulates memory formation. Neuron 53, 857–869 (2007).
Nelson, E.D., Kavalali, E.T. & Monteggia, L.M. Activity-dependent suppression of miniature neurotransmission through the regulation of DNA methylation. J. Neurosci. 28, 395–406 (2008).
Ma, D.K. et al. Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323, 1074–1077 (2009).
Feng, J. et al. Dnmt1 and Dnmt3a maintain DNA methylation and regulate synaptic function in adult forebrain neurons. Nat. Neurosci. 13, 423–430 (2010).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–434 (2011).
Guo, J.U. et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat. Neurosci. 14, 1345–1351 (2011).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Emerging roles of TET proteins and 5-hydroxymethylcytosines in active DNA demethylation and beyond. Cell Cycle 10, 2662–2668 (2011).
Guo, J.U. et al. Genome-wide antagonism between 5-hydroxymethylcytosine and DNA methylation in the adult mouse brain. Front. Biol. (Beijing) 9, 66–74 (2014).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Pollen, A.A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
Pastor, W.A., Aravind, L. & Rao, A. TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat. Rev. Mol. Cell Biol. 14, 341–356 (2013).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009).
Szulwach, K.E. et al. 5-hmC–mediated epigenetic dynamics during postnatal neurodevelopment and aging. Nat. Neurosci. 14, 1607–1616 (2011).
Yao, B. et al. Genome-wide alteration of 5-hydroxymethylcytosine in a mouse model of fragile X-associated tremor/ataxia syndrome. Hum. Mol. Genet. 23, 1095–1107 (2014).
Kaas, G.A. et al. TET1 controls CNS 5-methylcytosine hydroxylation, active DNA demethylation, gene transcription and memory formation. Neuron 79, 1086–1093 (2013).
Rudenko, A. et al. Tet1 is critical for neuronal activity-regulated gene expression and memory extinction. Neuron 79, 1109–1122 (2013).
Todarello, G. et al. Incomplete penetrance of NRXN1 deletions in families with schizophrenia. Schizophr. Res. 155, 1–7 (2014).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Hajkova, P. et al. Genome-wide reprogramming in the mouse germ line entails the base excision repair pathway. Science 329, 78–82 (2010).
Turrigiano, G.G. The self-tuning neuron: synaptic scaling of excitatory synapses. Cell 135, 422–435 (2008).
Aoto, J., Nam, C.I., Poon, M.M., Ting, P. & Chen, L. Synaptic signaling by all-trans retinoic acid in homeostatic synaptic plasticity. Neuron 60, 308–320 (2008).
Lee, H.K. Ca-permeable AMPA receptors in homeostatic synaptic plasticity. Front. Mol. Neurosci. 5, 17 (2012).
Koike, M., Iino, M. & Ozawa, S. Blocking effect of 1-naphthyl acetyl spermine on Ca2+-permeable AMPA receptors in cultured rat hippocampal neurons. Neurosci. Res. 29, 27–36 (1997).
Shepherd, J.D. et al. Arc/Arg3.1 mediates homeostatic synaptic scaling of AMPA receptors. Neuron 52, 475–484 (2006).
Gao, M. et al. A specific requirement of Arc/Arg3.1 for visual experience-induced homeostatic synaptic plasticity in mouse primary visual cortex. J. Neurosci. 30, 7168–7178 (2010).
Rial Verde, E.M., Lee-Osbourne, J., Worley, P.F., Malinow, R. & Cline, H.T. Increased expression of the immediate-early gene arc/arg3.1 reduces AMPA receptor–mediated synaptic transmission. Neuron 52, 461–474 (2006).
Poo, M.M. Neurotrophins as synaptic modulators. Nat. Rev. Neurosci. 2, 24–32 (2001).
Rutherford, L.C., Nelson, S.B. & Turrigiano, G.G. BDNF has opposite effects on the quantal amplitude of pyramidal neuron and interneuron excitatory synapses. Neuron 21, 521–530 (1998).
Madabhushi, R., Pan, L. & Tsai, L.H. DNA damage and its links to neurodegeneration. Neuron 83, 266–282 (2014).
Gu, T.P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Shen, L. et al. Tet3 and DNA replication mediate demethylation of both the maternal and paternal genomes in mouse zygotes. Cell Stem Cell 15, 459–470 (2014).
Guo, F. et al. Active and passive demethylation of male and female pronuclear DNA in the Mammalian zygote. Cell Stem Cell 15, 447–458 (2014).
Xu, Y. et al. Tet3 CXXC domain and dioxygenase activity cooperatively regulate key genes for Xenopus eye and neural development. Cell 151, 1200–1213 (2012).
Hahn, M.A. et al. Dynamics of 5-hydroxymethylcytosine and chromatin marks in Mammalian neurogenesis. Cell Reports 3, 291–300 (2013).
Zhang, R.R. et al. Tet1 regulates adult hippocampal neurogenesis and cognition. Cell Stem Cell 13, 237–245 (2013).
Ibata, K., Sun, Q. & Turrigiano, G.G. Rapid synaptic scaling induced by changes in postsynaptic firing. Neuron 57, 819–826 (2008).
Abraham, W.C. & Bear, M.F. Metaplasticity: the plasticity of synaptic plasticity. Trends Neurosci. 19, 126–130 (1996).
Song, H., Stevens, C.F. & Gage, F.H. Astroglia induce neurogenesis from adult neural stem cells. Nature 417, 39–44 (2002).
Ge, S. et al. GABA regulates synaptic integration of newly generated neurons in the adult brain. Nature 439, 589–593 (2006).
Turrigiano, G.G., Leslie, K.R., Desai, N.S., Rutherford, L.C. & Nelson, S.B. Activity-dependent scaling of quantal amplitude in neocortical neurons. Nature 391, 892–896 (1998).
Jang, M.H. et al. Secreted frizzled-related protein 3 regulates activity-dependent adult hippocampal neurogenesis. Cell Stem Cell 12, 215–223 (2013).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Song, H.J., Stevens, C.F. & Gage, F.H. Neural stem cells from adult hippocampus develop essential properties of functional CNS neurons. Nat. Neurosci. 5, 438–445 (2002).
Kim, J.Y. et al. DISC1 regulates new neuron development in the adult brain via modulation of AKT-mTOR signaling through KIAA1212. Neuron 63, 761–773 (2009).
Acknowledgements
We thank R. Huganir, P. Worley, G.C. Turrigiano and L. Chen for suggestions, members of the Ming and Song laboratories for help and critical comments, G.L. Xu for Tet3 antibodies, and L. Liu, Q. Hussaini and Y. Cai for technical support. This work was supported by the US National Institutes of Health (NS047344 to H.S., NS048271 and MH105128 to G.-l.M., NS062691 to G.C. and D.H.G.), a grant from the Simons Foundation (SFARI240011 to H.S.), the Brain and Behavior Research Foundation (H.S., G.-l.M. and Y.S.), Maryland Stem Cell Research Foundation (G.-l.M. and H.S.), and by the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (G.-l.M., D.H.G. and G.C.). J.S. was supported by a Samsung fellowship.
Author information
Authors and Affiliations
Contributions
H.Y. performed electrophysiological analyses. Y.S. performed biochemical and DNA methylation analyses. J.S. and J.U.G. performed bioinformatics analysis. C.Z. and Y.-L.W. generated AAV. F.G., D.H.G. and G.C. performed RNA-seq. G.-l.M. and H.S. designed the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Basic characterization of neurons with Tet3 knockdown, overexpression of Tet1-CD or Tet1-mCD.
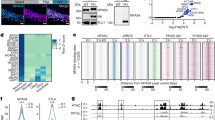
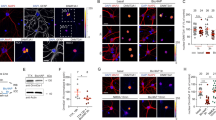
(a) Nuclear localization of Tet3 protein. Shown is a sample image of immunostaining of hippocampal neurons for Tet3, synapsin I and DAPI. Scale bar: 10 μm. (b) Schematic diagrams of AAV constructs used to infect cultured hippocampal neurons. (c) Sample images of neuronal cultures infected with AAV co-expressing EYFP and shRNAs. Scale bar: 20 μm. Note that over 99% neurons were EYFP+, indicating very high infection efficacy. Also shown is q-PCR analysis of mRNA expression of Tet1, 2, 3 after AAV-mediated expression of different shRNAs. Value represent mean + s.e.m. (n = 3; *P < 0.01; ANOVA; P = 0.03, sh-control vs sh-Tet1; P = 0.002, sh-control vs sh-Tet2; P = 0.01, sh-control vs sh-Tet3-1; P = 0.01 sh-control vs sh-Tet3-2). (d) Dot blot analysis of 5hmC levels in neurons under different conditions. Different cultures were treated with TTX (1 μM) or bicuculline (20 μM) for 48 hours before analyses. (e) Lack of differences in the excitability of neurons with or without Tet3 KD. Shown are sample current-clamp whole-cell recording traces in response to injection of different amount of currents (top panels) and summary of numbers of action potentials (bottom panel). Values represent mean + s.e.m. (f) Lack of differences in the density of synapsin 1+ boutons in neurons upon Tet3 knockdown or Tet1-CD expression. Shown on the top panel are sample confocal images of Tuj1 and synapsin I immunostaining and DAPI staining. Scale bar: 50 μm. Shown on the bottom panel is a summary of density of synapsin I+ boutons on Tuj1+ neurites. Values represent mean + s.e.m. (n > 3 cultures).
Supplementary Figure 2 Rank plots of mEPSC amplitudes to calculate scaling factors for neurons with Tet3 KD under different conditions.
(a) Tet3 KD scaled up mEPSC amplitudes multiplicatively. Shown are plots of ranked mEPSC amplitudes recorded from neurons expressing sh-Tet3-1 (left) or sh-Tet3-2 (right) against ranked mEPSC amplitudes from neurons expressing sh-control. The data are best described by a multiplicative increase in mEPSC amplitude with a scaling factor of 1.48 (sh-Tet3-1) and 1.98 (sh-Tet3-2). (b-e) Lack of significant TTX-induced scaling-up (b-c) and bicuculline-induced scaling-down (d-e) in Tet3 KD neurons. Similar as in (a), shown are plots of ranked mEPSC amplitudes recorded from neurons expressing different shRNAs with TTX (1 μM; b-c) or bicuculline treatment (20 μM; d-e) for 48 hours against ranked mEPSC amplitudes from neurons with the saline treatment. Scaling factor for each condition is listed at the top left corner. (f-g) RA (1 μm, 2 hour treatment) and Tet3 KD induced scaling-up occludes each other.
Supplementary Figure 3 Rank plots of mEPSC amplitudes to calculate scaling factors for neurons with treatment of BER inhibitors.
(a) Plots of ranked mEPSC amplitudes from neurons with treatment of ABT (50 μM, left) or CRT (50 μM, right) for 48 hours against ranked mEPSC amplitudes from neurons with the vehicle treatment. The data are best described by a multiplicative increase in mEPSC amplitude with a scaling factor of 1.40 (ABT) and 1.45 (CRT). (b-e) Lack of significant TTX-induced scaling up (b-c) and bicuculline-induced scaling-down (d-e) upon inhibition of DNA repair. Similar as in (a), shown are plots of ranked mEPSC amplitudes recorded from neurons under different drug treatments that were concurrently treated with TTX (1 μM; b-c) or bicuculline (20 μM; d-e) for 48 hours against ranked mEPSC amplitudes from those neurons without the TTX or bicuculline treatment. Scaling factor for each condition is listed at the top left corner.
Supplementary Figure 4 Rank plots of mEPSC amplitudes to calculate scaling factors for neurons with overexpression of Tet3.
(a) Plot of ranked mEPSC amplitudes from neurons co-expressing EYFP and Tet3 against ranked mEPSC amplitudes from neurons expressing EYFP alone. The data are best described by a multiplicative decrease in mEPSC amplitude with a scaling factor of 0.47. (b-e) Lack of significant TTX-induced scaling up (b-c) and bicuculline-induced scaling-down (d-e) upon Tet3 overexpression. Similar as in (a), shown are plots of ranked mEPSC amplitudes recorded from neurons over-expressing Tet3 (b, d) or EYFP (c, e) with TTX (1 μM; b-c) or bicuculline treatment (20 μM; d-e) for 48 hours against ranked mEPSC amplitudes from those neurons with the saline treatment. Scaling factor for each condition is listed at the top left corner.
Supplementary Figure 5 Rank plots of mEPSC amplitudes to calculate scaling factors for neurons with overexpression of Tet1-CD or Tet1-mCD.
(a) Plot of ranked mEPSC amplitudes from neurons co-expressing EYFP and Tet1-CD against ranked mEPSC amplitudes from neurons expressing EYFP alone with a scaling factor of 0.77. (b-e) Lack of significant TTX-induced scaling up (b-c) and bicuculline-induced scaling-down (d-e) upon Tet1-CD expression. Similar as in (a), shown are plots of ranked mEPSC amplitudes recorded from neurons over-expressing Tet1-CD (b, d) or EYFP (c, e) with TTX (1 μM; b-c) or bicuculline treatment (20 μM; d-e) for 48 hours against ranked mEPSC amplitudes from those neurons with the saline treatment. Scaling factor for each condition is listed at the top left corner.
Supplementary Figure 6 Total GluR1 and Arc protein levels under different conditions.
Shown are sample cropped Western blot images and quantification of total GluR1 (a) or Arc (b) levels under different conditions. The same biological samples as in Fig. 5c were used for a. Values represent mean + s.e.m. (n = 3 experiments; *P < 0.05; #P > 0.1; ANOVA; b: P = 0.002, sh-control vs sh-control + TTX; P = 0.007, sh-control vs sh-control + Bicu; P = 0.03, sh-control vs sh-Tet3-2). Full-length blots are presented in Supplementary Figure 11.
Supplementary Figure 7 Functional change of GluR2-lacking AMPA receptors upon changes in Tet3 expression and DNA demethylation signalling pathway.
(a) Lack of changes of surface or total GluR2 levels upon Tet3 KD or Tet1-CD expression. Shown are sample cropped Western blot images and quantification of surface and total GluR2 levels under different conditions. The same biological samples were used. Values represent mean + s.e.m. (n = 3). Full-length blots are presented in Supplementary Figure 11. (b) Effect of NASPM treatment on the mEPSC amplitude of neurons expressing sh-control or sh-Tet3-2. Values represent mean + s.e.m. (*P < 0.05; **P < 0.01; ANOVA; P = 0.03, before vs after in sh-control group; P = 0.003, before vs after in sh-Tet3-2 group). (c) Summary of decay time of mEPSCs recorded under different conditions. Same cells as in Fig. 1c-d & 2b-c. Values represent mean + s.e.m. (***P < 0.001; **P < 0.01; *P < 0.05; ANOVA; Left: P = 0.000006, sh-control vs sh-Tet3-1; P = 0.00005, sh-control vs sh-Tet3-2; Middle: P = 0.001, EYFP vs EYFP/Tet3 OE; P = 0.03, EYFP vs Tet1-CD; P = 0.02, Tet1-CD vs Tet1-mCD; right: P = 0.02, vehicle vs ABT; P = 0.002, vehicle vs CRT).
Supplementary Figure 8 RNA-seq analysis of neurons under different conditions.
(a) Confirmation of Tet3 knockdown efficacy upon AAV-mediated expression of sh-Tet3-2 in samples for RNA-seq. FPKM reads from three samples each are shown. (b) Heat-map of Pearson correlations among different RNA-seq samples. A total of 17 samples were subject to RNA-seq analysis and the top 2000 highly expressed genes were used for calculation of Pearson correlation. (c) Multidimensional scaling plot. edgeR was used for the analysis of 17 RNA-seq samples. Note clustering of biological replicates, but clear segregation of gene expression from neurons expressing sh-control and neurons expressing sh-Tet3-2. For neurons expressing sh-control, TTX and bicuculline treatment induced the opposite direction of gene expression alterations. Importantly, in Tet3-KD neurons, such changes were either attenuated (for bicuculline treatment) or abolished (for TTX treatment).
Supplementary Figure 9 Tet3 regulates expression of genes associated with synapses and synaptic transmission.
Pathway diagrams are modified from Kyoto Encyclopedia of Genes and Genomes (KEGG). Genes that exhibited significant changes of expression in Tet3-KD neurons (FDR < 0.05) are indicated by stars (red: genes activated by Tet3; blue: genes inhibited by Tet3). Data is based on RNA-seq analyses of neurons expressing sh-Tet3-2 compared to those expressing sh-control (n = 3 samples each).
Supplementary Figure 10 Lack of effect of Tet3 KD on immediate early genes.
(a) Q-PCR analysis of expression of immediate early genes in neurons expressing sh-control or sh-Tet3-2 at 4 hours after bicuculline treatment (20 μM). Values represent mean + s.e.m. (n = 3; *** P < 0.001; **P < 0.01; *P < 0.05; #P > 0.1; Student’s t-test; Arc: P = 0.02, vehicle vs Bicu in sh-control group; P = 0.01, vehicle vs Bicu in sh-Tet3-2 group; Egr4: P = 0.02, vehicle vs Bicu in sh-control group; P = 0.03, vehicle vs Bicu in sh-Tet3-2 group; c-fos: P = 0.006, vehicle vs Bicu in sh-control group; P = 0.00001, vehicle vs Bicu in sh-Tet3-2 group; Npas4: P = 0.002, vehicle vs Bicu in sh-control group; P = 0.02, vehicle vs Bicu in sh-Tet3-2 group). (b) Methylation status of Arc and Npas4 promoters and intragenic regions in neurons expressing sh-control or sh-Tet3-2. Each row represents one allele showing methylation status of individual CpG sites (open circle: unmethylated; closed circle methylated).
Supplementary Figure 11 Full length blots sample blots shown in other figures.
Shown are full blots for cropped sample gel images shown in Figures 1b, 5c, 7c and Supplementary Figures 6a, 6b and 7a.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 (PDF 1424 kb)
Supplementary Table 1
Summary of shRNA and primer sequences used in the current study. (a) Primer sequences used for qPCR analysis; (b) shRNA sequences; (c) Primer sequences for bisulfite sequencing analysis; (d) Primer sequences for ChIP-PCR analyses. (XLS 41 kb)
Supplementary Table 2
Summary of RNA-seq analysis. (a) RNA-seq run and mapping information; (b) Gene list and information on differentially expressed genes upon Tet3 knockdown. (c-d) Gene lists and information on bicuculline-regulated genes in neurons expressing sh-control (c) and those expressing sh-Tet3-2 (d). (e-f) Gene lists and information on TTX-regulated genes in neurons expressing shRNA-control (e) and those expressing sh-Tet3-2 (f). (g) KEGG pathway analysis. (XLS 904 kb)
Rights and permissions
About this article
Cite this article
Yu, H., Su, Y., Shin, J. et al. Tet3 regulates synaptic transmission and homeostatic plasticity via DNA oxidation and repair. Nat Neurosci 18, 836–843 (2015). https://doi.org/10.1038/nn.4008
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4008
This article is cited by
-
Formation of memory assemblies through the DNA-sensing TLR9 pathway
Nature (2024)
-
Multiomic analysis implicates nuclear hormone receptor signalling in clustering epilepsy
Translational Psychiatry (2024)
-
Tet Enzyme-Mediated Response in Environmental Stress and Stress-Related Psychiatric Diseases
Molecular Neurobiology (2023)
-
The threat of programmed DNA damage to neuronal genome integrity and plasticity
Nature Genetics (2022)
-
Tet3 Deletion in Adult Brain Neurons of Female Mice Results in Anxiety-like Behavior and Cognitive Impairments
Molecular Neurobiology (2022)